Robert W. Williams
Enabling clinical use of foundation models in histopathology
Feb 25, 2026Abstract:Foundation models in histopathology are expected to facilitate the development of high-performing and generalisable deep learning systems. However, current models capture not only biologically relevant features, but also pre-analytic and scanner-specific variation that bias the predictions of task-specific models trained from the foundation model features. Here we show that introducing novel robustness losses during training of downstream task-specific models reduces sensitivity to technical variability. A purpose-designed comprehensive experimentation setup with 27,042 WSIs from 6155 patients is used to train thousands of models from the features of eight popular foundation models for computational pathology. In addition to a substantial improvement in robustness, we observe that prediction accuracy improves by focusing on biologically relevant features. Our approach successfully mitigates robustness issues of foundation models for computational pathology without retraining the foundation models themselves, enabling development of robust computational pathology models applicable to real-world data in routine clinical practice.
Artificial Intelligence Models for Cell Type and Subtype Identification Based on Single-Cell RNA Sequencing Data in Vision Science
Sep 26, 2022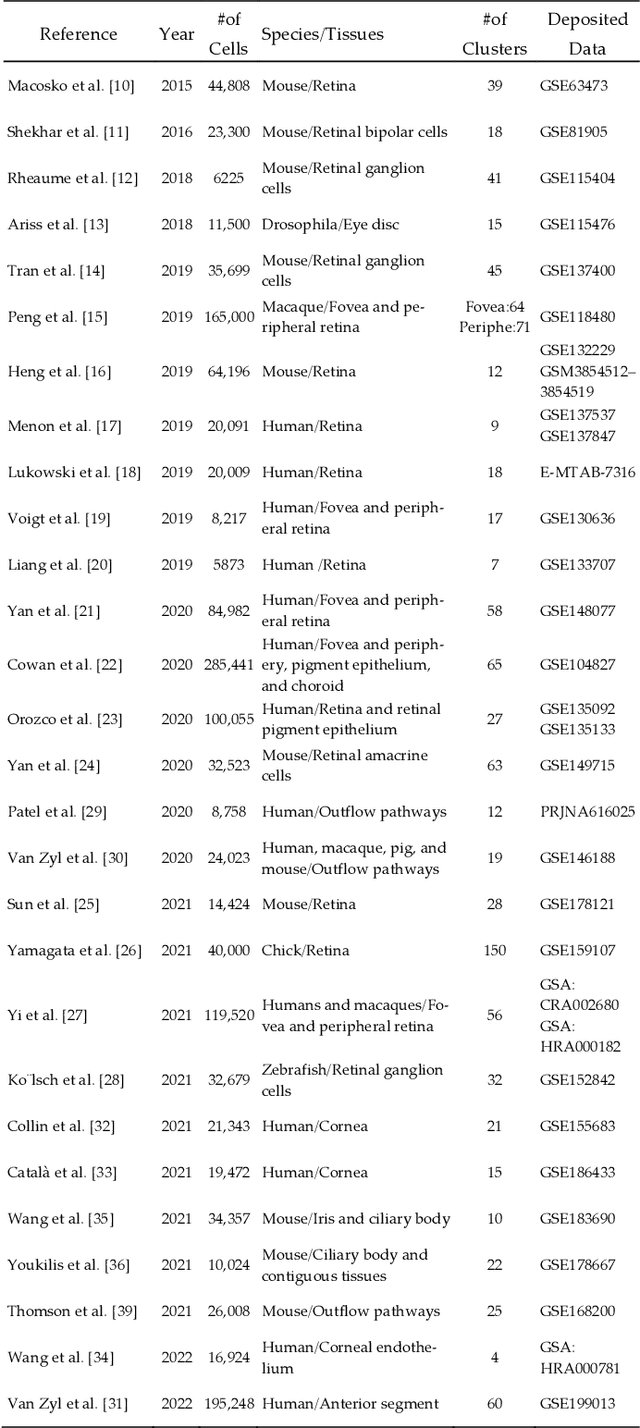
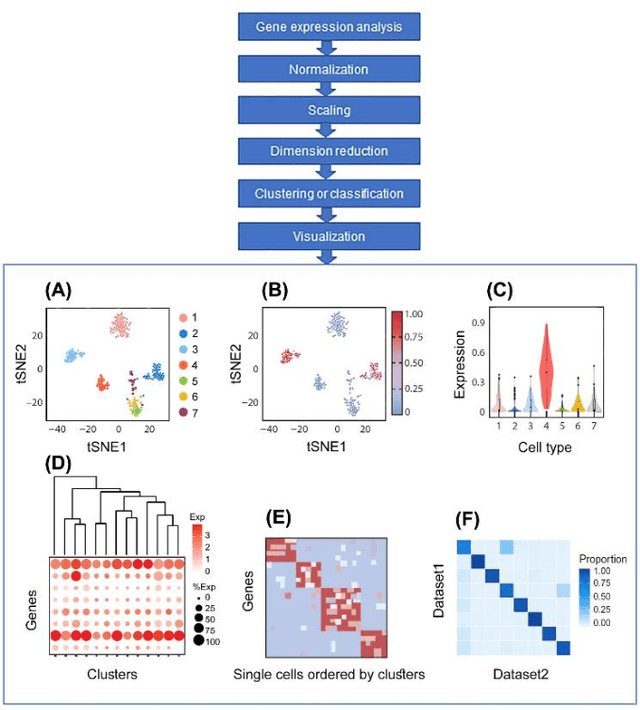
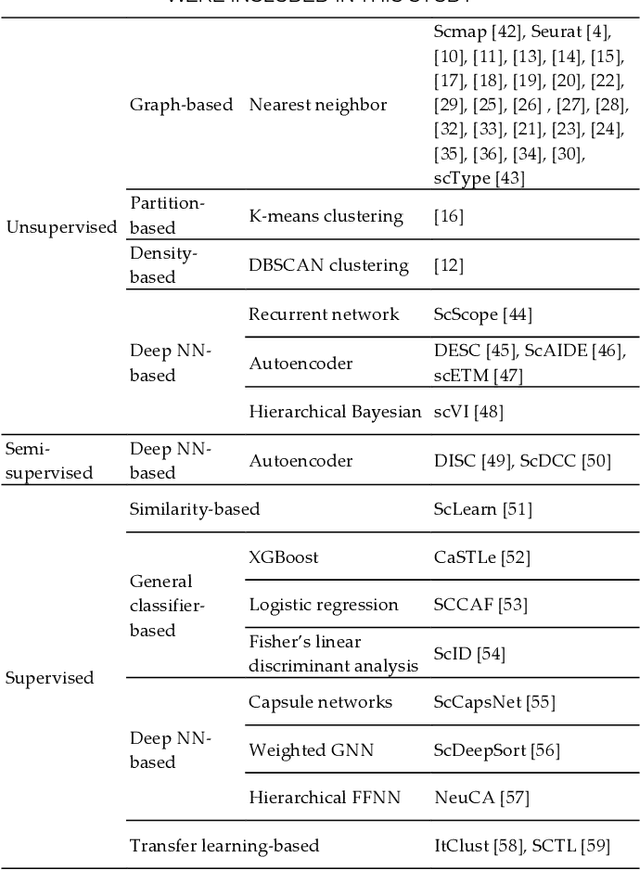
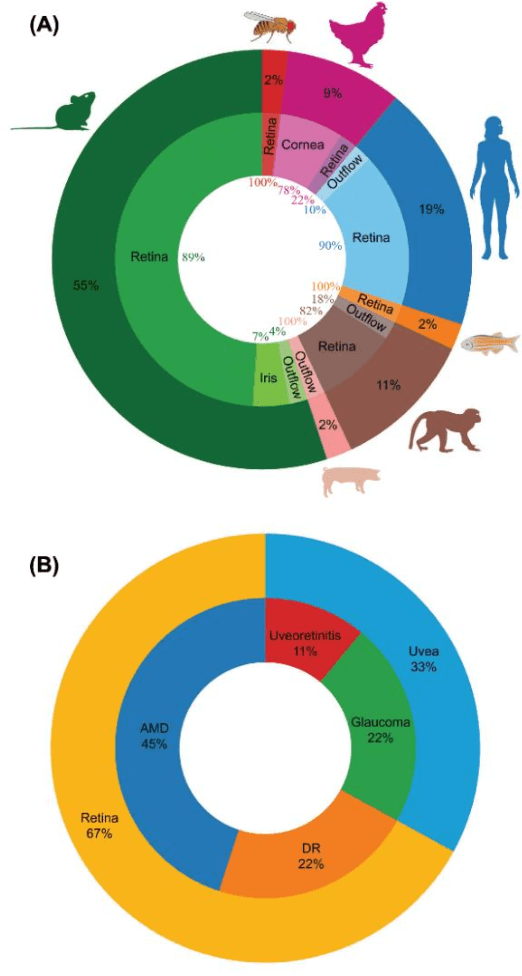
Abstract:Single-cell RNA sequencing (scRNA-seq) provides a high throughput, quantitative and unbiased framework for scientists in many research fields to identify and characterize cell types within heterogeneous cell populations from various tissues. However, scRNA-seq based identification of discrete cell-types is still labor intensive and depends on prior molecular knowledge. Artificial intelligence has provided faster, more accurate, and user-friendly approaches for cell-type identification. In this review, we discuss recent advances in cell-type identification methods using artificial intelligence techniques based on single-cell and single-nucleus RNA sequencing data in vision science.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge