Santu Rana
Plug and Play, Model-Based Reinforcement Learning
Aug 20, 2021
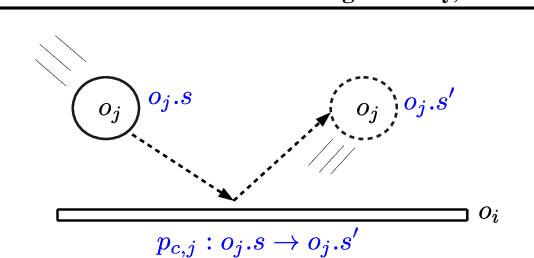
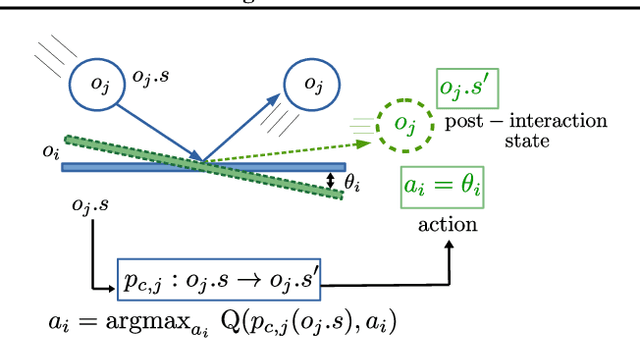
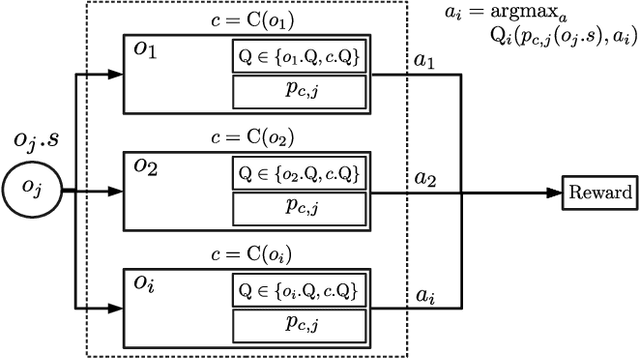
Abstract:Sample-efficient generalisation of reinforcement learning approaches have always been a challenge, especially, for complex scenes with many components. In this work, we introduce Plug and Play Markov Decision Processes, an object-based representation that allows zero-shot integration of new objects from known object classes. This is achieved by representing the global transition dynamics as a union of local transition functions, each with respect to one active object in the scene. Transition dynamics from an object class can be pre-learnt and thus would be ready to use in a new environment. Each active object is also endowed with its reward function. Since there is no central reward function, addition or removal of objects can be handled efficiently by only updating the reward functions of objects involved. A new transfer learning mechanism is also proposed to adapt reward function in such cases. Experiments show that our representation can achieve sample-efficiency in a variety of set-ups.
Combining Online Learning and Offline Learning for Contextual Bandits with Deficient Support
Jul 24, 2021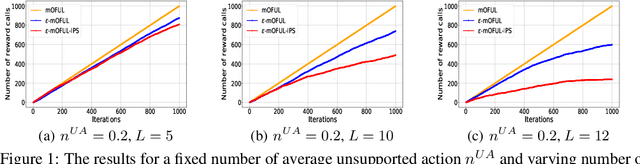
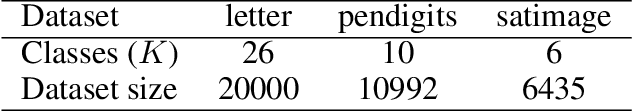

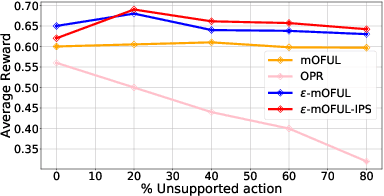
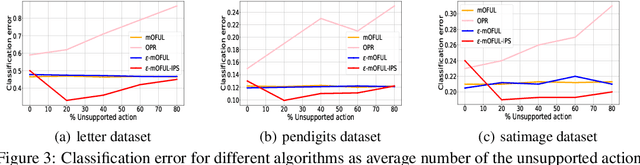
Abstract:We address policy learning with logged data in contextual bandits. Current offline-policy learning algorithms are mostly based on inverse propensity score (IPS) weighting requiring the logging policy to have \emph{full support} i.e. a non-zero probability for any context/action of the evaluation policy. However, many real-world systems do not guarantee such logging policies, especially when the action space is large and many actions have poor or missing rewards. With such \emph{support deficiency}, the offline learning fails to find optimal policies. We propose a novel approach that uses a hybrid of offline learning with online exploration. The online exploration is used to explore unsupported actions in the logged data whilst offline learning is used to exploit supported actions from the logged data avoiding unnecessary explorations. Our approach determines an optimal policy with theoretical guarantees using the minimal number of online explorations. We demonstrate our algorithms' effectiveness empirically on a diverse collection of datasets.
A New Representation of Successor Features for Transfer across Dissimilar Environments
Jul 18, 2021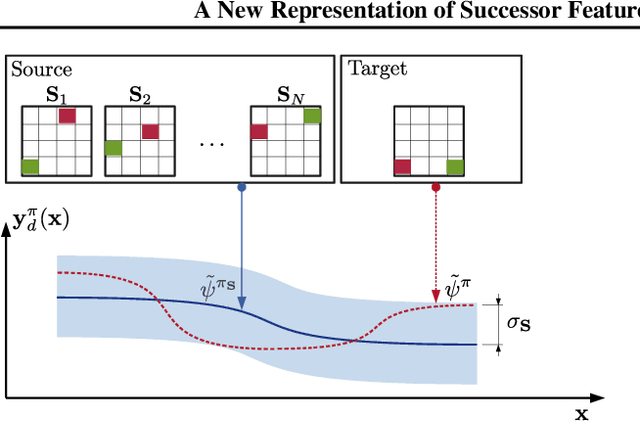
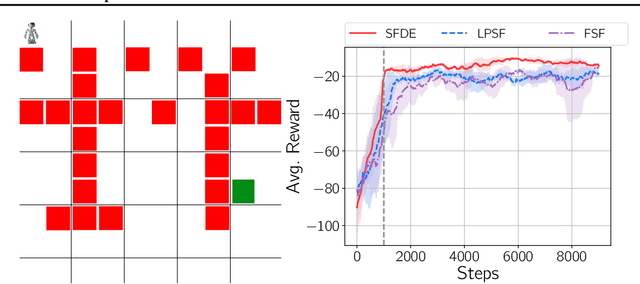
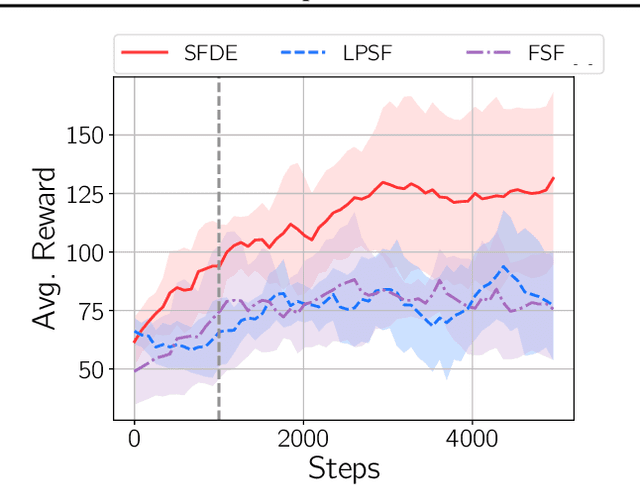
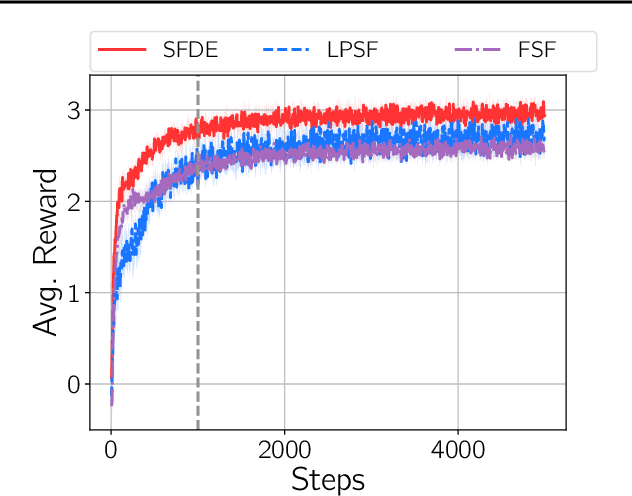
Abstract:Transfer in reinforcement learning is usually achieved through generalisation across tasks. Whilst many studies have investigated transferring knowledge when the reward function changes, they have assumed that the dynamics of the environments remain consistent. Many real-world RL problems require transfer among environments with different dynamics. To address this problem, we propose an approach based on successor features in which we model successor feature functions with Gaussian Processes permitting the source successor features to be treated as noisy measurements of the target successor feature function. Our theoretical analysis proves the convergence of this approach as well as the bounded error on modelling successor feature functions with Gaussian Processes in environments with both different dynamics and rewards. We demonstrate our method on benchmark datasets and show that it outperforms current baselines.
Bayesian Optimistic Optimisation with Exponentially Decaying Regret
May 10, 2021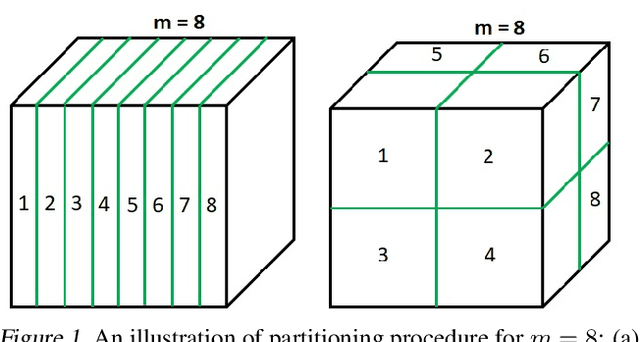
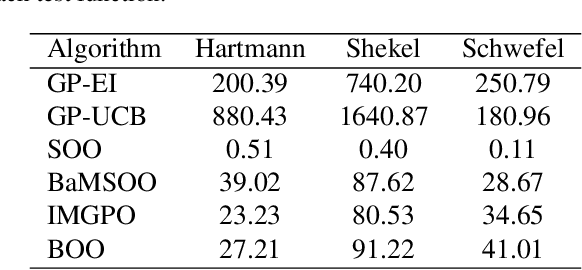
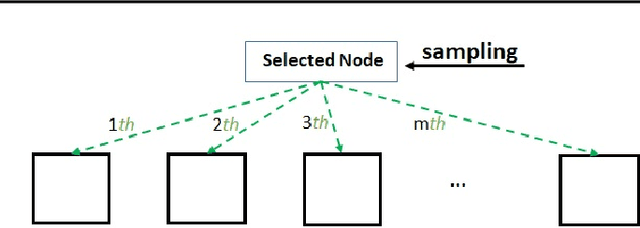
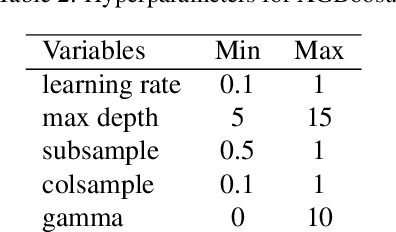
Abstract:Bayesian optimisation (BO) is a well-known efficient algorithm for finding the global optimum of expensive, black-box functions. The current practical BO algorithms have regret bounds ranging from $\mathcal{O}(\frac{logN}{\sqrt{N}})$ to $\mathcal O(e^{-\sqrt{N}})$, where $N$ is the number of evaluations. This paper explores the possibility of improving the regret bound in the noiseless setting by intertwining concepts from BO and tree-based optimistic optimisation which are based on partitioning the search space. We propose the BOO algorithm, a first practical approach which can achieve an exponential regret bound with order $\mathcal O(N^{-\sqrt{N}})$ under the assumption that the objective function is sampled from a Gaussian process with a Mat\'ern kernel with smoothness parameter $\nu > 4 +\frac{D}{2}$, where $D$ is the number of dimensions. We perform experiments on optimisation of various synthetic functions and machine learning hyperparameter tuning tasks and show that our algorithm outperforms baselines.
Intuitive Physics Guided Exploration for Sample Efficient Sim2real Transfer
Apr 18, 2021
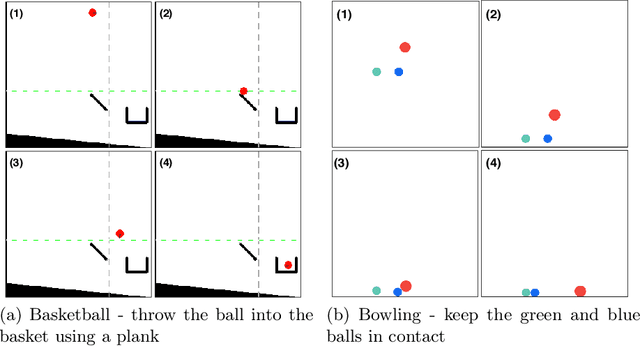
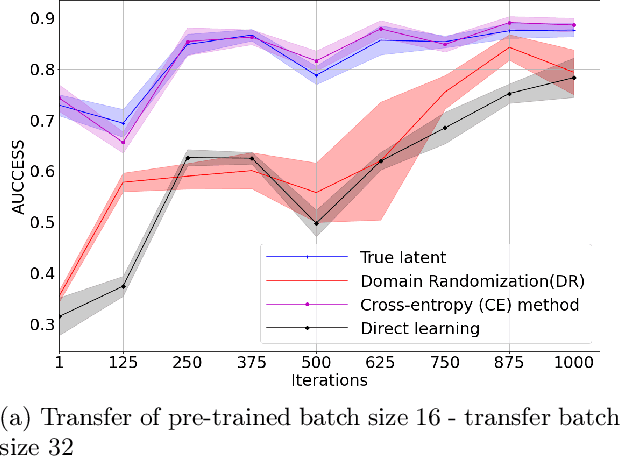
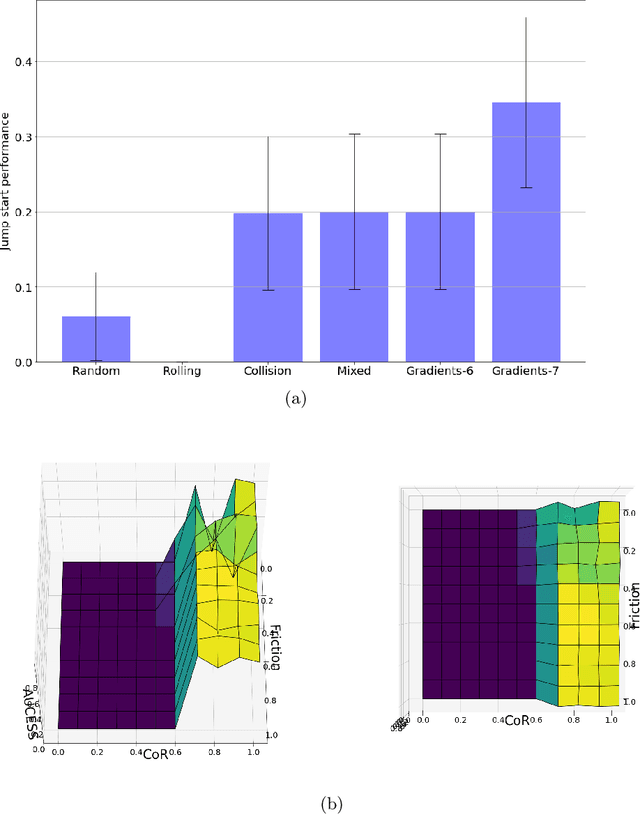
Abstract:Physics-based reinforcement learning tasks can benefit from simplified physics simulators as they potentially allow near-optimal policies to be learned in simulation. However, such simulators require the latent factors (e.g. mass, friction coefficient etc.) of the associated objects and other environment-specific factors (e.g. wind speed, air density etc.) to be accurately specified, without which, it could take considerable additional learning effort to adapt the learned simulation policy to the real environment. As such a complete specification can be impractical, in this paper, we instead, focus on learning task-specific estimates of latent factors which allow the approximation of real world trajectories in an ideal simulation environment. Specifically, we propose two new concepts: a) action grouping - the idea that certain types of actions are closely associated with the estimation of certain latent factors, and; b) partial grounding - the idea that simulation of task-specific dynamics may not need precise estimation of all the latent factors. We first introduce intuitive action groupings based on human physics knowledge and experience, which is then used to design novel strategies for interacting with the real environment. Next, we describe how prior knowledge of a task in a given environment can be used to extract the relative importance of different latent factors, and how this can be used to inform partial grounding, which enables efficient learning of the task in any arbitrary environment. We demonstrate our approach in a range of physics based tasks, and show that it achieves superior performance relative to other baselines, using only a limited number of real-world interactions.
ALT-MAS: A Data-Efficient Framework for Active Testing of Machine Learning Algorithms
Apr 11, 2021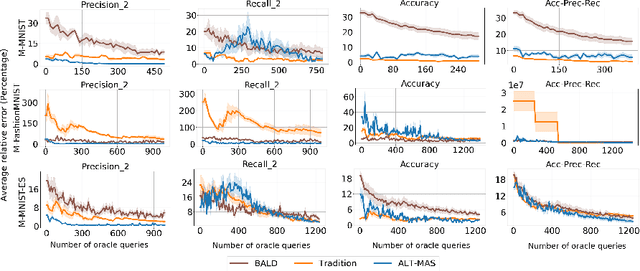
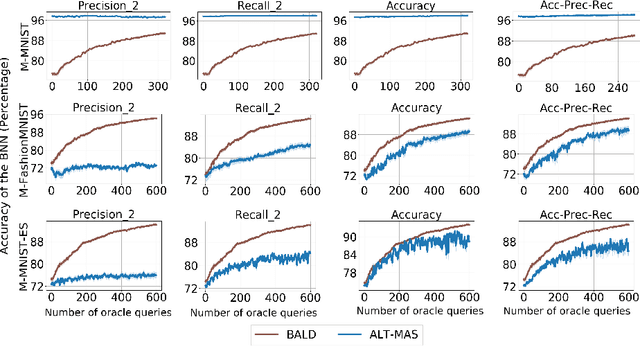
Abstract:Machine learning models are being used extensively in many important areas, but there is no guarantee a model will always perform well or as its developers intended. Understanding the correctness of a model is crucial to prevent potential failures that may have significant detrimental impact in critical application areas. In this paper, we propose a novel framework to efficiently test a machine learning model using only a small amount of labeled test data. The idea is to estimate the metrics of interest for a model-under-test using Bayesian neural network (BNN). We develop a novel data augmentation method helping to train the BNN to achieve high accuracy. We also devise a theoretic information based sampling strategy to sample data points so as to achieve accurate estimations for the metrics of interest. Finally, we conduct an extensive set of experiments to test various machine learning models for different types of metrics. Our experiments show that the metrics estimations by our method are significantly better than existing baselines.
High Dimensional Level Set Estimation with Bayesian Neural Network
Dec 17, 2020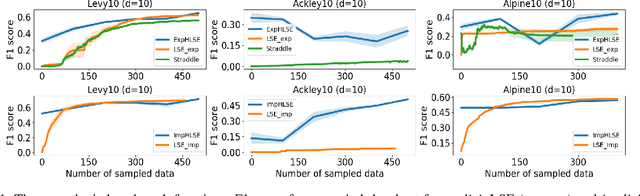
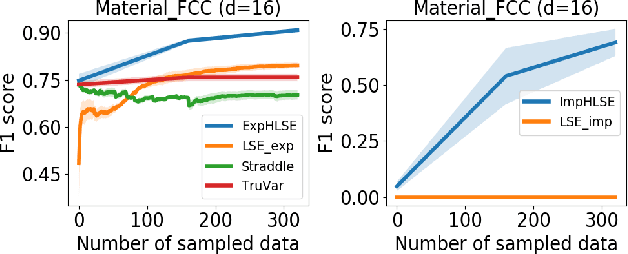
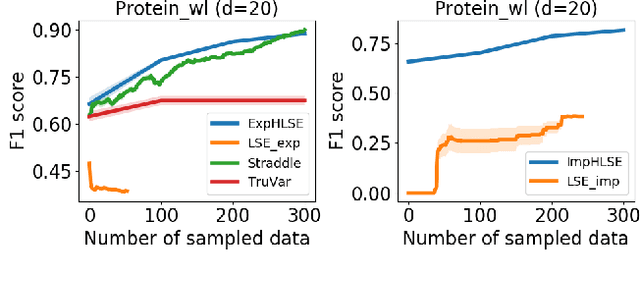
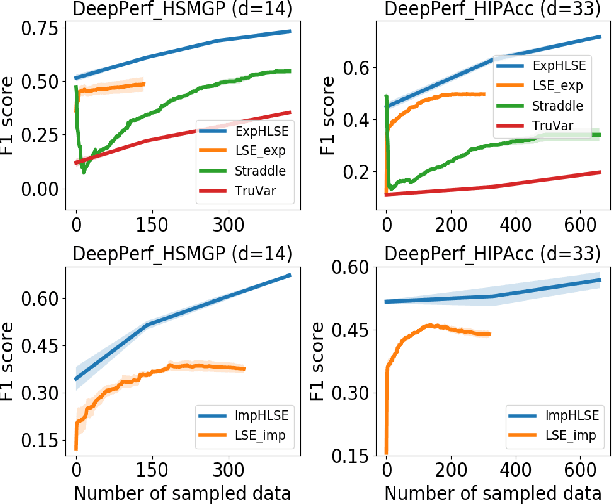
Abstract:Level Set Estimation (LSE) is an important problem with applications in various fields such as material design, biotechnology, machine operational testing, etc. Existing techniques suffer from the scalability issue, that is, these methods do not work well with high dimensional inputs. This paper proposes novel methods to solve the high dimensional LSE problems using Bayesian Neural Networks. In particular, we consider two types of LSE problems: (1) \textit{explicit} LSE problem where the threshold level is a fixed user-specified value, and, (2) \textit{implicit} LSE problem where the threshold level is defined as a percentage of the (unknown) maximum of the objective function. For each problem, we derive the corresponding theoretic information based acquisition function to sample the data points so as to maximally increase the level set accuracy. Furthermore, we also analyse the theoretical time complexity of our proposed acquisition functions, and suggest a practical methodology to efficiently tune the network hyper-parameters to achieve high model accuracy. Numerical experiments on both synthetic and real-world datasets show that our proposed method can achieve better results compared to existing state-of-the-art approaches.
Logically Consistent Loss for Visual Question Answering
Nov 19, 2020
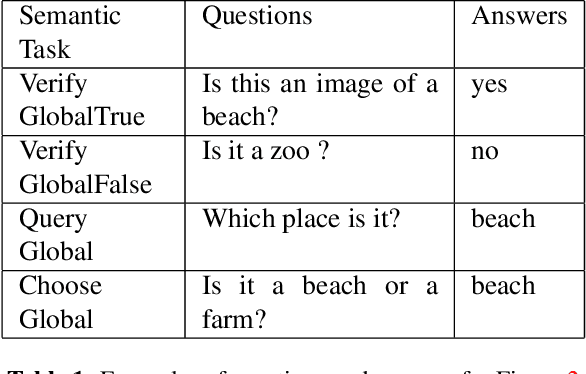
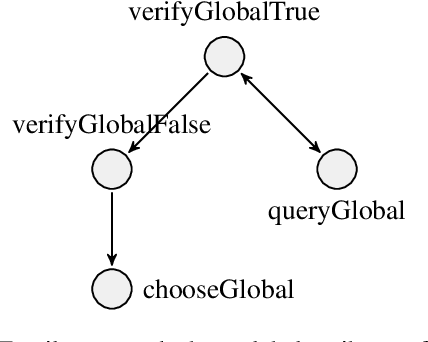
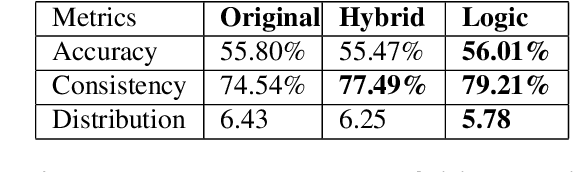
Abstract:Given an image, a back-ground knowledge, and a set of questions about an object, human learners answer the questions very consistently regardless of question forms and semantic tasks. The current advancement in neural-network based Visual Question Answering (VQA), despite their impressive performance, cannot ensure such consistency due to identically distribution (i.i.d.) assumption. We propose a new model-agnostic logic constraint to tackle this issue by formulating a logically consistent loss in the multi-task learning framework as well as a data organisation called family-batch and hybrid-batch. To demonstrate usefulness of this proposal, we train and evaluate MAC-net based VQA machines with and without the proposed logically consistent loss and the proposed data organization. The experiments confirm that the proposed loss formulae and introduction of hybrid-batch leads to more consistency as well as better performance. Though the proposed approach is tested with MAC-net, it can be utilised in any other QA methods whenever the logical consistency between answers exist.
Unsupervised Anomaly Detection on Temporal Multiway Data
Sep 20, 2020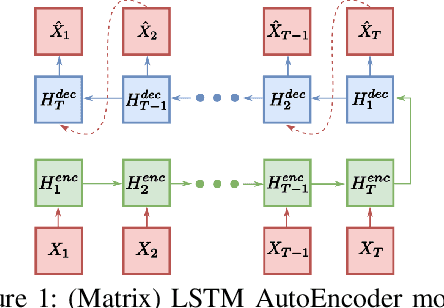
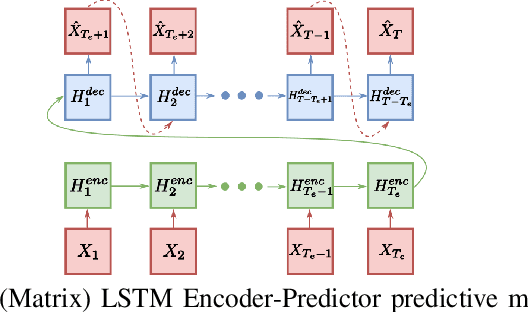
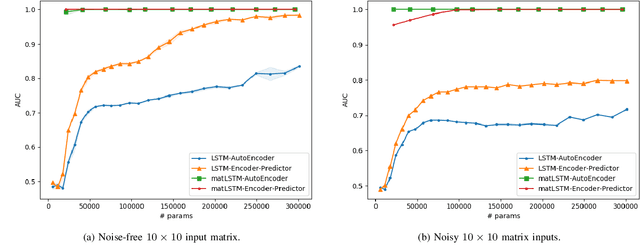
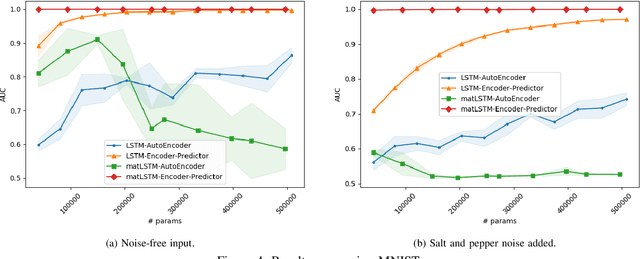
Abstract:Temporal anomaly detection looks for irregularities over space-time. Unsupervised temporal models employed thus far typically work on sequences of feature vectors, and much less on temporal multiway data. We focus our investigation on two-way data, in which a data matrix is observed at each time step. Leveraging recent advances in matrix-native recurrent neural networks, we investigated strategies for data arrangement and unsupervised training for temporal multiway anomaly detection. These include compressing-decompressing, encoding-predicting, and temporal data differencing. We conducted a comprehensive suite of experiments to evaluate model behaviors under various settings on synthetic data, moving digits, and ECG recordings. We found interesting phenomena not previously reported. These include the capacity of the compact matrix LSTM to compress noisy data near perfectly, making the strategy of compressing-decompressing data ill-suited for anomaly detection under the noise. Also, long sequence of vectors can be addressed directly by matrix models that allow very long context and multiple step prediction. Overall, the encoding-predicting strategy works very well for the matrix LSTMs in the conducted experiments, thanks to its compactness and better fit to the data dynamics.
Sub-linear Regret Bounds for Bayesian Optimisation in Unknown Search Spaces
Sep 09, 2020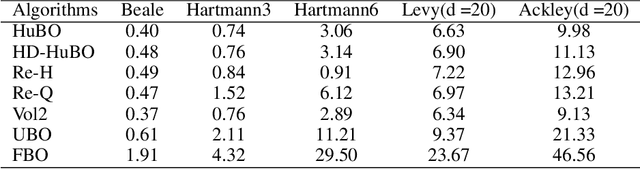
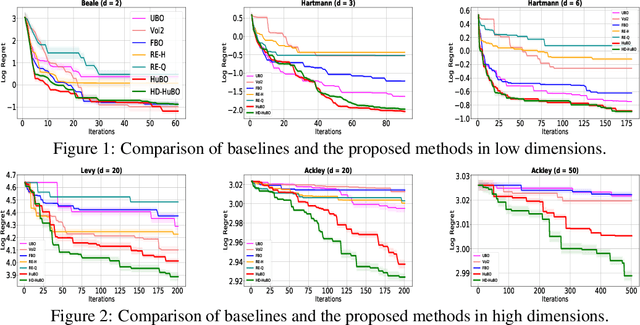
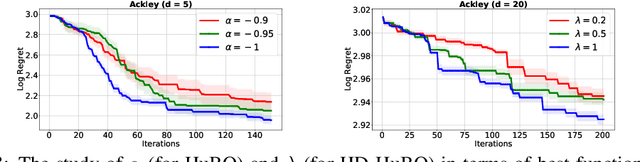
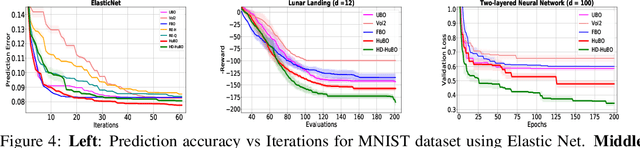
Abstract:Bayesian optimisation is a popular method for efficient optimisation of expensive black-box functions. Traditionally, BO assumes that the search space is known. However, in many problems, this assumption does not hold. To this end, we propose a novel BO algorithm which expands (and shifts) the search space over iterations based on controlling the expansion rate thought a hyperharmonic series. Further, we propose another variant of our algorithm that scales to high dimensions. We show theoretically that for both our algorithms, the cumulative regret grows at sub-linear rates. Our experiments with synthetic and real-world optimisation tasks demonstrate the superiority of our algorithms over the current state-of-the-art methods for Bayesian optimisation in unknown search space.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge