Shu-Min Kao
Toward Drug-Target Interaction Prediction via Ensemble Modeling and Transfer Learning
Jul 28, 2021
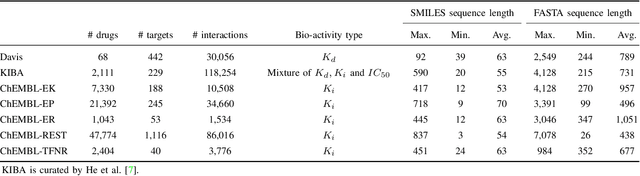
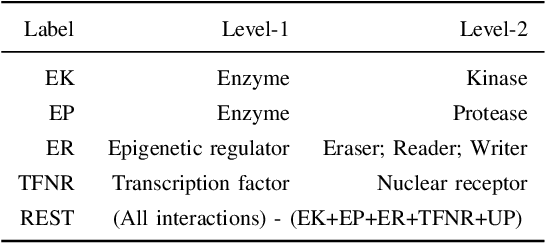

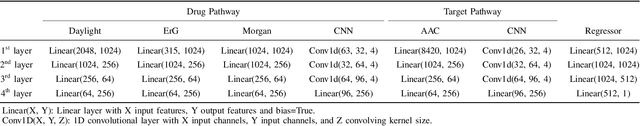
Abstract:Drug-target interaction (DTI) prediction plays a crucial role in drug discovery, and deep learning approaches have achieved state-of-the-art performance in this field. We introduce an ensemble of deep learning models (EnsembleDLM) for DTI prediction. EnsembleDLM only uses the sequence information of chemical compounds and proteins, and it aggregates the predictions from multiple deep neural networks. This approach not only achieves state-of-the-art performance in Davis and KIBA datasets but also reaches cutting-edge performance in the cross-domain applications across different bio-activity types and different protein classes. We also demonstrate that EnsembleDLM achieves a good performance (Pearson correlation coefficient and concordance index > 0.8) in the new domain with approximately 50% transfer learning data, i.e., the training set has twice as much data as the test set.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge