Thiru Siddharth
Predicting pathways for old and new metabolites through clustering
Nov 28, 2022Abstract:The diverse metabolic pathways are fundamental to all living organisms, as they harvest energy, synthesize biomass components, produce molecules to interact with the microenvironment, and neutralize toxins. While discovery of new metabolites and pathways continues, the prediction of pathways for new metabolites can be challenging. It can take vast amounts of time to elucidate pathways for new metabolites; thus, according to HMDB only 60% of metabolites get assigned to pathways. Here, we present an approach to identify pathways based on metabolite structure. We extracted 201 features from SMILES annotations, and identified new metabolites from PubMed abstracts and HMDB. After applying clustering algorithms to both groups of features, we quantified correlations between metabolites, and found the clusters accurately linked 92% of known metabolites to their respective pathways. Thus, this approach could be valuable for predicting metabolic pathways for new metabolites.
Plant Species Classification Using Transfer Learning by Pretrained Classifier VGG-19
Sep 07, 2022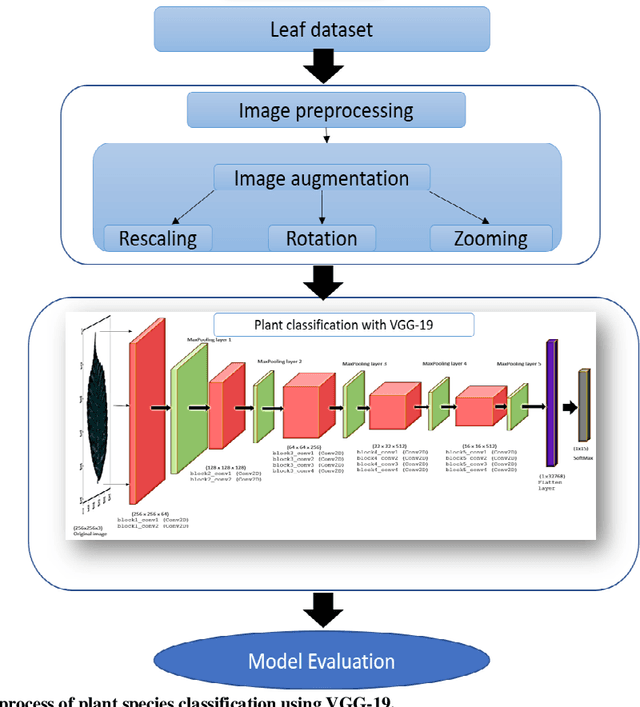
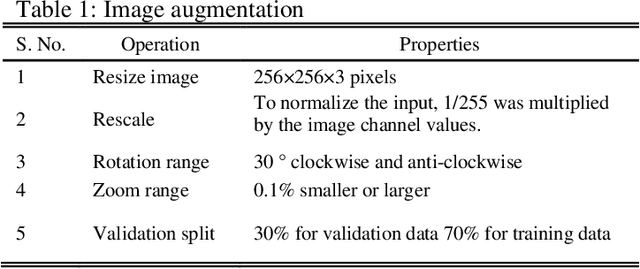
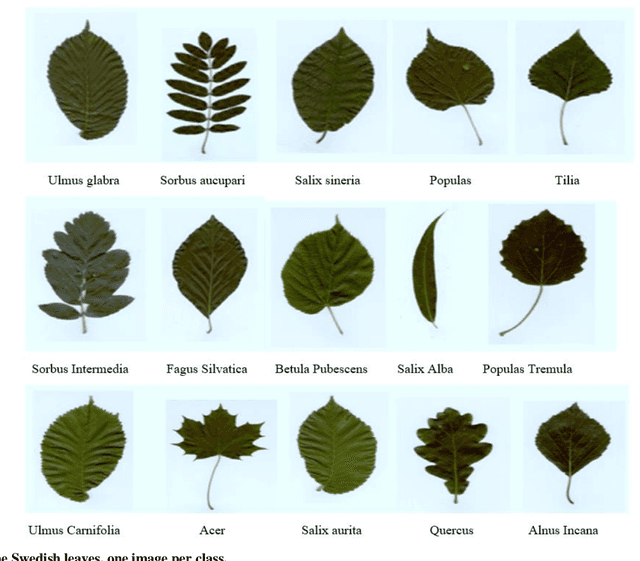


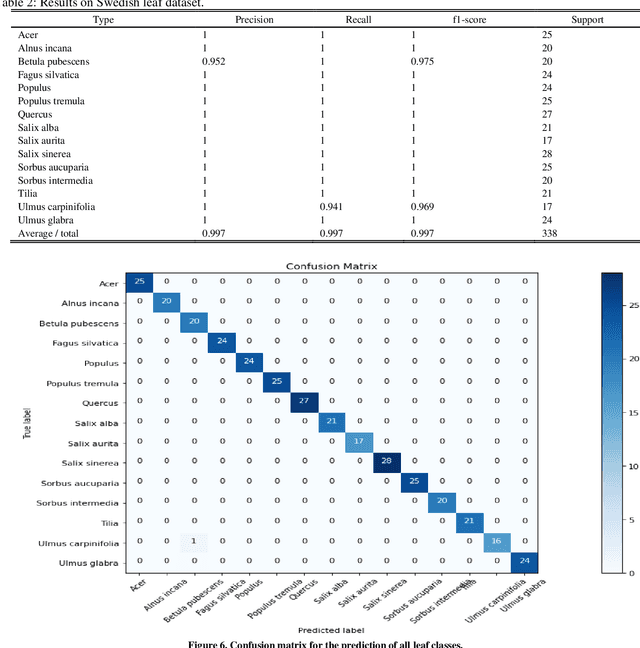
Abstract:Deep learning is currently the most important branch of machine learning, with applications in speech recognition, computer vision, image classification, and medical imaging analysis. Plant recognition is one of the areas where image classification can be used to identify plant species through their leaves. Botanists devote a significant amount of time to recognizing plant species by personally inspecting. This paper describes a method for dissecting color images of Swedish leaves and identifying plant species. To achieve higher accuracy, the task is completed using transfer learning with the help of pre-trained classifier VGG-19. The four primary processes of classification are image preprocessing, image augmentation, feature extraction, and recognition, which are performed as part of the overall model evaluation. The VGG-19 classifier grasps the characteristics of leaves by employing pre-defined hidden layers such as convolutional layers, max pooling layers, and fully connected layers, and finally uses the soft-max layer to generate a feature representation for all plant classes. The model obtains knowledge connected to aspects of the Swedish leaf dataset, which contains fifteen tree classes, and aids in predicting the proper class of an unknown plant with an accuracy of 99.70% which is higher than previous research works reported.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge