Talley Lambert
λSplit: Self-Supervised Content-Aware Spectral Unmixing for Fluorescence Microscopy
Mar 24, 2026Abstract:In fluorescence microscopy, spectral unmixing aims to recover individual fluorophore concentrations from spectral images that capture mixed fluorophore emissions. Since classical methods operate pixel-wise and rely on least-squares fitting, their performance degrades with increasingly overlapping emission spectra and higher levels of noise, suggesting that a data-driven approach that can learn and utilize a structural prior might lead to improved results. Learning-based approaches for spectral imaging do exist, but they are either not optimized for microscopy data or are developed for very specific cases that are not applicable to fluorescence microscopy settings. To address this, we propose λSplit, a physics-informed deep generative model that learns a conditional distribution over concentration maps using a hierarchical Variational Autoencoder. A fully differentiable Spectral Mixer enforces consistency with the image formation process, while the learned structural priors enable state-of-the-art unmixing and implicit noise removal. We demonstrate λSplit on 3 real-world datasets that we synthetically cast into a total of 66 challenging spectral unmixing benchmarks. We compare our results against a total of 10 baseline methods, including classical methods and a range of learning-based methods. Our results consistently show competitive performance and improved robustness in high noise regimes, when spectra overlap considerably, or when the spectral dimensionality is lowered, making λSplit a new state-of-the-art for spectral unmixing of fluorescent microscopy data. Importantly, λSplit is compatible with spectral data produced by standard confocal microscopes, enabling immediate adoption without specialized hardware modifications.
DeadNet: Identifying Phototoxicity from Label-free Microscopy Images of Cells using Deep ConvNets
Jan 22, 2017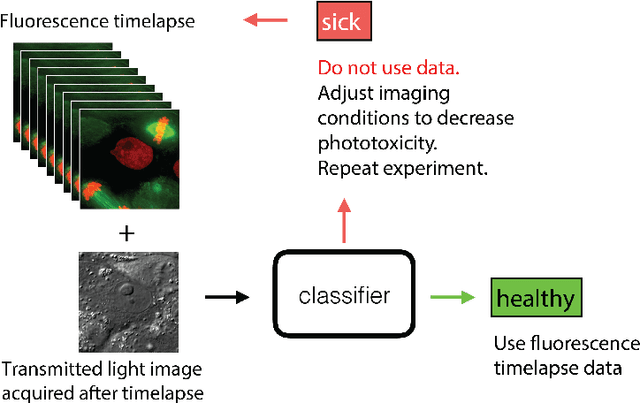
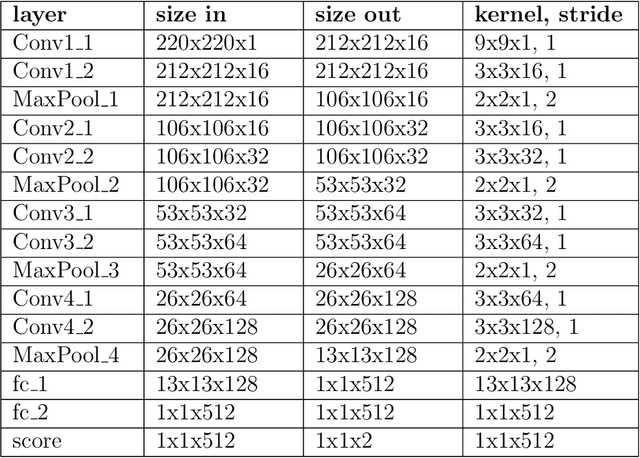
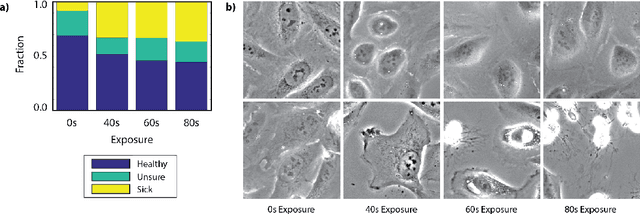
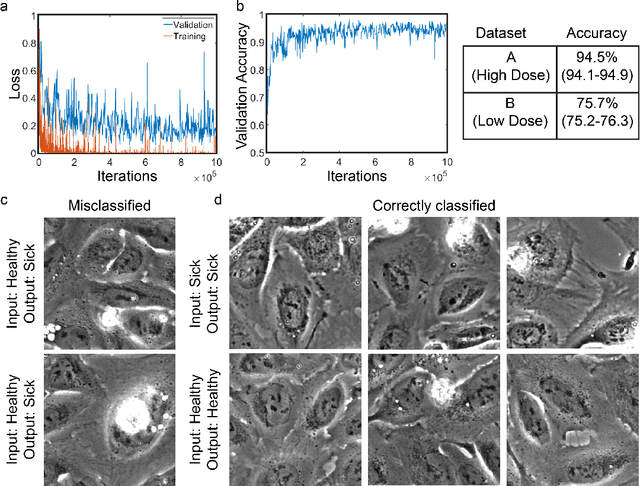
Abstract:Exposure to intense illumination light is an unavoidable consequence of fluorescence microscopy, and poses a risk to the health of the sample in every live-cell fluorescence microscopy experiment. Furthermore, the possible side-effects of phototoxicity on the scientific conclusions that are drawn from an imaging experiment are often unaccounted for. Previously, controlling for phototoxicity in imaging experiments required additional labels and experiments, limiting its widespread application. Here we provide a proof-of-principle demonstration that the phototoxic effects of an imaging experiment can be identified directly from a single phase-contrast image using deep convolutional neural networks (ConvNets). This lays the groundwork for an automated tool for assessing cell health in a wide range of imaging experiments. Interpretability of such a method is crucial for its adoption. We take steps towards interpreting the classification mechanism of the trained ConvNet by visualizing salient features of images that contribute to accurate classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge