Umaseh Sivanesan
TricycleGAN: Unsupervised Image Synthesis and Segmentation Based on Shape Priors
Feb 04, 2021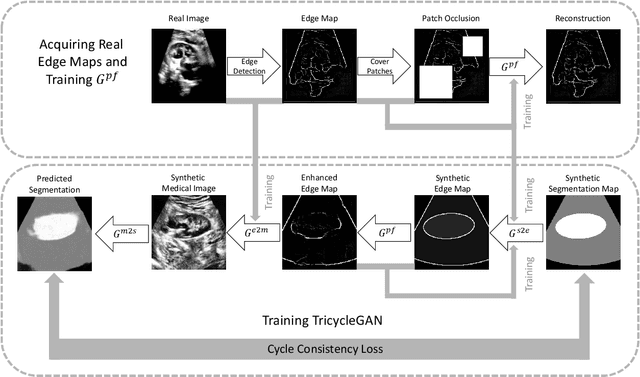
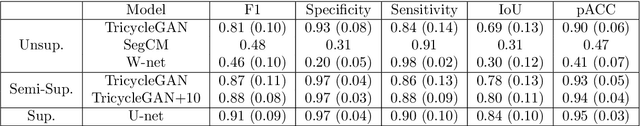
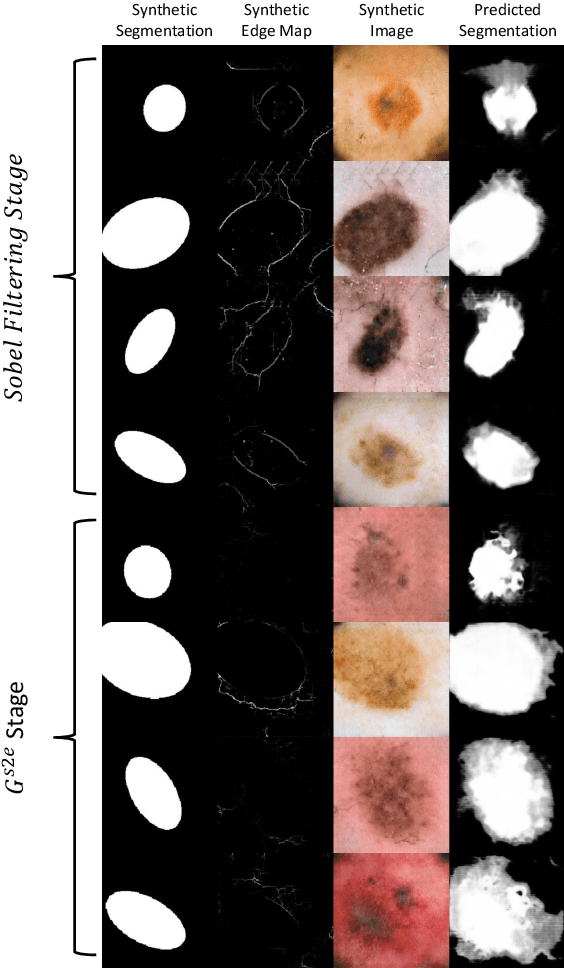
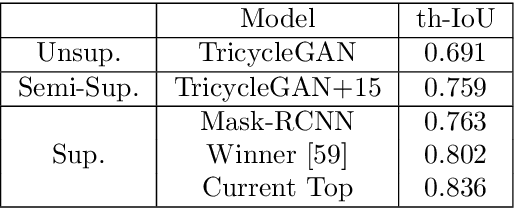
Abstract:Medical image segmentation is routinely performed to isolate regions of interest, such as organs and lesions. Currently, deep learning is the state of the art for automatic segmentation, but is usually limited by the need for supervised training with large datasets that have been manually segmented by trained clinicians. The goal of semi-superised and unsupervised image segmentation is to greatly reduce, or even eliminate, the need for training data and therefore to minimze the burden on clinicians when training segmentation models. To this end we introduce a novel network architecture for capable of unsupervised and semi-supervised image segmentation called TricycleGAN. This approach uses three generative models to learn translations between medical images and segmentation maps using edge maps as an intermediate step. Distinct from other approaches based on generative networks, TricycleGAN relies on shape priors rather than colour and texture priors. As such, it is particularly well-suited for several domains of medical imaging, such as ultrasound imaging, where commonly used visual cues may be absent. We present experiments with TricycleGAN on a clinical dataset of kidney ultrasound images and the benchmark ISIC 2018 skin lesion dataset.
Unsupervised Medical Image Segmentation with Adversarial Networks: From Edge Diagrams to Segmentation Maps
Nov 12, 2019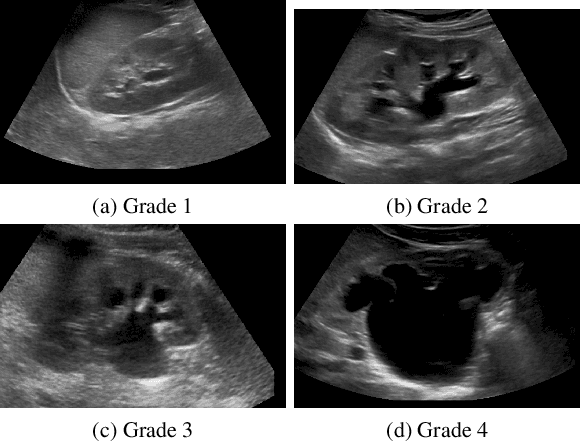
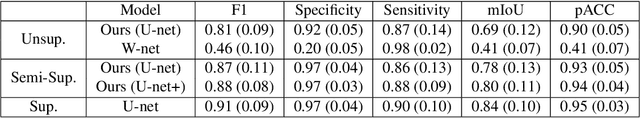
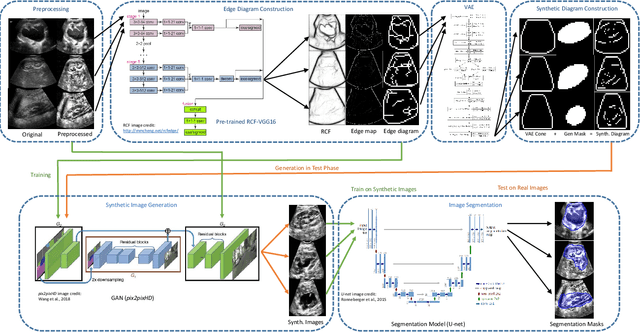
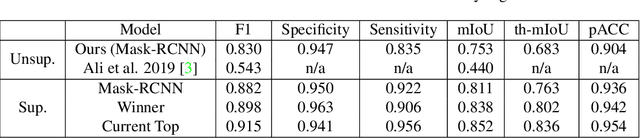
Abstract:We develop and approach to unsupervised semantic medical image segmentation that extends previous work with generative adversarial networks. We use existing edge detection methods to construct simple edge diagrams, train a generative model to convert them into synthetic medical images, and construct a dataset of synthetic images with known segmentations using variations on extracted edge diagrams. This synthetic dataset is then used to train a supervised image segmentation model. We test our approach on a clinical dataset of kidney ultrasound images and the benchmark ISIC 2018 skin lesion dataset. We show that our unsupervised approach is more accurate than previous unsupervised methods, and performs reasonably compared to supervised image segmentation models. All code and trained models are available at https://github.com/kiretd/Unsupervised-MIseg.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge