Thierry Delzescaux
Adversarial Stain Transfer to Study the Effect of Color Variation on Cell Instance Segmentation
Sep 01, 2022
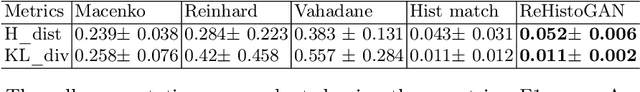
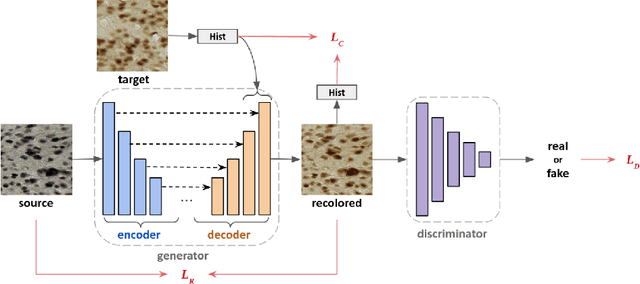
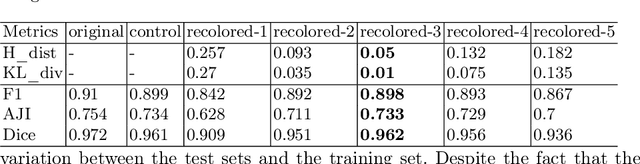
Abstract:Stain color variation in histological images, caused by a variety of factors, is a challenge not only for the visual diagnosis of pathologists but also for cell segmentation algorithms. To eliminate the color variation, many stain normalization approaches have been proposed. However, most were designed for hematoxylin and eosin staining images and performed poorly on immunohistochemical staining images. Current cell segmentation methods systematically apply stain normalization as a preprocessing step, but the impact brought by color variation has not been quantitatively investigated yet. In this paper, we produced five groups of NeuN staining images with different colors. We applied a deep learning image-recoloring method to perform color transfer between histological image groups. Finally, we altered the color of a segmentation set and quantified the impact of color variation on cell segmentation. The results demonstrated the necessity of color normalization prior to subsequent analysis.
End-to-end Neuron Instance Segmentation based on Weakly Supervised Efficient UNet and Morphological Post-processing
Feb 17, 2022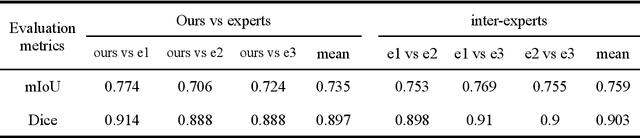
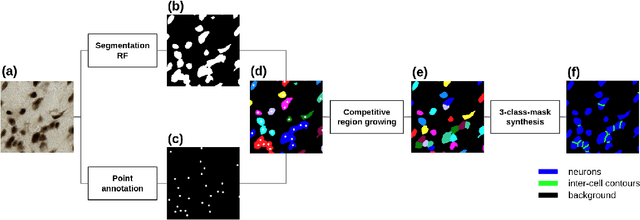
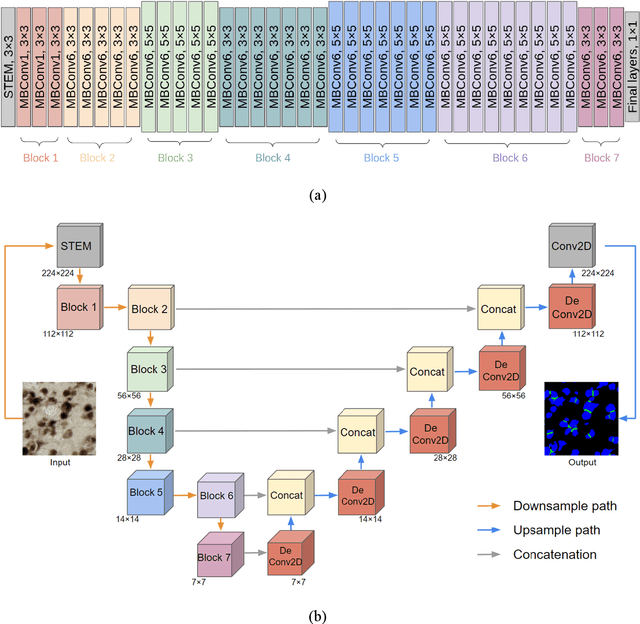
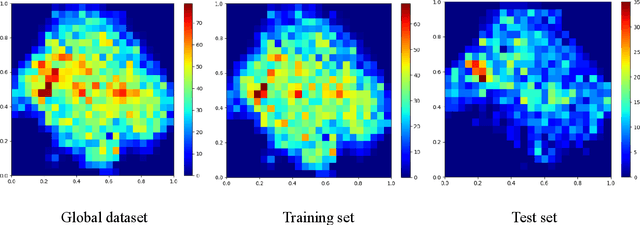
Abstract:Recent studies have demonstrated the superiority of deep learning in medical image analysis, especially in cell instance segmentation, a fundamental step for many biological studies. However, the good performance of the neural networks requires training on large unbiased dataset and annotations, which is labor-intensive and expertise-demanding. In this paper, we present an end-to-end weakly-supervised framework to automatically detect and segment NeuN stained neuronal cells on histological images using only point annotations. We integrate the state-of-the-art network, EfficientNet, into our U-Net-like architecture. Validation results show the superiority of our model compared to other recent methods. In addition, we investigated multiple post-processing schemes and proposed an original strategy to convert the probability map into segmented instances using ultimate erosion and dynamic reconstruction. This approach is easy to configure and outperforms other classical post-processing techniques.
Automated Atlas-based Segmentation of Single Coronal Mouse Brain Slices using Linear 2D-2D Registration
Nov 16, 2021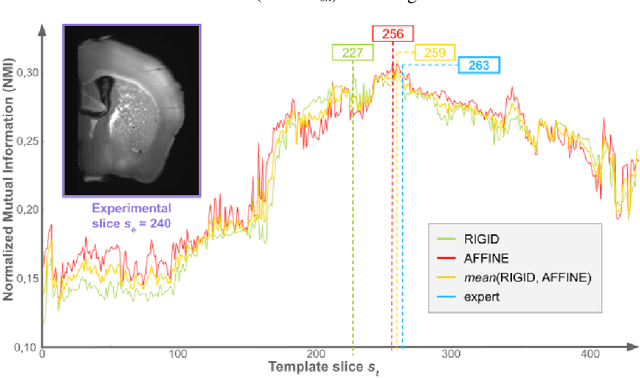
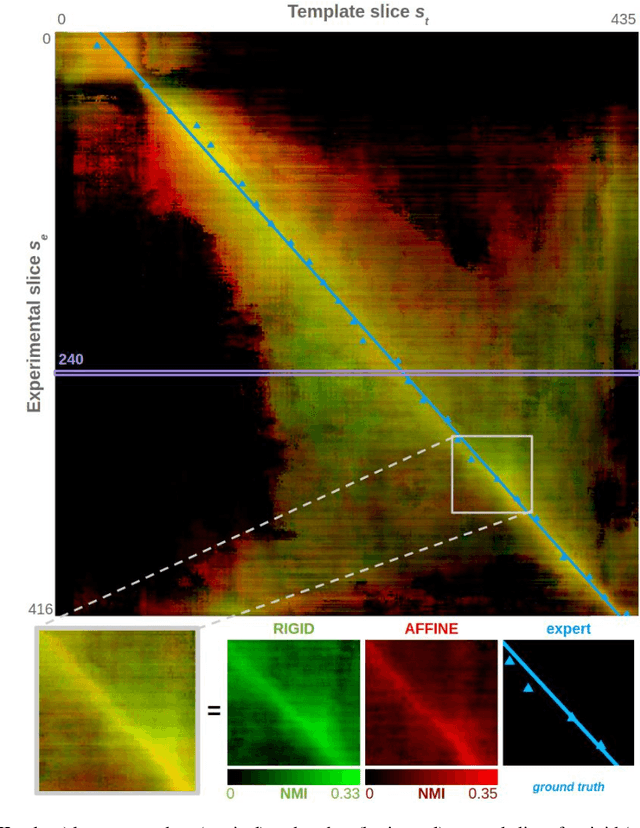

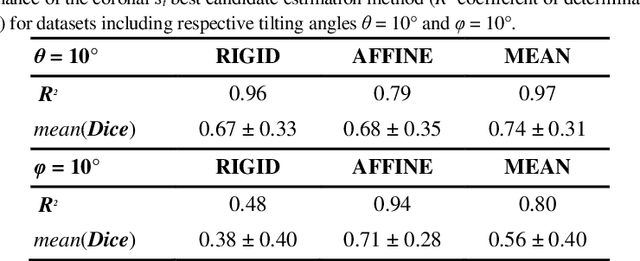
Abstract:A significant challenge for brain histological data analysis is to precisely identify anatomical regions in order to perform accurate local quantifications and evaluate therapeutic solutions. Usually, this task is performed manually, becoming therefore tedious and subjective. Another option is to use automatic or semi-automatic methods, among which segmentation using digital atlases co-registration. However, most available atlases are 3D, whereas digitized histological data are 2D. Methods to perform such 2D-3D segmentation from an atlas are required. This paper proposes a strategy to automatically and accurately segment single 2D coronal slices within a 3D volume of atlas, using linear registration. We validated its robustness and performance using an exploratory approach at whole-brain scale.
Evaluation of Deep Learning Topcoders Method for Neuron Individualization in Histological Macaque Brain Section
Nov 10, 2021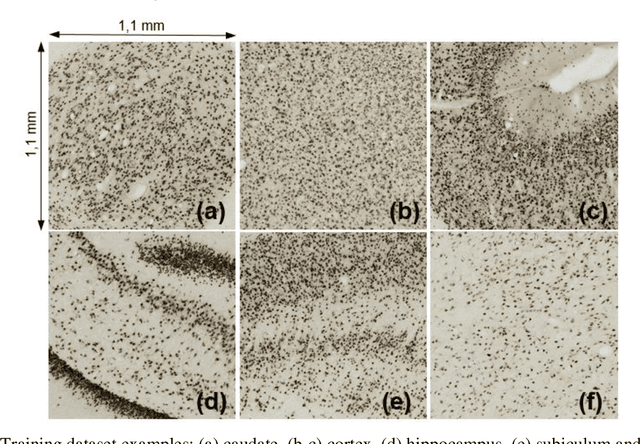
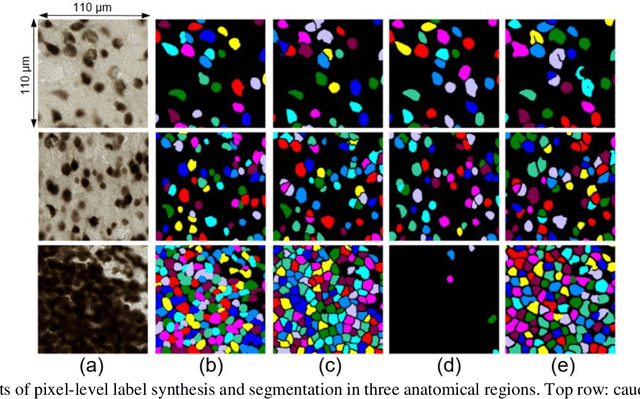
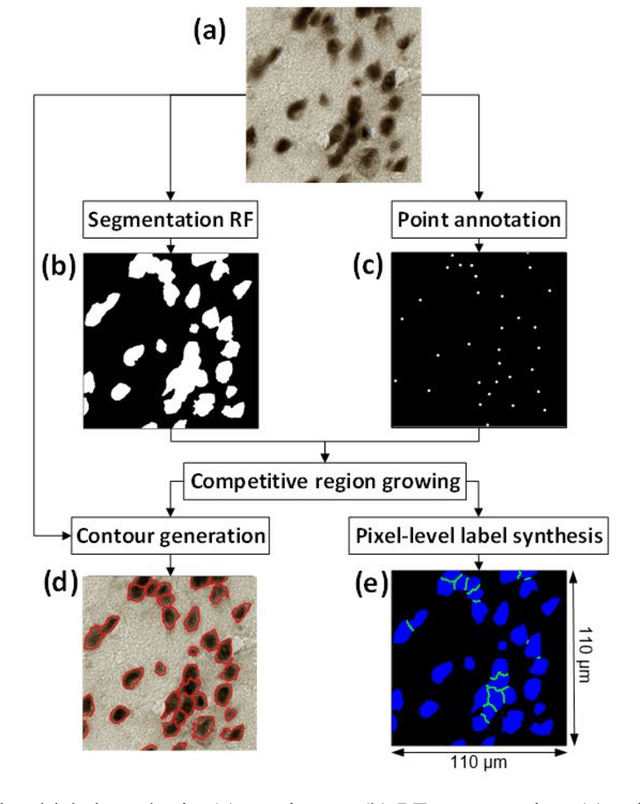

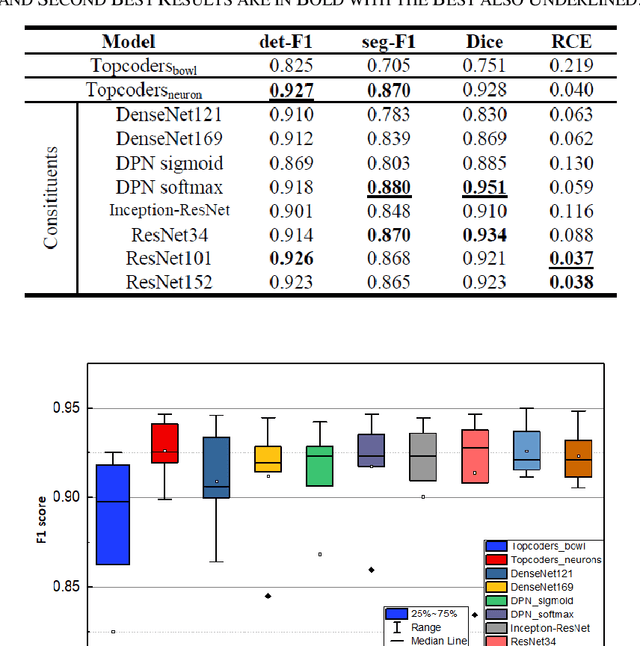
Abstract:Cell individualization has a vital role in digital pathology image analysis. Deep Learning is considered as an efficient tool for instance segmentation tasks, including cell individualization. However, the precision of the Deep Learning model relies on massive unbiased dataset and manual pixel-level annotations, which is labor intensive. Moreover, most applications of Deep Learning have been developed for processing oncological data. To overcome these challenges, i) we established a pipeline to synthesize pixel-level labels with only point annotations provided; ii) we tested an ensemble Deep Learning algorithm to perform cell individualization on neurological data. Results suggest that the proposed method successfully segments neuronal cells in both object-level and pixel-level, with an average detection accuracy of 0.93.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge