Ruichen Rong
Deep reflective reasoning in interdependence constrained structured data extraction from clinical notes for digital health
Mar 20, 2026Abstract:Extracting structured information from clinical notes requires navigating a dense web of interdependent variables where the value of one attribute logically constrains others. Existing Large Language Model (LLM)-based extraction pipelines often struggle to capture these dependencies, leading to clinically inconsistent outputs. We propose deep reflective reasoning, a large language model agent framework that iteratively self-critiques and revises structured outputs by checking consistency among variables, the input text, and retrieved domain knowledge, stopping when outputs converge. We extensively evaluate the proposed method in three diverse oncology applications: (1) On colorectal cancer synoptic reporting from gross descriptions (n=217), reflective reasoning improved average F1 across eight categorical synoptic variables from 0.828 to 0.911 and increased mean correct rate across four numeric variables from 0.806 to 0.895; (2) On Ewing sarcoma CD99 immunostaining pattern identification (n=200), the accuracy improved from 0.870 to 0.927; (3) On lung cancer tumor staging (n=100), tumor stage accuracy improved from 0.680 to 0.833 (pT: 0.842 -> 0.884; pN: 0.885 -> 0.948). The results demonstrate that deep reflective reasoning can systematically improve the reliability of LLM-based structured data extraction under interdependence constraints, enabling more consistent machine-operable clinical datasets and facilitating knowledge discovery with machine learning and data science towards digital health.
Large language models enabled multiagent ensemble method for efficient EHR data labeling
Oct 21, 2024



Abstract:This study introduces a novel multiagent ensemble method powered by LLMs to address a key challenge in ML - data labeling, particularly in large-scale EHR datasets. Manual labeling of such datasets requires domain expertise and is labor-intensive, time-consuming, expensive, and error-prone. To overcome this bottleneck, we developed an ensemble LLMs method and demonstrated its effectiveness in two real-world tasks: (1) labeling a large-scale unlabeled ECG dataset in MIMIC-IV; (2) identifying social determinants of health (SDOH) from the clinical notes of EHR. Trading off benefits and cost, we selected a pool of diverse open source LLMs with satisfactory performance. We treat each LLM's prediction as a vote and apply a mechanism of majority voting with minimal winning threshold for ensemble. We implemented an ensemble LLMs application for EHR data labeling tasks. By using the ensemble LLMs and natural language processing, we labeled MIMIC-IV ECG dataset of 623,566 ECG reports with an estimated accuracy of 98.2%. We applied the ensemble LLMs method to identify SDOH from social history sections of 1,405 EHR clinical notes, also achieving competitive performance. Our experiments show that the ensemble LLMs can outperform individual LLM even the best commercial one, and the method reduces hallucination errors. From the research, we found that (1) the ensemble LLMs method significantly reduces the time and effort required for labeling large-scale EHR data, automating the process with high accuracy and quality; (2) the method generalizes well to other text data labeling tasks, as shown by its application to SDOH identification; (3) the ensemble of a group of diverse LLMs can outperform or match the performance of the best individual LLM; and (4) the ensemble method substantially reduces hallucination errors. This approach provides a scalable and efficient solution to data-labeling challenges.
What evidence does deep learning model use to classify Skin Lesions?
Nov 02, 2018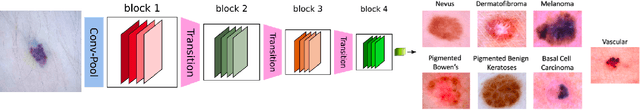
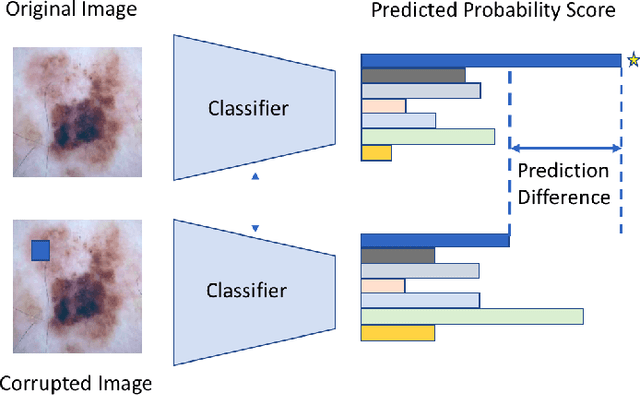
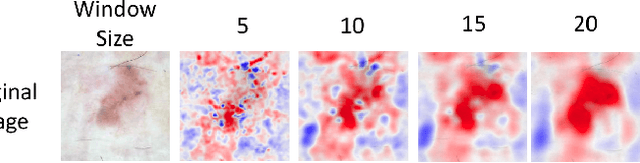
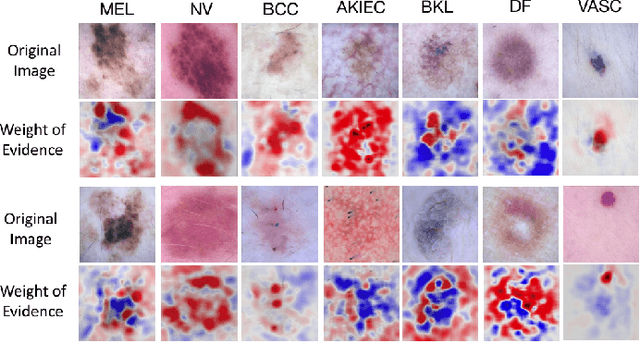
Abstract:Melanoma is a type of skin cancer with the most rapidly increasing incidence. Early detection of melanoma using dermoscopy images significantly increases patients' survival rate. However, accurately classifying skin lesions, especially in the early stage, is extremely challenging via dermatologists' observation. Hence, the discovery of reliable biomarkers for melanoma diagnosis will be meaningful. Recent years, deep learning empowered computer-assisted diagnosis has been shown its value in medical imaging-based decision making. However, lots of research focus on improving disease detection accuracy but not exploring the evidence of pathology. In this paper, we propose a method to interpret the deep learning classification findings. Firstly, we propose an accurate neural network architecture to classify skin lesion. Secondly, we utilize a prediction difference analysis method that examining each patch on the image through patch wised corrupting for detecting the biomarkers. Lastly, we validate that our biomarker findings are corresponding to the patterns in the literature. The findings might be significant to guide clinical diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge