Veronika Cheplygina
High-level Prior-based Loss Functions for Medical Image Segmentation: A Survey
Nov 22, 2020
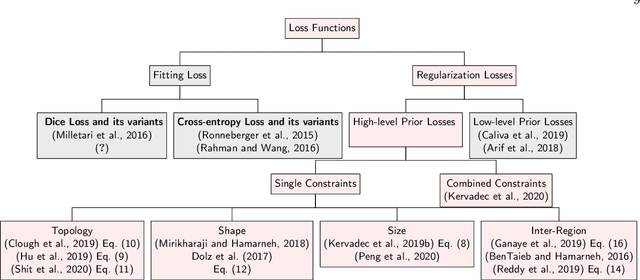
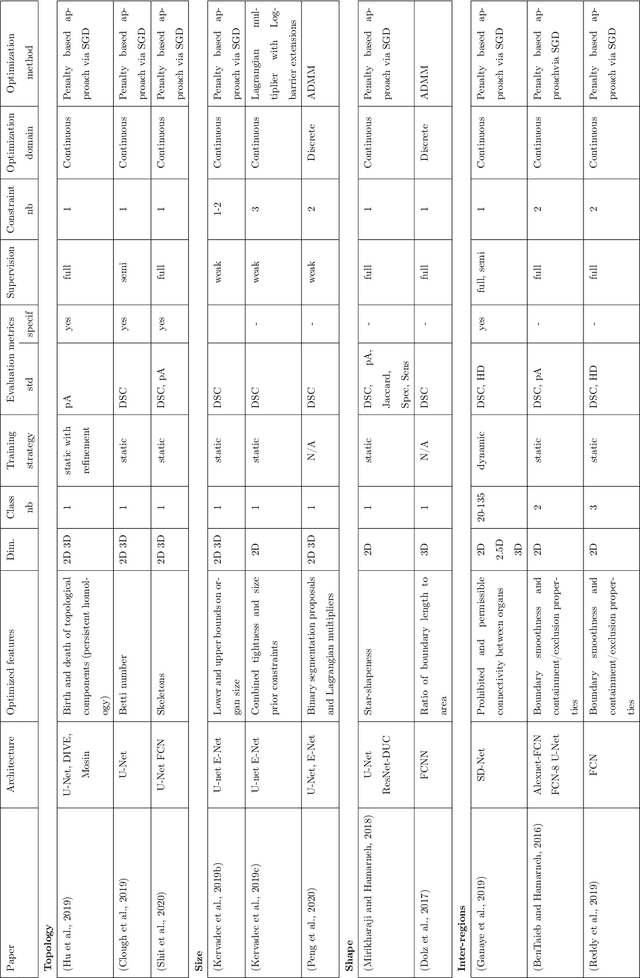
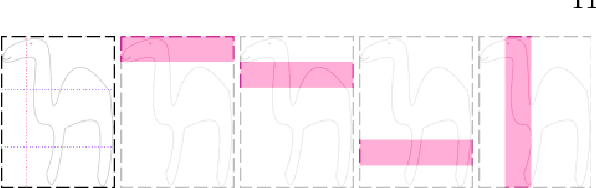
Abstract:Today, deep convolutional neural networks (CNNs) have demonstrated state of the art performance for supervised medical image segmentation, across various imaging modalities and tasks. Despite early success, segmentation networks may still generate anatomically aberrant segmentations, with holes or inaccuracies near the object boundaries. To mitigate this effect, recent research works have focused on incorporating spatial information or prior knowledge to enforce anatomically plausible segmentation. If the integration of prior knowledge in image segmentation is not a new topic in classical optimization approaches, it is today an increasing trend in CNN based image segmentation, as shown by the growing literature on the topic. In this survey, we focus on high level prior, embedded at the loss function level. We categorize the articles according to the nature of the prior: the object shape, size, topology, and the inter-regions constraints. We highlight strengths and limitations of current approaches, discuss the challenge related to the design and the integration of prior-based losses, and the optimization strategies, and draw future research directions.
Crowdsourcing Airway Annotations in Chest Computed Tomography Images
Nov 20, 2020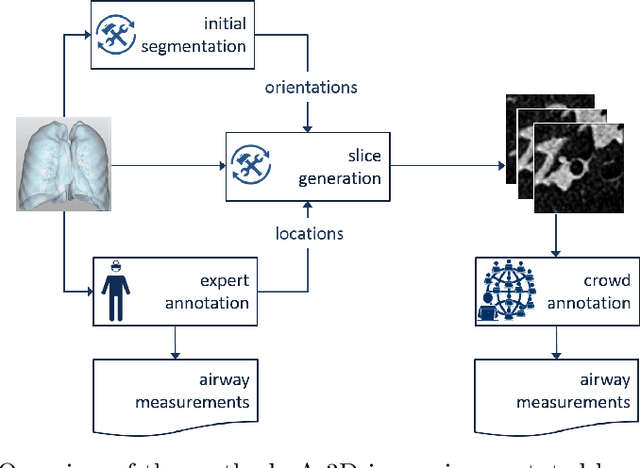
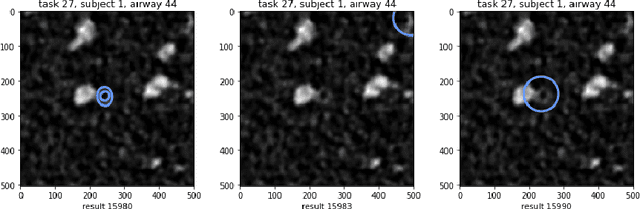
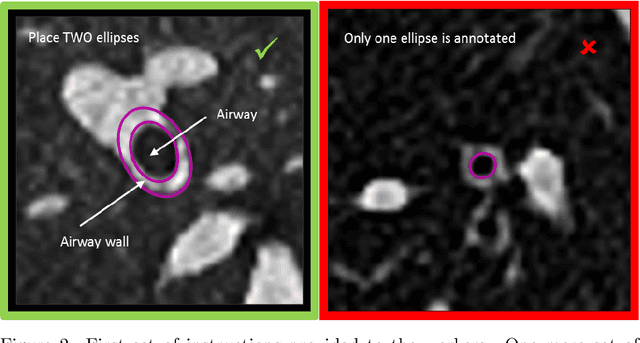
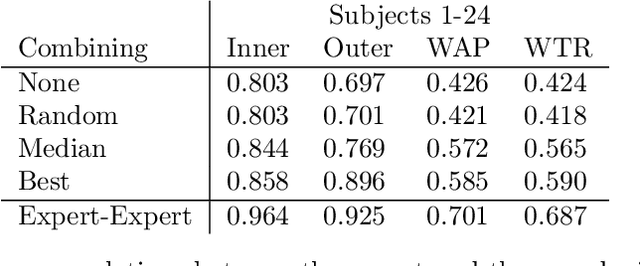

Abstract:Measuring airways in chest computed tomography (CT) scans is important for characterizing diseases such as cystic fibrosis, yet very time-consuming to perform manually. Machine learning algorithms offer an alternative, but need large sets of annotated scans for good performance. We investigate whether crowdsourcing can be used to gather airway annotations. We generate image slices at known locations of airways in 24 subjects and request the crowd workers to outline the airway lumen and airway wall. After combining multiple crowd workers, we compare the measurements to those made by the experts in the original scans. Similar to our preliminary study, a large portion of the annotations were excluded, possibly due to workers misunderstanding the instructions. After excluding such annotations, moderate to strong correlations with the expert can be observed, although these correlations are slightly lower than inter-expert correlations. Furthermore, the results across subjects in this study are quite variable. Although the crowd has potential in annotating airways, further development is needed for it to be robust enough for gathering annotations in practice. For reproducibility, data and code are available online: \url{http://github.com/adriapr/crowdairway.git}.
Primary Tumor Origin Classification of Lung Nodules in Spectral CT using Transfer Learning
Jun 30, 2020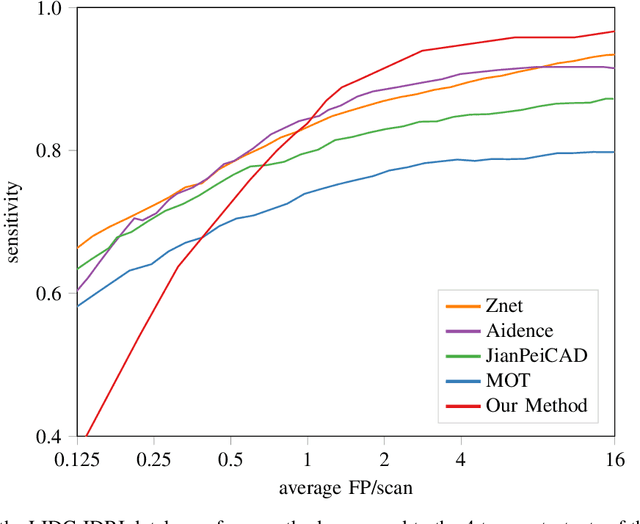
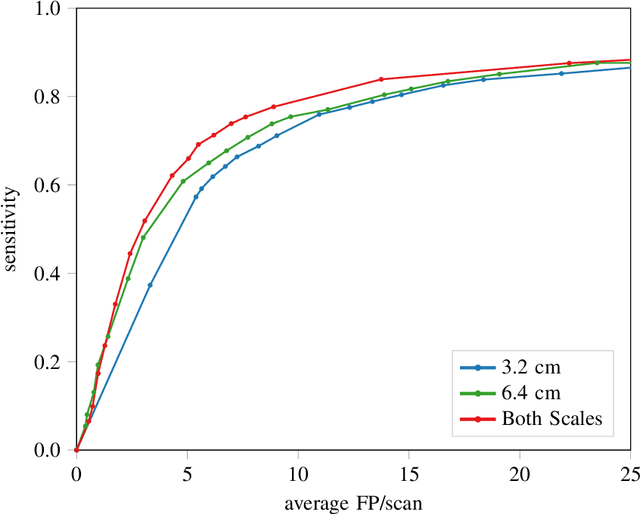
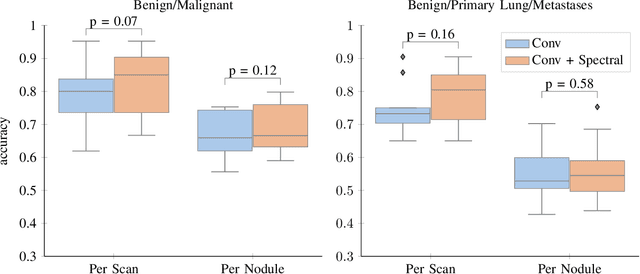
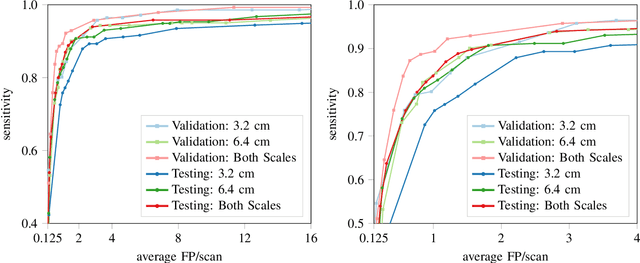
Abstract:Early detection of lung cancer has been proven to decrease mortality significantly. A recent development in computed tomography (CT), spectral CT, can potentially improve diagnostic accuracy, as it yields more information per scan than regular CT. However, the shear workload involved with analyzing a large number of scans drives the need for automated diagnosis methods. Therefore, we propose a detection and classification system for lung nodules in CT scans. Furthermore, we want to observe whether spectral images can increase classifier performance. For the detection of nodules we trained a VGG-like 3D convolutional neural net (CNN). To obtain a primary tumor classifier for our dataset we pre-trained a 3D CNN with similar architecture on nodule malignancies of a large publicly available dataset, the LIDC-IDRI dataset. Subsequently we used this pre-trained network as feature extractor for the nodules in our dataset. The resulting feature vectors were classified into two (benign/malignant) and three (benign/primary lung cancer/metastases) classes using support vector machine (SVM). This classification was performed both on nodule- and scan-level. We obtained state-of-the art performance for detection and malignancy regression on the LIDC-IDRI database. Classification performance on our own dataset was higher for scan- than for nodule-level predictions. For the three-class scan-level classification we obtained an accuracy of 78\%. Spectral features did increase classifier performance, but not significantly. Our work suggests that a pre-trained feature extractor can be used as primary tumor origin classifier for lung nodules, eliminating the need for elaborate fine-tuning of a new network and large datasets. Code is available at \url{https://github.com/tueimage/lung-nodule-msc-2018}.
Risk of Training Diagnostic Algorithms on Data with Demographic Bias
Jun 17, 2020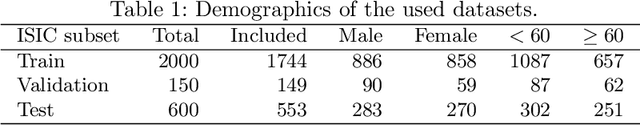
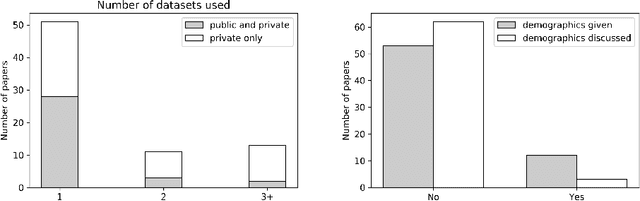
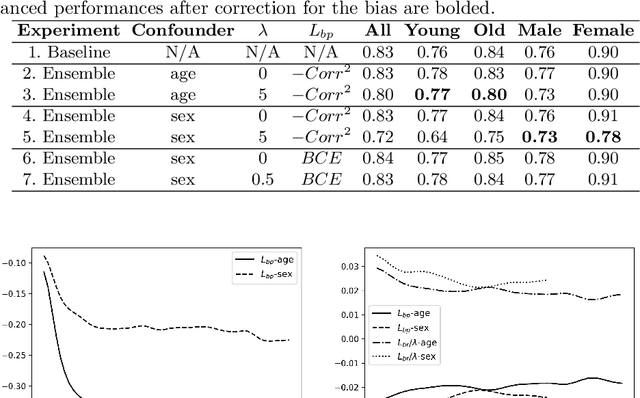
Abstract:One of the critical challenges in machine learning applications is to have fair predictions. There are numerous recent examples in various domains that convincingly show that algorithms trained with biased datasets can easily lead to erroneous or discriminatory conclusions. This is even more crucial in clinical applications where the predictive algorithms are designed mainly based on a limited or given set of medical images and demographic variables such as age, sex and race are not taken into account. In this work, we conduct a survey of the MICCAI 2018 proceedings to investigate the common practice in medical image analysis applications. Surprisingly, we found that papers focusing on diagnosis rarely describe the demographics of the datasets used, and the diagnosis is purely based on images. In order to highlight the importance of considering the demographics in diagnosis tasks, we used a publicly available dataset of skin lesions. We then demonstrate that a classifier with an overall area under the curve (AUC) of 0.83 has variable performance between 0.76 and 0.91 on subgroups based on age and sex, even though the training set was relatively balanced. Moreover, we show that it is possible to learn unbiased features by explicitly using demographic variables in an adversarial training setup, which leads to balanced scores per subgroups. Finally, we discuss the implications of these results and provide recommendations for further research.
Predicting Scores of Medical Imaging Segmentation Methods with Meta-Learning
May 08, 2020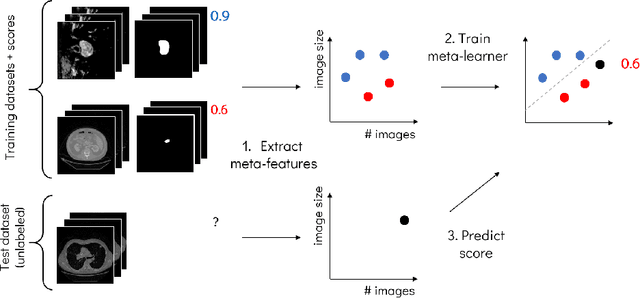
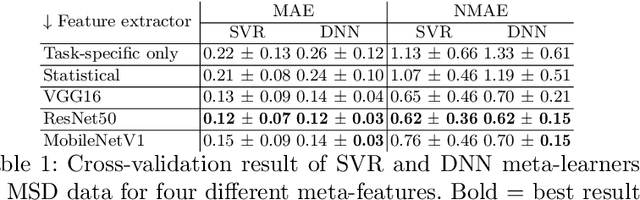

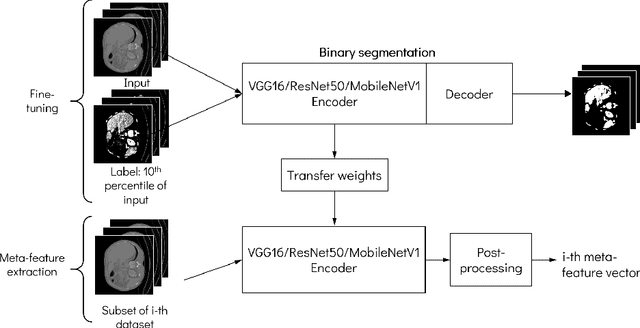
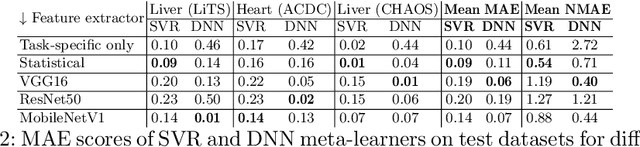
Abstract:Deep learning has led to state-of-the-art results for many medical imaging tasks, such as segmentation of different anatomical structures. With the increased numbers of deep learning publications and openly available code, the approach to choosing a model for a new task becomes more complicated, while time and (computational) resources are limited. A possible solution to choosing a model efficiently is meta-learning, a learning method in which prior performance of a model is used to predict the performance for new tasks. We investigate meta-learning for segmentation across ten datasets of different organs and modalities. We propose four ways to represent each dataset by meta-features: one based on statistical features of the images and three are based on deep learning features. We use support vector regression and deep neural networks to learn the relationship between the meta-features and prior model performance. On three external test datasets these methods give Dice scores within 0.10 of the true performance. These results demonstrate the potential of meta-learning in medical imaging.
Multi-task Learning with Crowdsourced Features Improves Skin Lesion Diagnosis
Apr 28, 2020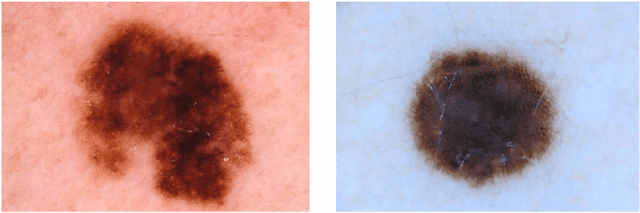
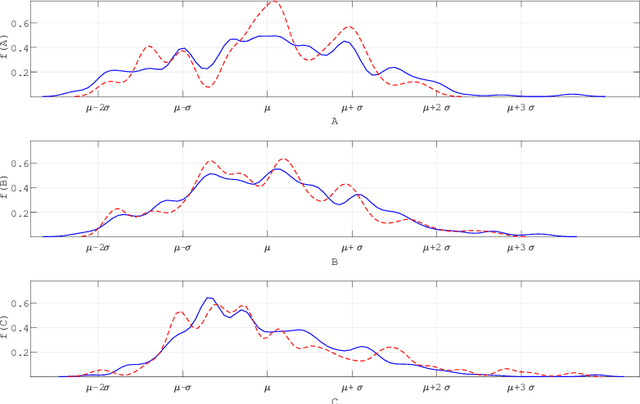
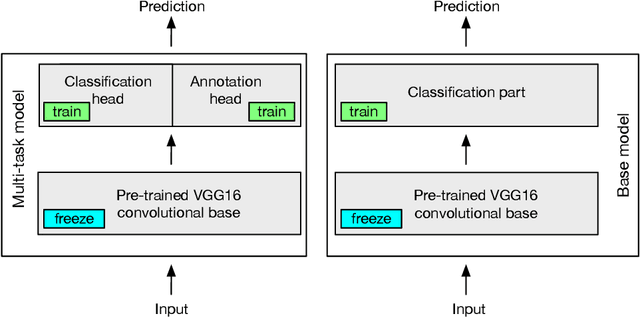
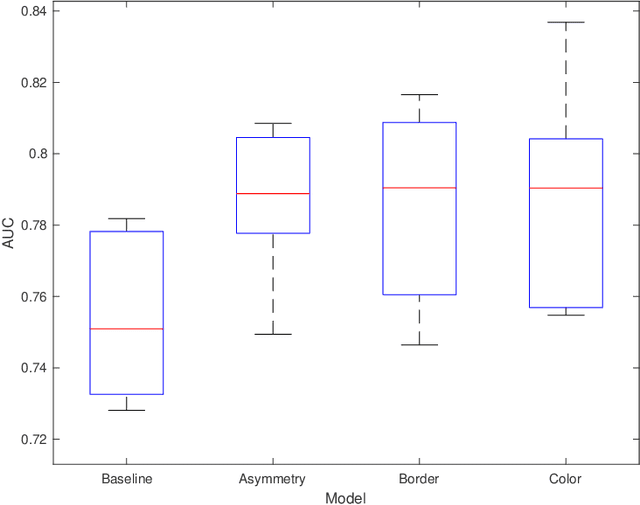
Abstract:Machine learning has a recognised need for large amounts of annotated data. Due to the high cost of expert annotations, crowdsourcing, where non-experts are asked to label or outline images, has been proposed as an alternative. Although many promising results are reported, the quality of diagnostic crowdsourced labels is still lacking. We propose to address this by instead asking the crowd about visual features of the images, which can be provided more intuitively, and by using these features in a multi-task learning framework. We compare our proposed approach to a baseline model with a set of 2000 skin lesions from the ISIC 2017 challenge dataset. The baseline model only predicts a binary label from the skin lesion image, while our multi-task model also predicts one of the following features: asymmetry of the lesion, border irregularity and color. We show that crowd features in combination with multi-task learning leads to improved generalisation. The area under the receiver operating characteristic curve is 0.754 for the baseline model and 0.782, 0.785 and 0.789 for multi-task models with border, color and asymmetry respectively. Finally, we discuss the findings, identify some limitations and recommend directions for further research.
A Survey of Crowdsourcing in Medical Image Analysis
Feb 25, 2019
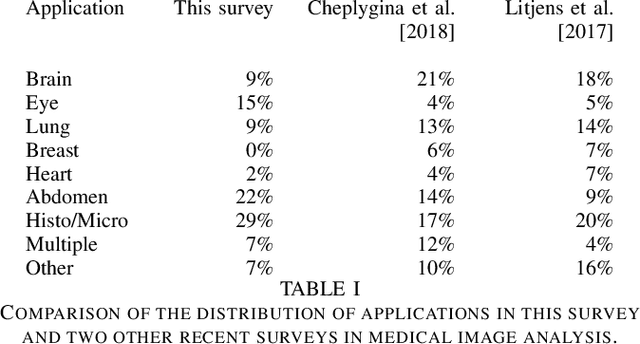

Abstract:Rapid advances in image processing capabilities have been seen across many domains, fostered by the application of machine learning algorithms to "big-data". However, within the realm of medical image analysis, advances have been curtailed, in part, due to the limited availability of large-scale, well-annotated datasets. One of the main reasons for this is the high cost often associated with producing large amounts of high-quality meta-data. Recently, there has been growing interest in the application of crowdsourcing for this purpose; a technique that has proven effective for creating large-scale datasets across a range of disciplines, from computer vision to astrophysics. Despite the growing popularity of this approach, there has not yet been a comprehensive literature review to provide guidance to researchers considering using crowdsourcing methodologies in their own medical imaging analysis. In this survey, we review studies applying crowdsourcing to the analysis of medical images, published prior to July 2018. We identify common approaches, challenges and considerations, providing guidance of utility to researchers adopting this approach. Finally, we discuss future opportunities for development within this emerging domain.
Cats or CAT scans: transfer learning from natural or medical image source datasets?
Oct 12, 2018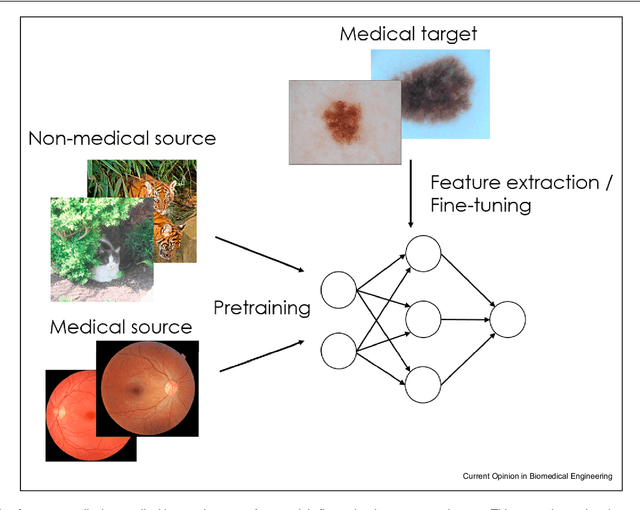
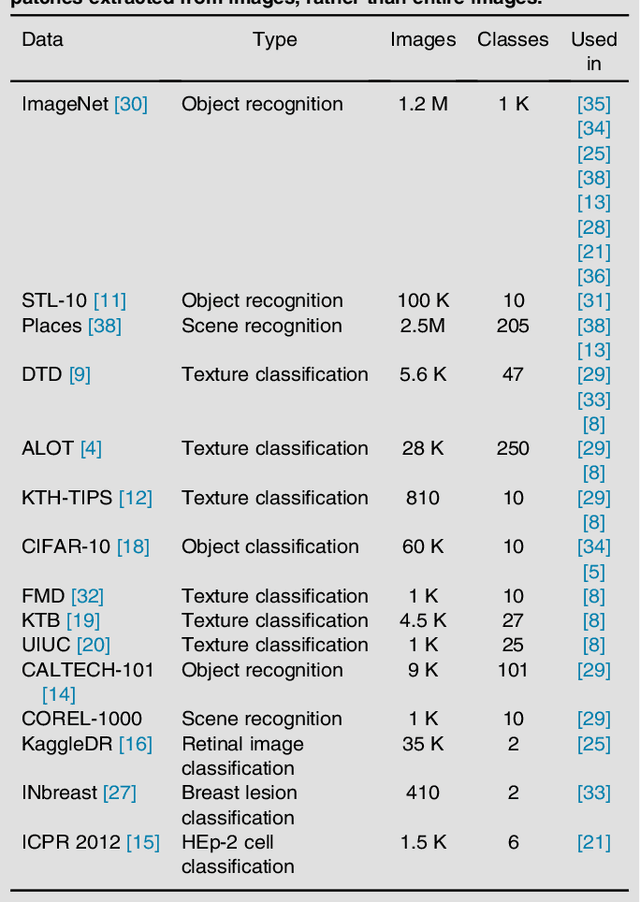
Abstract:Transfer learning is a widely used strategy in medical image analysis. Instead of only training a network with a limited amount of data from the target task of interest, we can first train the network with other, potentially larger source datasets, creating a more robust model. The source datasets do not have to be related to the target task. For a classification task in lung CT images, we could use both head CT images, or images of cats, as the source. While head CT images appear more similar to lung CT images, the number and diversity of cat images might lead to a better model overall. In this survey we review a number of papers that have performed similar comparisons. Although the answer to which strategy is best seems to be "it depends", we discuss a number of research directions we need to take as a community, to gain more understanding of this topic.
Automatic Emphysema Detection using Weakly Labeled HRCT Lung Images
Oct 01, 2018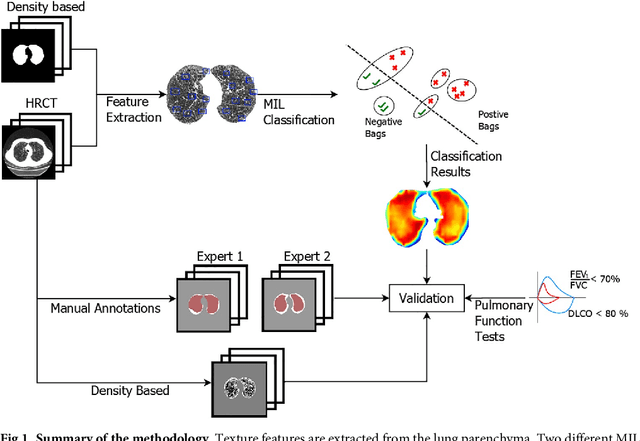
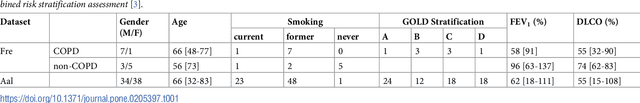
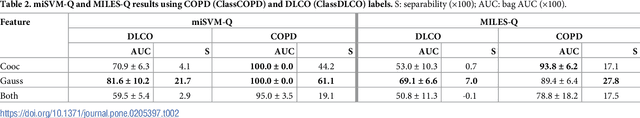
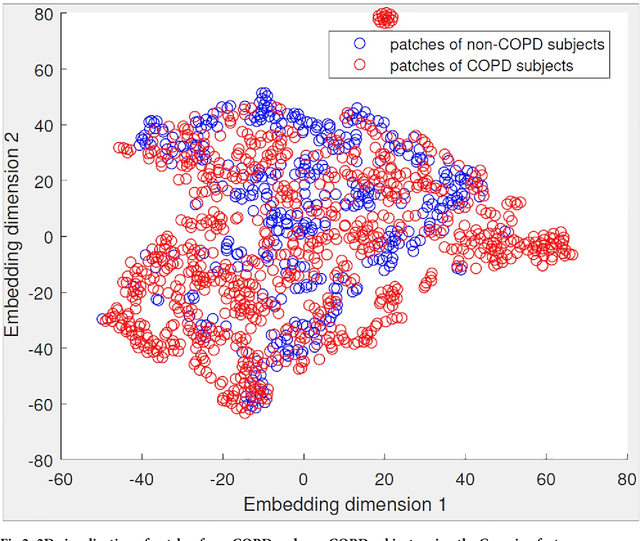
Abstract:A method for automatically quantifying emphysema regions using High-Resolution Computed Tomography (HRCT) scans of patients with chronic obstructive pulmonary disease (COPD) that does not require manually annotated scans for training is presented. HRCT scans of controls and of COPD patients with diverse disease severity are acquired at two different centers. Textural features from co-occurrence matrices and Gaussian filter banks are used to characterize the lung parenchyma in the scans. Two robust versions of multiple instance learning (MIL) classifiers, miSVM and MILES, are investigated. The classifiers are trained with the weak labels extracted from the forced expiratory volume in one minute (FEV$_1$) and diffusing capacity of the lungs for carbon monoxide (DLCO). At test time, the classifiers output a patient label indicating overall COPD diagnosis and local labels indicating the presence of emphysema. The classifier performance is compared with manual annotations by two radiologists, a classical density based method, and pulmonary function tests (PFTs). The miSVM classifier performed better than MILES on both patient and emphysema classification. The classifier has a stronger correlation with PFT than the density based method, the percentage of emphysema in the intersection of annotations from both radiologists, and the percentage of emphysema annotated by one of the radiologists. The correlation between the classifier and the PFT is only outperformed by the second radiologist. The method is therefore promising for facilitating assessment of emphysema and reducing inter-observer variability.
Not-so-supervised: a survey of semi-supervised, multi-instance, and transfer learning in medical image analysis
Sep 14, 2018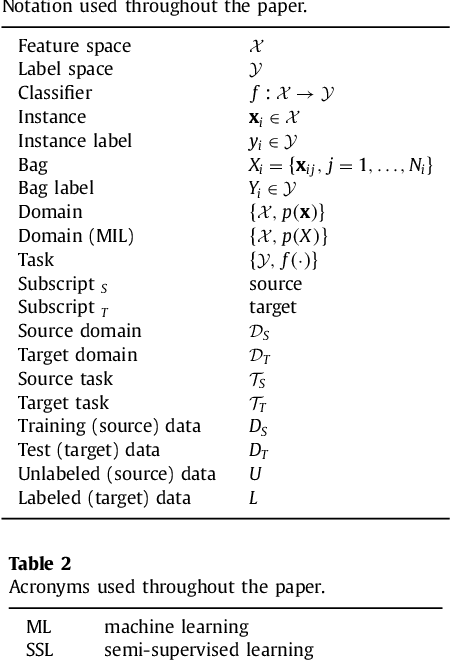
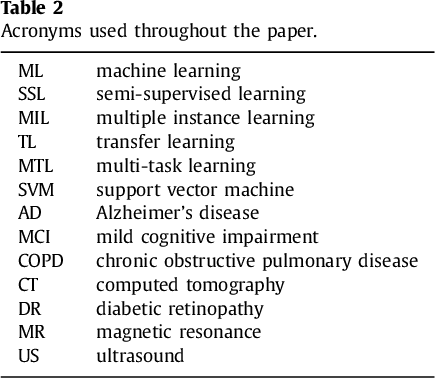
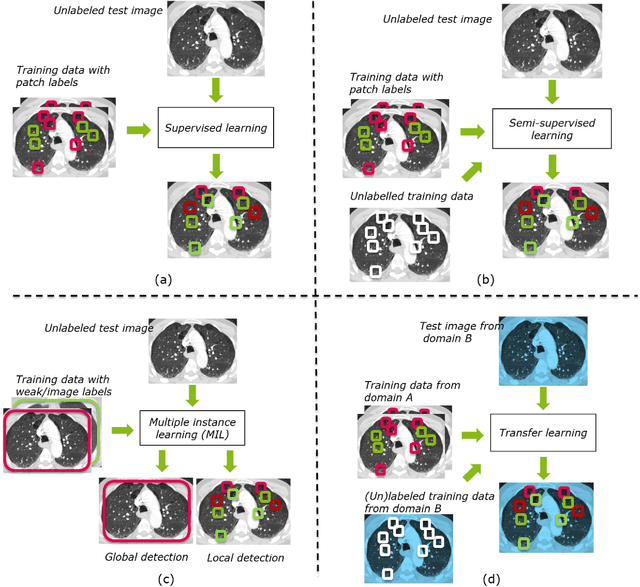

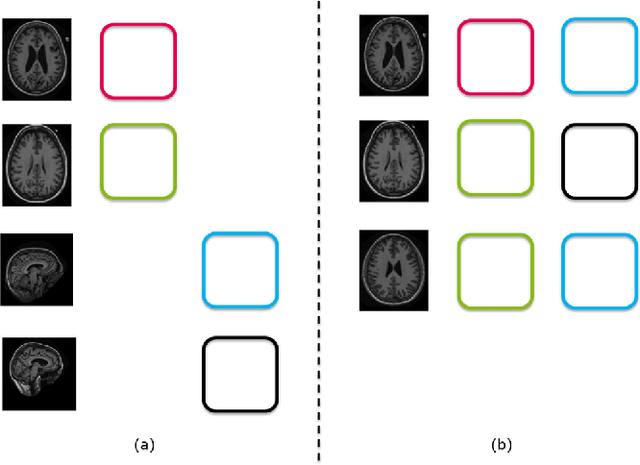
Abstract:Machine learning (ML) algorithms have made a tremendous impact in the field of medical imaging. While medical imaging datasets have been growing in size, a challenge for supervised ML algorithms that is frequently mentioned is the lack of annotated data. As a result, various methods which can learn with less/other types of supervision, have been proposed. We review semi-supervised, multiple instance, and transfer learning in medical imaging, both in diagnosis/detection or segmentation tasks. We also discuss connections between these learning scenarios, and opportunities for future research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge