Yassien Shaalan
Once-for-All Channel Mixers (HYPERTINYPW): Generative Compression for TinyML
Mar 26, 2026Abstract:Deploying neural networks on microcontrollers is constrained by kilobytes of flash and SRAM, where 1x1 pointwise (PW) mixers often dominate memory even after INT8 quantization across vision, audio, and wearable sensing. We present HYPER-TINYPW, a compression-as-generation approach that replaces most stored PW weights with generated weights: a shared micro-MLP synthesizes PW kernels once at load time from tiny per-layer codes, caches them, and executes them with standard integer operators. This preserves commodity MCU runtimes and adds only a one-off synthesis cost; steady-state latency and energy match INT8 separable CNN baselines. Enforcing a shared latent basis across layers removes cross-layer redundancy, while keeping PW1 in INT8 stabilizes early, morphology-sensitive mixing. We contribute (i) TinyML-faithful packed-byte accounting covering generator, heads/factorization, codes, kept PW1, and backbone; (ii) a unified evaluation with validation-tuned t* and bootstrap confidence intervals; and (iii) a deployability analysis covering integer-only inference and boot versus lazy synthesis. On three ECG benchmarks (Apnea-ECG, PTB-XL, MIT-BIH), HYPER-TINYPW shifts the macro-F1 versus flash Pareto frontier: at about 225 kB it matches a roughly 1.4 MB CNN while being 6.31x smaller (84.15% fewer bytes), retaining at least 95% of large-model macro-F1. Under 32-64 kB budgets it sustains balanced detection where compact baselines degrade. The mechanism applies broadly to other 1D biosignals, on-device speech, and embedded sensing tasks where per-layer redundancy dominates, indicating a wider role for compression-as-generation in resource-constrained ML systems. Beyond ECG, HYPER-TINYPW transfers to TinyML audio: on Speech Commands it reaches 96.2% test accuracy (98.2% best validation), supporting broader applicability to embedded sensing workloads where repeated linear mixers dominate memory.
* 12 pages, 5 figures. Accepted at MLSys 2026. TinyML / on-device learning paper on hypernetwork-based compression for ECG and other 1D biosignals, with integer-only inference on commodity MCUs. Evaluated on Apnea-ECG, PTB-XL, and MIT-BIH. Camera-ready version with additional datasets, experiments, and insights will appear after May 2026
Collaborative residual learners for automatic icd10 prediction using prescribed medications
Dec 16, 2020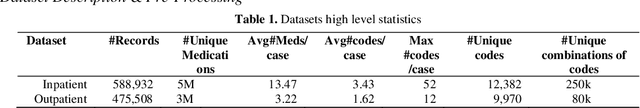
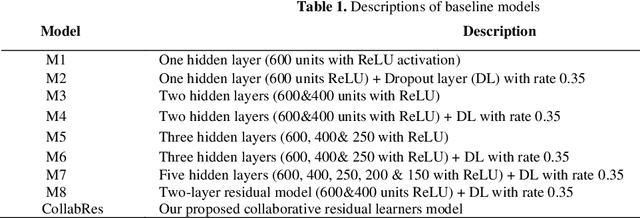
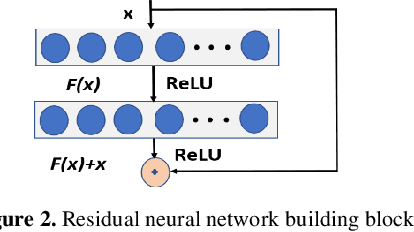
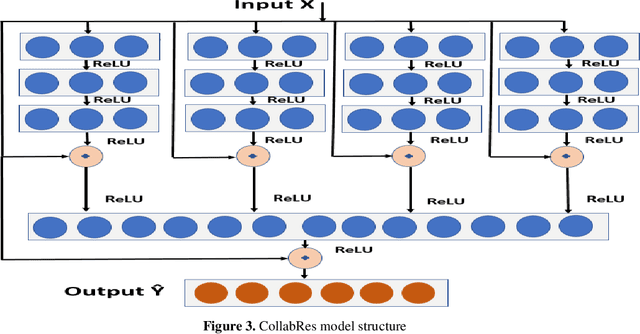
Abstract:Clinical coding is an administrative process that involves the translation of diagnostic data from episodes of care into a standard code format such as ICD10. It has many critical applications such as billing and aetiology research. The automation of clinical coding is very challenging due to data sparsity, low interoperability of digital health systems, complexity of real-life diagnosis coupled with the huge size of ICD10 code space. Related work suffer from low applicability due to reliance on many data sources, inefficient modelling and less generalizable solutions. We propose a novel collaborative residual learning based model to automatically predict ICD10 codes employing only prescriptions data. Extensive experiments were performed on two real-world clinical datasets (outpatient & inpatient) from Maharaj Nakorn Chiang Mai Hospital with real case-mix distributions. We obtain multi-label classification accuracy of 0.71 and 0.57 of average precision, 0.57 and 0.38 of F1-score and 0.73 and 0.44 of accuracy in predicting principal diagnosis for inpatient and outpatient datasets respectively.
* 6 Pages, 5 Figures and 4 tables. Presented at AIDH (Australian Institute of Digital Health) Conference 2020
Ensemble model for pre-discharge icd10 coding prediction
Dec 16, 2020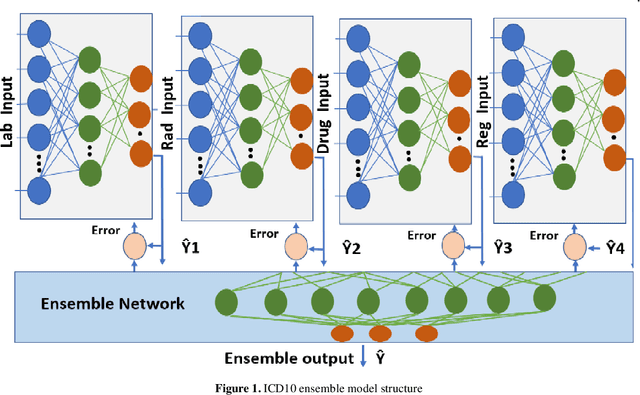
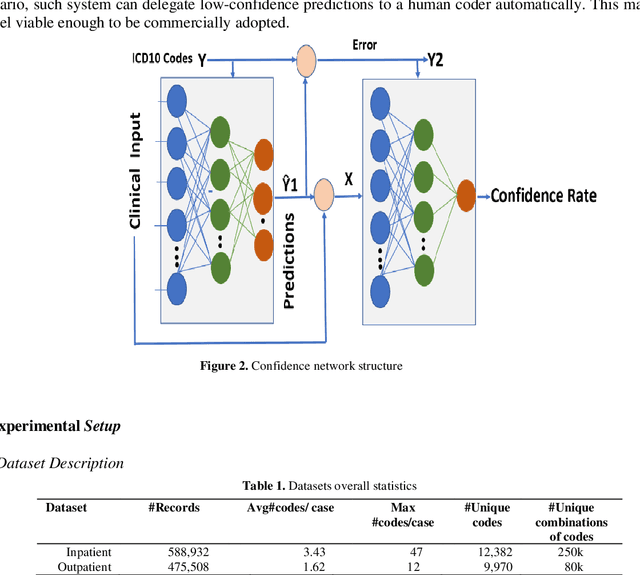
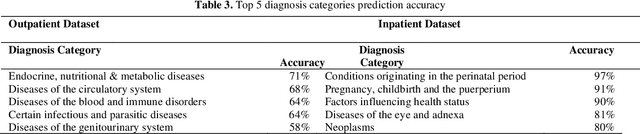
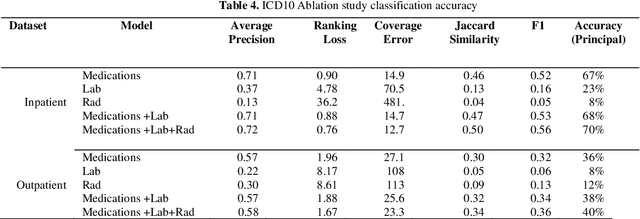
Abstract:The translation of medical diagnosis to clinical coding has wide range of applications in billing, aetiology analysis, and auditing. Currently, coding is a manual effort while the automation of such task is not straight forward. Among the challenges are the messy and noisy clinical records, case complexities, along with the huge ICD10 code space. Previous work mainly relied on discharge notes for prediction and was applied to a very limited data scale. We propose an ensemble model incorporating multiple clinical data sources for accurate code predictions. We further propose an assessment mechanism to provide confidence rates in predicted outcomes. Extensive experiments were performed on two new real-world clinical datasets (inpatient & outpatient) with unaltered case-mix distributions from Maharaj Nakorn Chiang Mai Hospital. We obtain multi-label classification accuracies of 0.73 and 0.58 for average precision, 0.56 and 0.35 for F1-scores and 0.71 and 0.4 accuracy in predicting principal diagnosis for inpatient and outpatient datasets respectively.
* 6 Pages, 2 Figures and 5 tables. Presented at AIDH (Australian Institute of Digital Health) Conference 2020
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge