ZhenZhou Wang
A Comparative Review of the Histogram-based Image Segmentation Methods
Feb 23, 2025Abstract:The histogram of an image is the accurate graphical representation of the numerical grayscale distribution and it is also an estimate of the probability distribution of image pixels. Therefore, histogram has been widely adopted to calculate the clustering means and partitioning thresholds for image segmentation. There have been many classical histogram-based image segmentation methods proposed and played important roles in both academics and industry. In this article, the histories and recent advances of the histogram-based image segmentation techniques are first reviewed and then they are divided into four categories: (1), the means-based method; (2), the Gaussian-mixture-model-based method; (3), the entropy-based method and (4) the feature-points-based method. The principles of the classical histogram-based image segmentation methods are described at first and then their performances are compared objectively. In addition, the histogram-based image segmentation methods are compared with the general-purpose deep learning methods in segmenting objects with uniform or simple backgrounds. The histogram-based image segmentation methods are more accurate than the universal deep-learning methods without special training in segmenting many types of images.
Fully Automated Segmentation of the Left Ventricle in Magnetic Resonance Images
Jul 21, 2020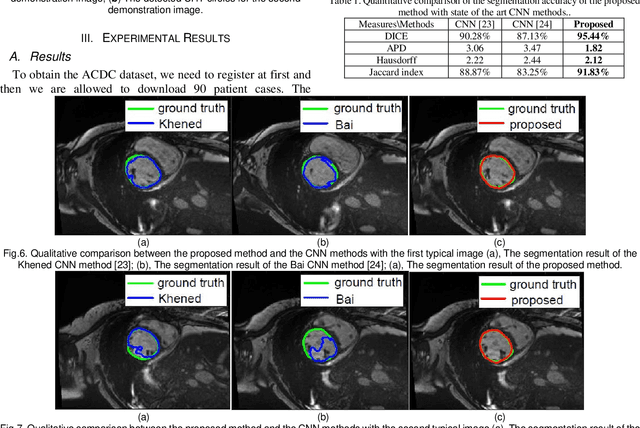
Abstract:Automatic and robust segmentation of the left ventricle (LV) in magnetic resonance images (MRI) has remained challenging for many decades. With the great success of deep learning in object detection and classification, the research focus of LV segmentation has changed to convolutional neural network (CNN) in recent years. However, LV segmentation is a pixel-level classification problem and its categories are intractable compared to object detection and classification. Although lots of CNN based methods have been proposed for LV segmentation, no robust and reproducible results are achieved yet. In this paper, we try to reproduce the CNN based LV segmentation methods with their disclosed codes and trained CNN models. Not surprisingly, the reproduced results are significantly worse than their claimed accuracies. We also proposed a fully automated LV segmentation method based on slope difference distribution (SDD) threshold selection to compare with the reproduced CNN methods. The proposed method achieved 95.44% DICE score on the test set of automated cardiac diagnosis challenge (ACDC) while the two compared CNN methods achieved 90.28% and 87.13% DICE scores. Our achieved accuracy is also higher than the best accuracy reported in the published literatures. The MATLAB codes of our proposed method are freely available on line.
Bottleneck detection by slope difference distribution: a robust approach for separating overlapped cells
Dec 11, 2019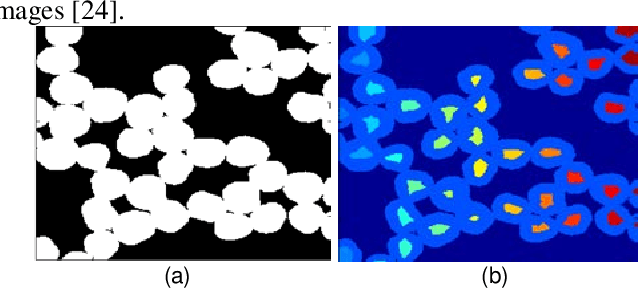
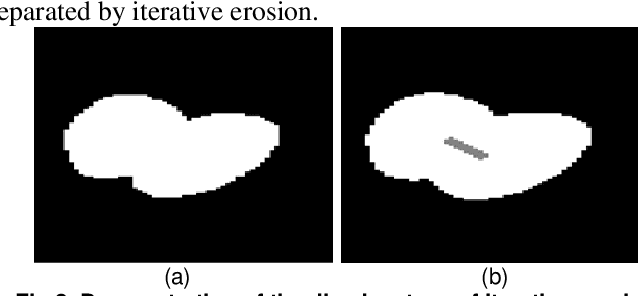
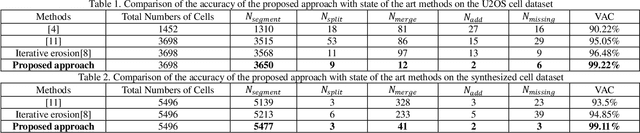

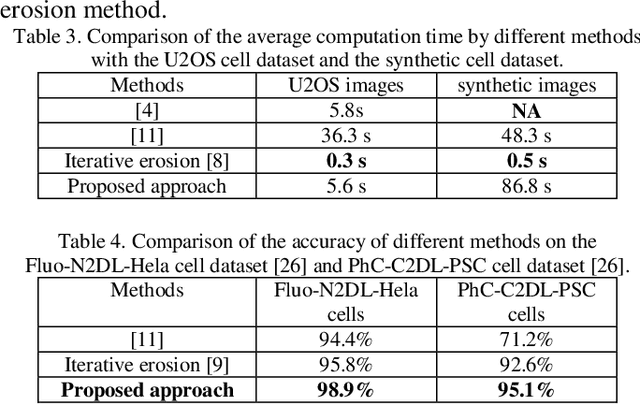
Abstract:To separate the overlapped cells, a bottleneck detection approach is proposed in this paper. The cell image is segmented by slope difference distribution (SDD) threshold selection. For each segmented binary clump, its one-dimensional boundary is computed as the distance distribution between its centroid and each point on the two-dimensional boundary. The bottleneck points of the one-dimensional boundary is detected by SDD and then transformed back into two-dimensional bottleneck points. Two largest concave parts of the binary clump are used to select the valid bottleneck points. Two bottleneck points from different concave parts with the minimum Euclidean distance is connected to separate the binary clump with minimum-cut. The binary clumps are separated iteratively until the number of computed concave parts is smaller than two. We use four types of open-accessible cell datasets to verify the effectiveness of the proposed approach and experimental results showed that the proposed approach is significantly more robust than state of the art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge