Truyen Tran
Learning to Abstract and Predict Human Actions
Aug 20, 2020
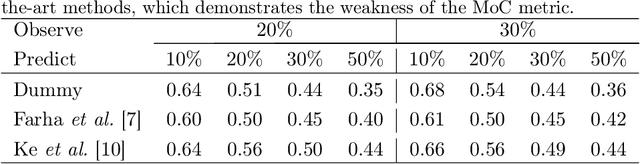
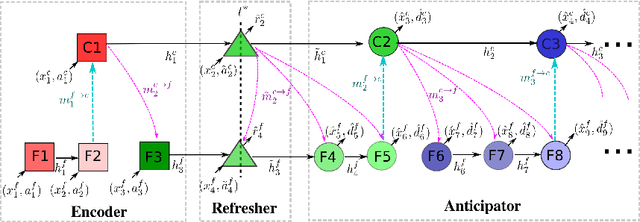
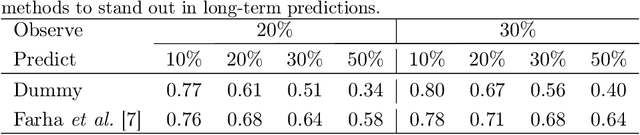
Abstract:Human activities are naturally structured as hierarchies unrolled over time. For action prediction, temporal relations in event sequences are widely exploited by current methods while their semantic coherence across different levels of abstraction has not been well explored. In this work we model the hierarchical structure of human activities in videos and demonstrate the power of such structure in action prediction. We propose Hierarchical Encoder-Refresher-Anticipator, a multi-level neural machine that can learn the structure of human activities by observing a partial hierarchy of events and roll-out such structure into a future prediction in multiple levels of abstraction. We also introduce a new coarse-to-fine action annotation on the Breakfast Actions videos to create a comprehensive, consistent, and cleanly structured video hierarchical activity dataset. Through our experiments, we examine and rethink the settings and metrics of activity prediction tasks toward unbiased evaluation of prediction systems, and demonstrate the role of hierarchical modeling toward reliable and detailed long-term action forecasting.
HyperVAE: A Minimum Description Length Variational Hyper-Encoding Network
May 18, 2020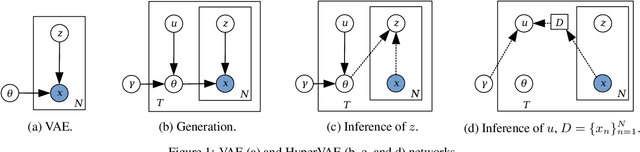
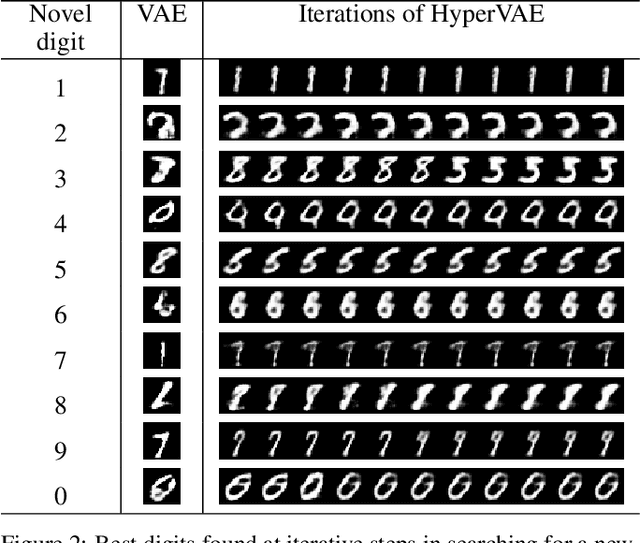
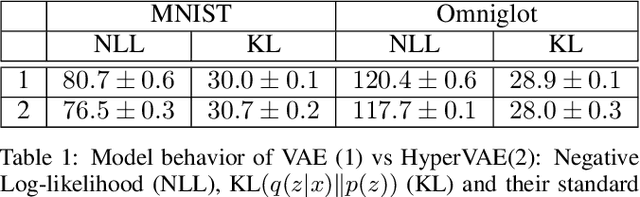

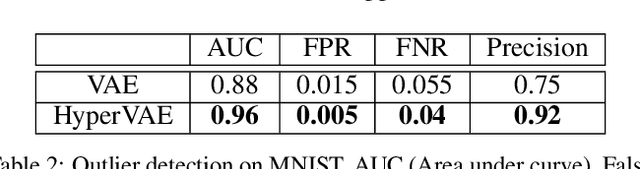
Abstract:We propose a framework called HyperVAE for encoding distributions of distributions. When a target distribution is modeled by a VAE, its neural network parameters \theta is drawn from a distribution p(\theta) which is modeled by a hyper-level VAE. We propose a variational inference using Gaussian mixture models to implicitly encode the parameters \theta into a low dimensional Gaussian distribution. Given a target distribution, we predict the posterior distribution of the latent code, then use a matrix-network decoder to generate a posterior distribution q(\theta). HyperVAE can encode the parameters \theta in full in contrast to common hyper-networks practices, which generate only the scale and bias vectors as target-network parameters. Thus HyperVAE preserves much more information about the model for each task in the latent space. We discuss HyperVAE using the minimum description length (MDL) principle and show that it helps HyperVAE to generalize. We evaluate HyperVAE in density estimation tasks, outlier detection and discovery of novel design classes, demonstrating its efficacy.
Dynamic Language Binding in Relational Visual Reasoning
Apr 30, 2020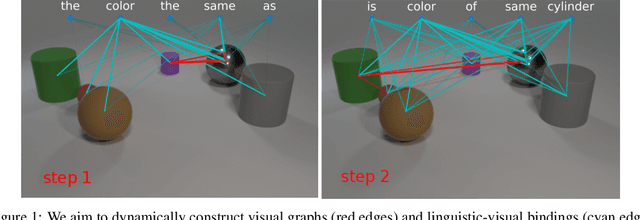
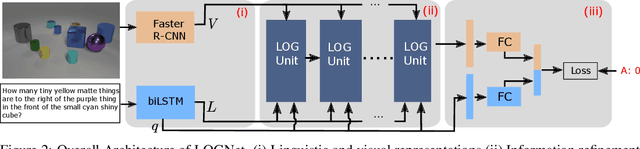
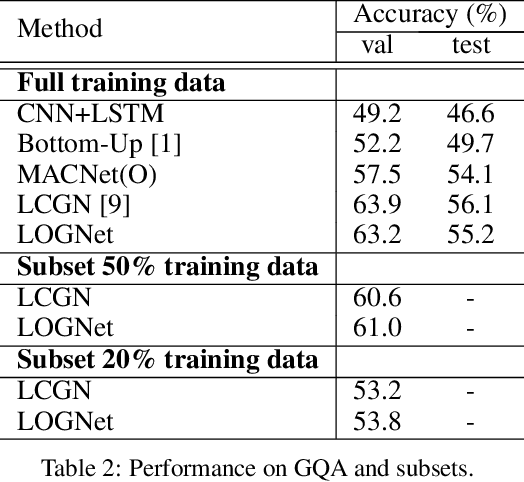
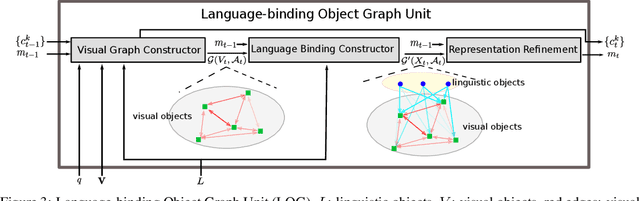
Abstract:We present Language-binding Object Graph Network, the first neural reasoning method with dynamic relational structures across both visual and textual domains with applications in visual question answering. Relaxing the common assumption made by current models that the object predicates pre-exist and stay static, passive to the reasoning process, we propose that these dynamic predicates expand across the domain borders to include pair-wise visual-linguistic object binding. In our method, these contextualized object links are actively found within each recurrent reasoning step without relying on external predicative priors. These dynamic structures reflect the conditional dual-domain object dependency given the evolving context of the reasoning through co-attention. Such discovered dynamic graphs facilitate multi-step knowledge combination and refinements that iteratively deduce the compact representation of the final answer. The effectiveness of this model is demonstrated on image question answering demonstrating favorable performance on major VQA datasets. Our method outperforms other methods in sophisticated question-answering tasks wherein multiple object relations are involved. The graph structure effectively assists the progress of training, and therefore the network learns efficiently compared to other reasoning models.
Hierarchical Conditional Relation Networks for Video Question Answering
Mar 17, 2020
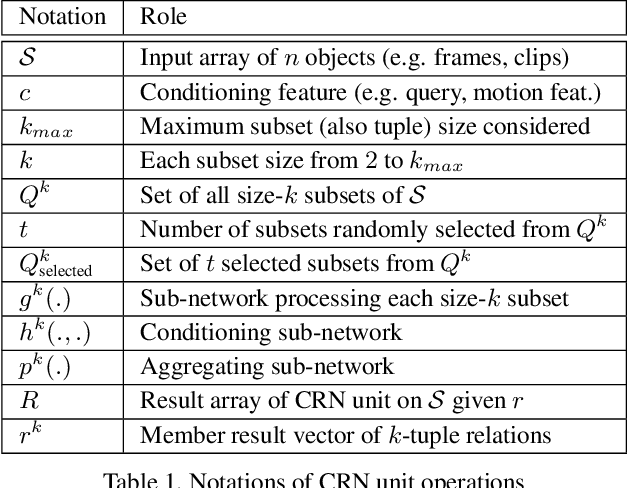
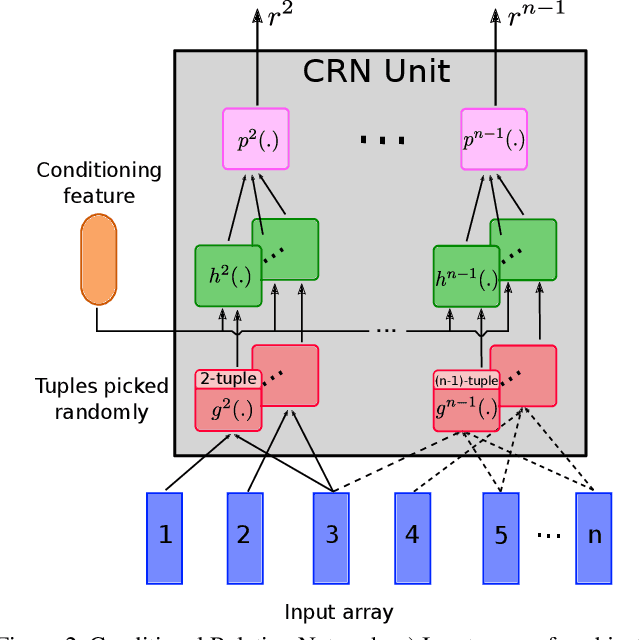
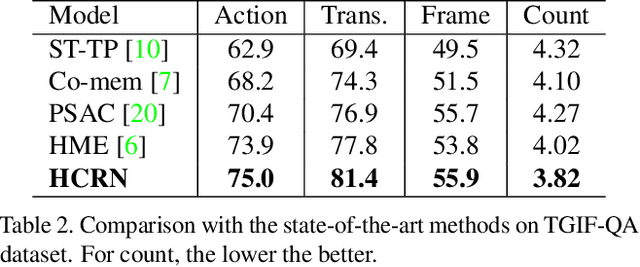
Abstract:Video question answering (VideoQA) is challenging as it requires modeling capacity to distill dynamic visual artifacts and distant relations and to associate them with linguistic concepts. We introduce a general-purpose reusable neural unit called Conditional Relation Network (CRN) that serves as a building block to construct more sophisticated structures for representation and reasoning over video. CRN takes as input an array of tensorial objects and a conditioning feature, and computes an array of encoded output objects. Model building becomes a simple exercise of replication, rearrangement and stacking of these reusable units for diverse modalities and contextual information. This design thus supports high-order relational and multi-step reasoning. The resulting architecture for VideoQA is a CRN hierarchy whose branches represent sub-videos or clips, all sharing the same question as the contextual condition. Our evaluations on well-known datasets achieved new SoTA results, demonstrating the impact of building a general-purpose reasoning unit on complex domains such as VideoQA.
* Check out our code on GitHub at https://github.com/thaolmk54/hcrn-videoqa
Self-Attentive Associative Memory
Feb 11, 2020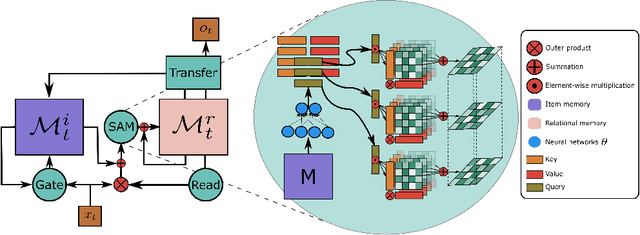
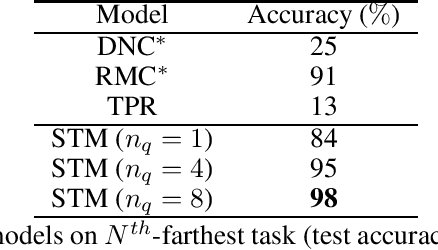
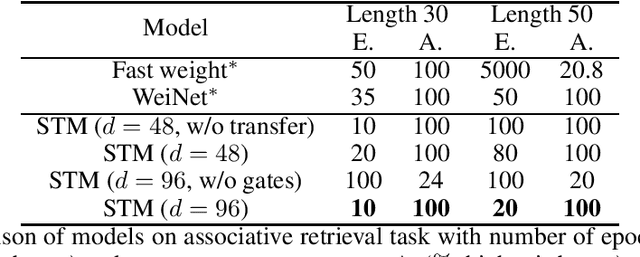

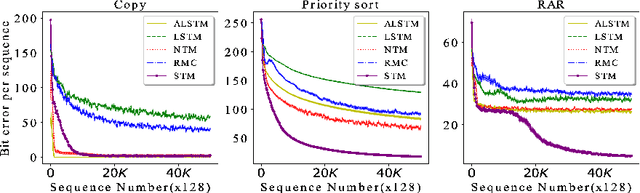
Abstract:Heretofore, neural networks with external memory are restricted to single memory with lossy representations of memory interactions. A rich representation of relationships between memory pieces urges a high-order and segregated relational memory. In this paper, we propose to separate the storage of individual experiences (item memory) and their occurring relationships (relational memory). The idea is implemented through a novel Self-attentive Associative Memory (SAM) operator. Found upon outer product, SAM forms a set of associative memories that represent the hypothetical high-order relationships between arbitrary pairs of memory elements, through which a relational memory is constructed from an item memory. The two memories are wired into a single sequential model capable of both memorization and relational reasoning. We achieve competitive results with our proposed two-memory model in a diversity of machine learning tasks, from challenging synthetic problems to practical testbeds such as geometry, graph, reinforcement learning, and question answering.
Learning Transferable Domain Priors for Safe Exploration in Reinforcement Learning
Sep 11, 2019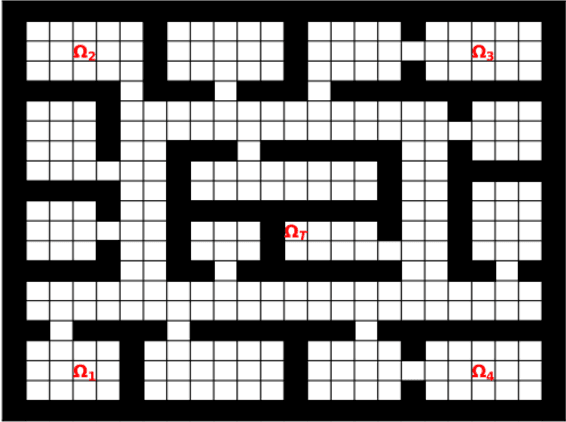
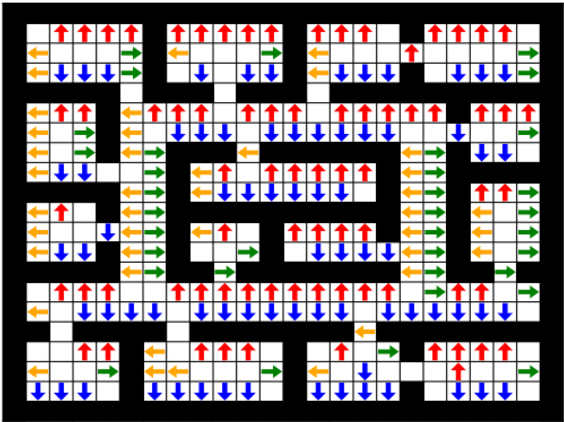
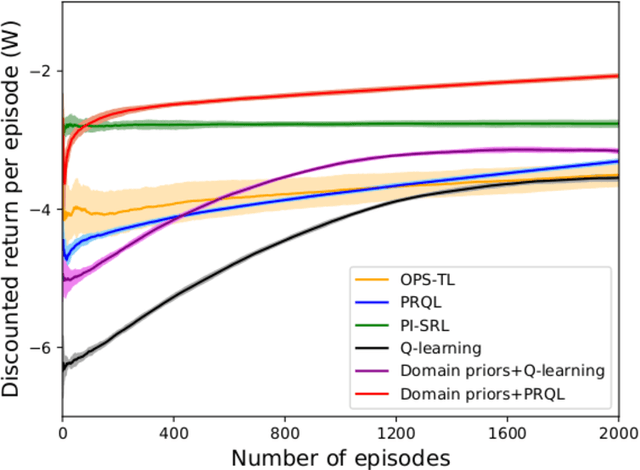
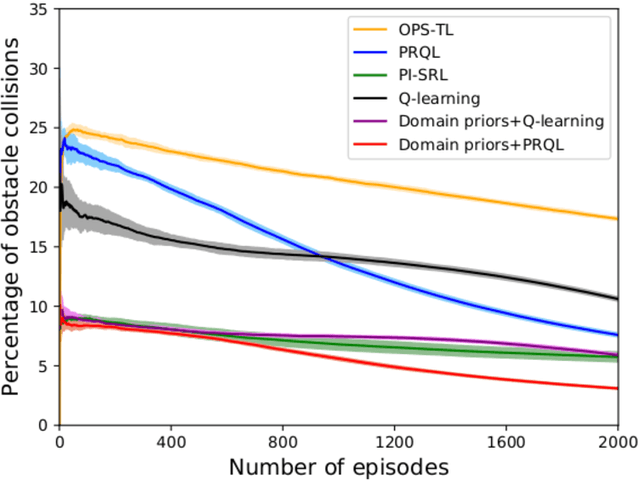
Abstract:Prior access to domain knowledge could significantly improve the performance of a reinforcement learning agent. In particular, it could help agents avoid potentially catastrophic exploratory actions, which would otherwise have to be experienced during learning. In this work, we identify consistently undesirable actions in a set of previously learned tasks, and use pseudo-rewards associated with them to learn a prior policy. In addition to enabling safe exploratory behaviors in subsequent tasks in the domain, these priors are transferable to similar environments, and can be learned off-policy and in parallel with the learning of other tasks in the domain. We compare our approach to established, state-of-the-art algorithms in a grid-world navigation environment, and demonstrate that it exhibits a superior performance with respect to avoiding unsafe actions while learning to perform arbitrary tasks in the domain. We also present some theoretical analysis to support these results, and discuss the implications and some alternative formulations of this approach, which could also be useful to accelerate learning in certain scenarios.
Theory and Evaluation Metrics for Learning Disentangled Representations
Aug 26, 2019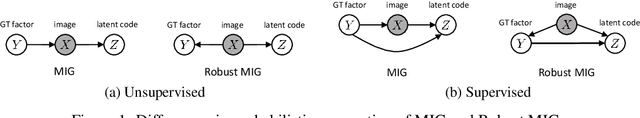
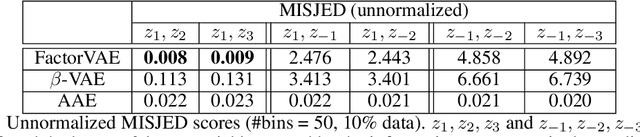
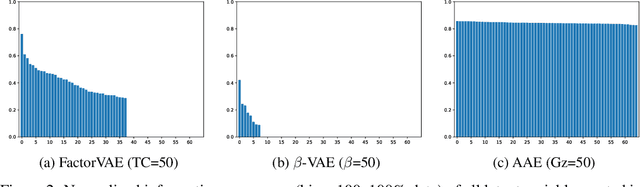
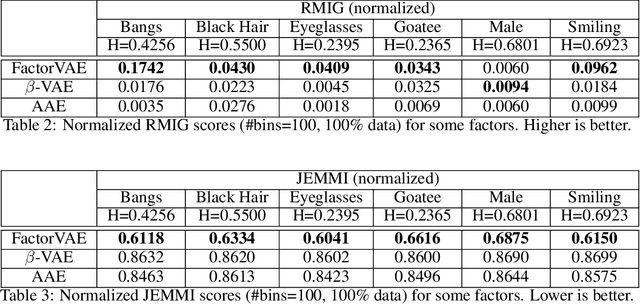
Abstract:We make two theoretical contributions to disentanglement learning by (a) defining precise semantics of disentangled representations, and (b) establishing robust metrics for evaluation. First, we characterize the concept "disentangled representations" used in supervised and unsupervised methods along three dimensions-informativeness, separability and interpretability - which can be expressed and quantified explicitly using information-theoretic constructs. This helps explain the behaviors of several well-known disentanglement learning models. We then propose robust metrics for measuring informativeness, separability and interpretability. Through a comprehensive suite of experiments, we show that our metrics correctly characterize the representations learned by different methods and are consistent with qualitative (visual) results. Thus, the metrics allow disentanglement learning methods to be compared on a fair ground. We also empirically uncovered new interesting properties of VAE-based methods and interpreted them with our formulation. These findings are promising and hopefully will encourage the design of more theoretically driven models for learning disentangled representations.
Learning to Reason with Relational Video Representation for Question Answering
Jul 10, 2019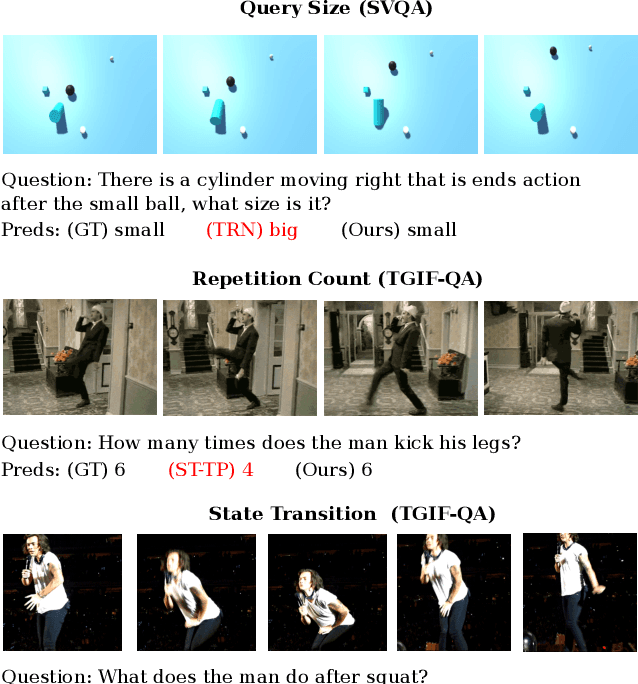
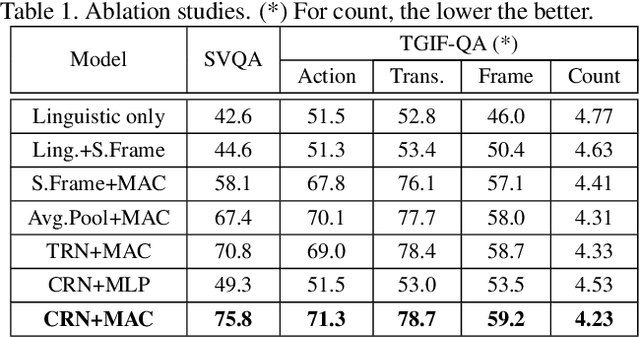
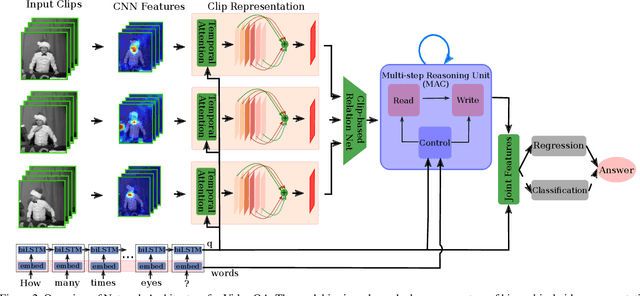
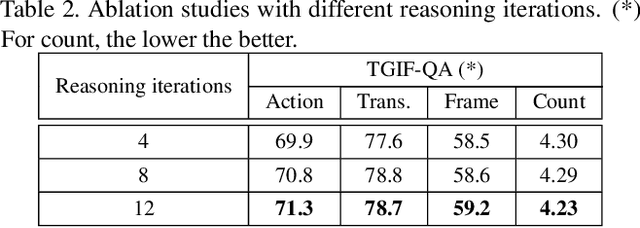
Abstract:How does machine learn to reason about the content of a video in answering a question? A Video QA system must simultaneously understand language, represent visual content over space-time, and iteratively transform these representations in response to lingual content in the query, and finally arriving at a sensible answer. While recent advances in textual and visual question answering have come up with sophisticated visual representation and neural reasoning mechanisms, major challenges in Video QA remain on dynamic grounding of concepts, relations and actions to support the reasoning process. We present a new end-to-end layered architecture for Video QA, which is composed of a question-guided video representation layer and a generic reasoning layer to produce answer. The video is represented using a hierarchical model that encodes visual information about objects, actions and relations in space-time given the textual cues from the question. The encoded representation is then passed to a reasoning module, which in this paper, is implemented as a MAC net. The system is evaluated on the SVQA (synthetic) and TGIF-QA datasets (real), demonstrating state-of-the-art results, with a large margin in the case of multi-step reasoning.
Learning Regularity in Skeleton Trajectories for Anomaly Detection in Videos
Mar 08, 2019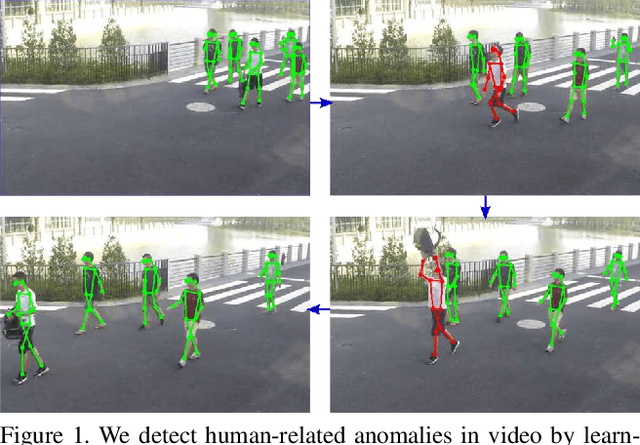
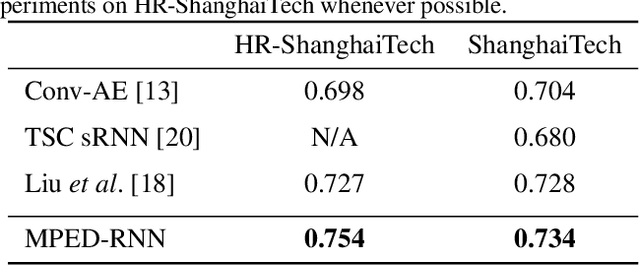
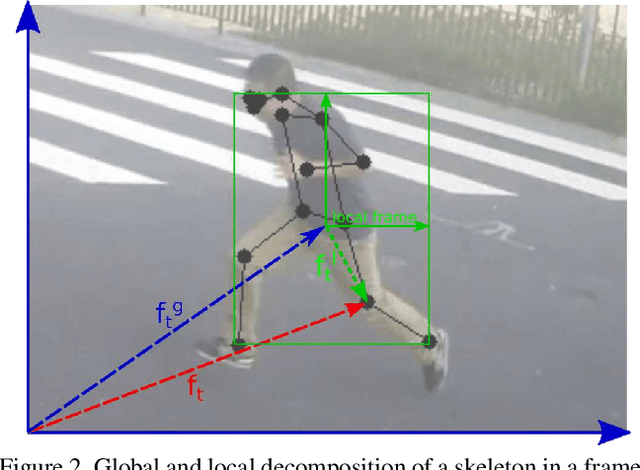
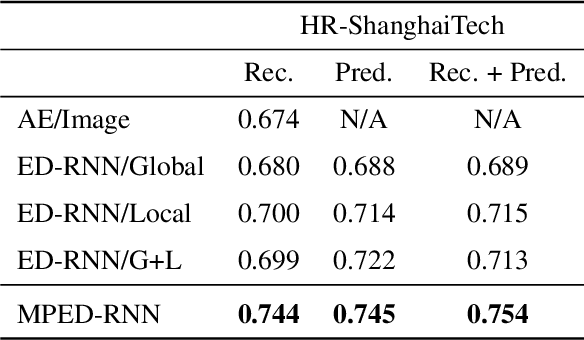
Abstract:Appearance features have been widely used in video anomaly detection even though they contain complex entangled factors. We propose a new method to model the normal patterns of human movements in surveillance video for anomaly detection using dynamic skeleton features. We decompose the skeletal movements into two sub-components: global body movement and local body posture. We model the dynamics and interaction of the coupled features in our novel Message-Passing Encoder-Decoder Recurrent Network. We observed that the decoupled features collaboratively interact in our spatio-temporal model to accurately identify human-related irregular events from surveillance video sequences. Compared to traditional appearance-based models, our method achieves superior outlier detection performance. Our model also offers "open-box" examination and decision explanation made possible by the semantically understandable features and a network architecture supporting interpretability.
Improving Generalization and Stability of Generative Adversarial Networks
Feb 11, 2019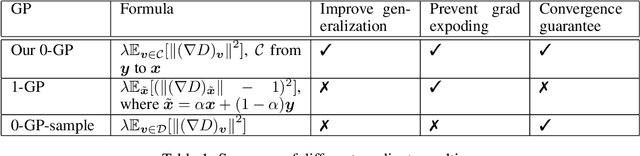
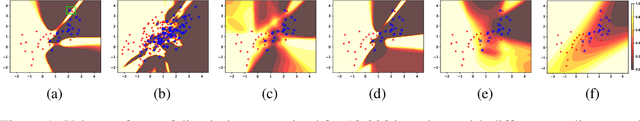
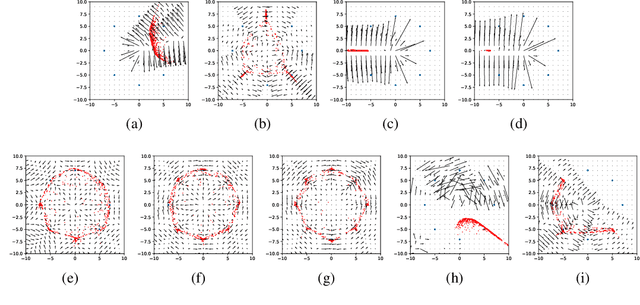

Abstract:Generative Adversarial Networks (GANs) are one of the most popular tools for learning complex high dimensional distributions. However, generalization properties of GANs have not been well understood. In this paper, we analyze the generalization of GANs in practical settings. We show that discriminators trained on discrete datasets with the original GAN loss have poor generalization capability and do not approximate the theoretically optimal discriminator. We propose a zero-centered gradient penalty for improving the generalization of the discriminator by pushing it toward the optimal discriminator. The penalty guarantees the generalization and convergence of GANs. Experiments on synthetic and large scale datasets verify our theoretical analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge