Rhydian Windsor
Automated Spinal MRI Labelling from Reports Using a Large Language Model
Oct 22, 2024Abstract:We propose a general pipeline to automate the extraction of labels from radiology reports using large language models, which we validate on spinal MRI reports. The efficacy of our labelling method is measured on five distinct conditions: spinal cancer, stenosis, spondylolisthesis, cauda equina compression and herniation. Using open-source models, our method equals or surpasses GPT-4 on a held-out set of reports. Furthermore, we show that the extracted labels can be used to train imaging models to classify the identified conditions in the accompanying MR scans. All classifiers trained using automated labels achieve comparable performance to models trained using scans manually annotated by clinicians. Code can be found at https://github.com/robinyjpark/AutoLabelClassifier.
Contouring by Unit Vector Field Regression
May 26, 2023



Abstract:This work introduces a simple deep-learning based method to delineate contours by `walking' along learnt unit vector fields. We demonstrate the effectiveness of our pipeline on the unique case of open contours on the task of delineating the sacroiliac joints (SIJs) in spinal MRIs. We show that: (i) 95% of the time the average root mean square error of the predicted contour against the original ground truth is below 4.5 pixels (2.5mm for a standard T1-weighted SIJ MRI), and (ii) the proposed method is better than the baseline of regressing vertices or landmarks of contours.
Vision-Language Modelling For Radiological Imaging and Reports In The Low Data Regime
Mar 30, 2023



Abstract:This paper explores training medical vision-language models (VLMs) -- where the visual and language inputs are embedded into a common space -- with a particular focus on scenarios where training data is limited, as is often the case in clinical datasets. We explore several candidate methods to improve low-data performance, including: (i) adapting generic pre-trained models to novel image and text domains (i.e. medical imaging and reports) via unimodal self-supervision; (ii) using local (e.g. GLoRIA) & global (e.g. InfoNCE) contrastive loss functions as well as a combination of the two; (iii) extra supervision during VLM training, via: (a) image- and text-only self-supervision, and (b) creating additional positive image-text pairs for training through augmentation and nearest-neighbour search. Using text-to-image retrieval as a benchmark, we evaluate the performance of these methods with variable sized training datasets of paired chest X-rays and radiological reports. Combined, they significantly improve retrieval compared to fine-tuning CLIP, roughly equivalent to training with the data. A similar pattern is found in the downstream task classification of CXR-related conditions with our method outperforming CLIP and also BioVIL, a strong CXR VLM benchmark, in the zero-shot and linear probing settings. We conclude with a set of recommendations for researchers aiming to train vision-language models on other medical imaging modalities when training data is scarce. To facilitate further research, we will make our code and models publicly available.
Context-Aware Transformers For Spinal Cancer Detection and Radiological Grading
Jun 27, 2022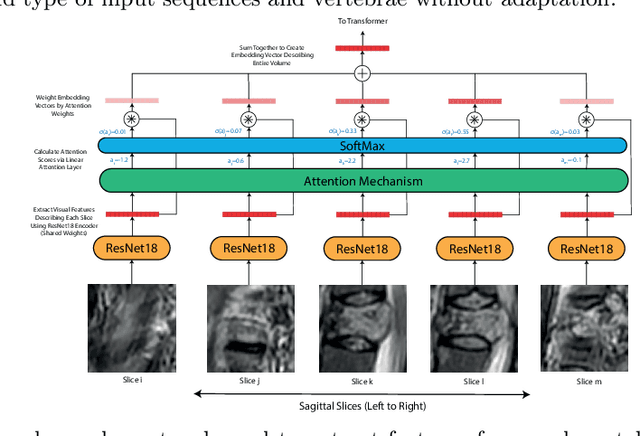
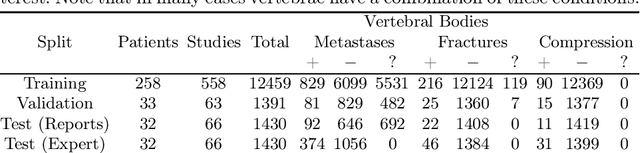

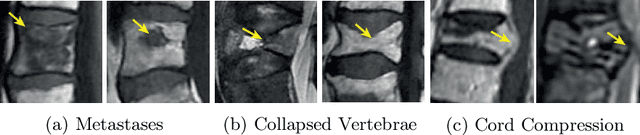
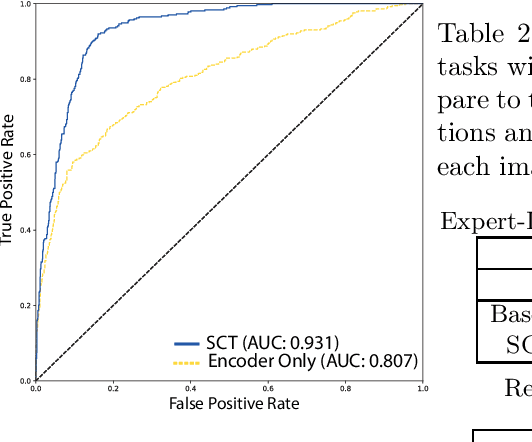
Abstract:This paper proposes a novel transformer-based model architecture for medical imaging problems involving analysis of vertebrae. It considers two applications of such models in MR images: (a) detection of spinal metastases and the related conditions of vertebral fractures and metastatic cord compression, (b) radiological grading of common degenerative changes in intervertebral discs. Our contributions are as follows: (i) We propose a Spinal Context Transformer (SCT), a deep-learning architecture suited for the analysis of repeated anatomical structures in medical imaging such as vertebral bodies (VBs). Unlike previous related methods, SCT considers all VBs as viewed in all available image modalities together, making predictions for each based on context from the rest of the spinal column and all available imaging modalities. (ii) We apply the architecture to a novel and important task: detecting spinal metastases and the related conditions of cord compression and vertebral fractures/collapse from multi-series spinal MR scans. This is done using annotations extracted from free-text radiological reports as opposed to bespoke annotation. However, the resulting model shows strong agreement with vertebral-level bespoke radiologist annotations on the test set. (iii) We also apply SCT to an existing problem: radiological grading of inter-vertebral discs (IVDs) in lumbar MR scans for common degenerative changes.We show that by considering the context of vertebral bodies in the image, SCT improves the accuracy for several gradings compared to previously published model.
SpineNetV2: Automated Detection, Labelling and Radiological Grading Of Clinical MR Scans
May 03, 2022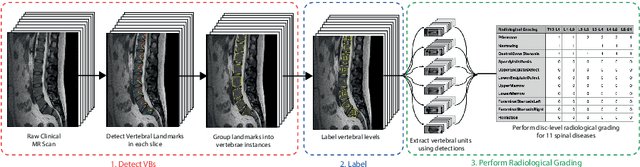
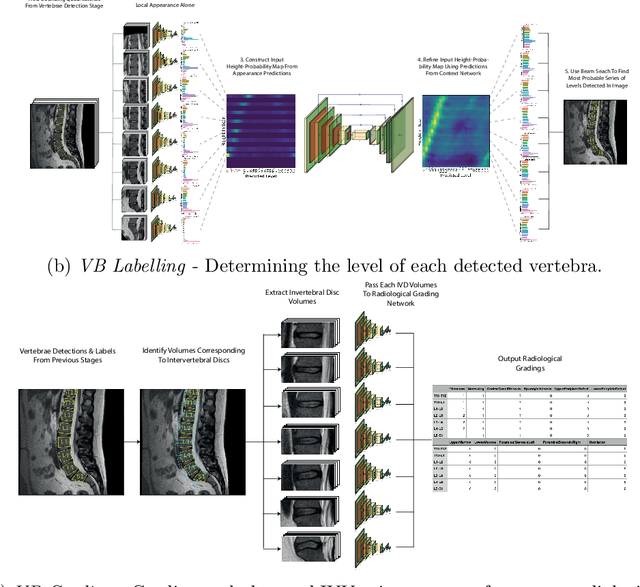
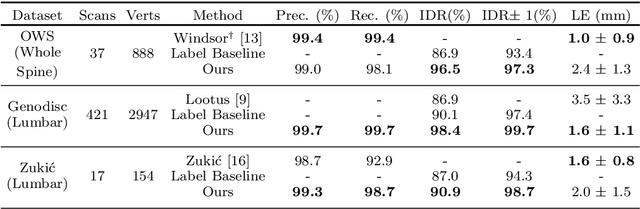
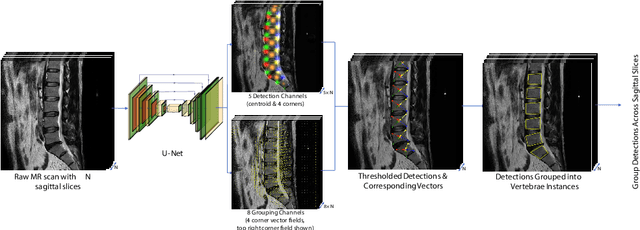
Abstract:This technical report presents SpineNetV2, an automated tool which: (i) detects and labels vertebral bodies in clinical spinal magnetic resonance (MR) scans across a range of commonly used sequences; and (ii) performs radiological grading of lumbar intervertebral discs in T2-weighted scans for a range of common degenerative changes. SpineNetV2 improves over the original SpineNet software in two ways: (1) The vertebral body detection stage is significantly faster, more accurate and works across a range of fields-of-view (as opposed to just lumbar scans). (2) Radiological grading adopts a more powerful architecture, adding several new grading schemes without loss in performance. A demo of the software is available at the project website: http://zeus.robots.ox.ac.uk/spinenet2/.
Self-Supervised Multi-Modal Alignment for Whole Body Medical Imaging
Aug 06, 2021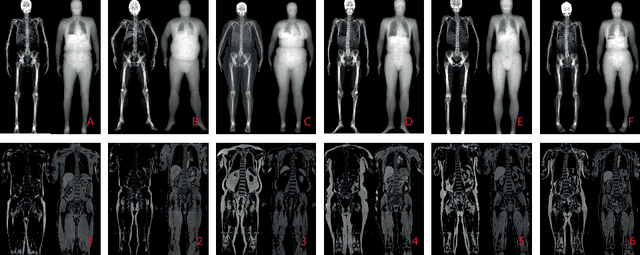
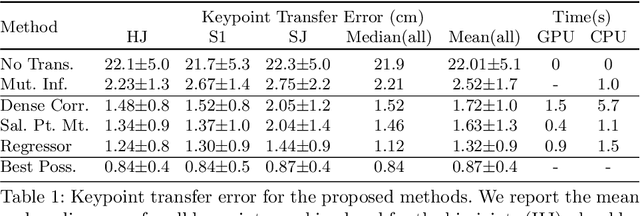
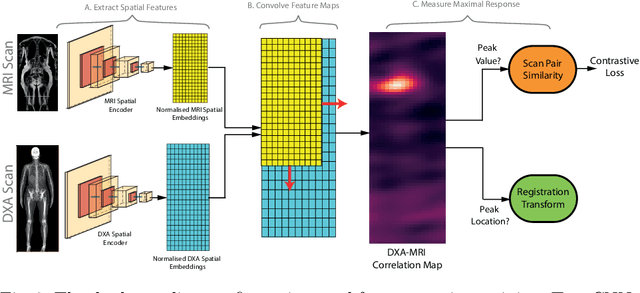
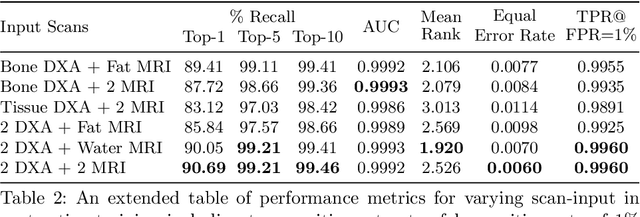
Abstract:This paper explores the use of self-supervised deep learning in medical imaging in cases where two scan modalities are available for the same subject. Specifically, we use a large publicly-available dataset of over 20,000 subjects from the UK Biobank with both whole body Dixon technique magnetic resonance (MR) scans and also dual-energy x-ray absorptiometry (DXA) scans. We make three contributions: (i) We introduce a multi-modal image-matching contrastive framework, that is able to learn to match different-modality scans of the same subject with high accuracy. (ii) Without any adaption, we show that the correspondences learnt during this contrastive training step can be used to perform automatic cross-modal scan registration in a completely unsupervised manner. (iii) Finally, we use these registrations to transfer segmentation maps from the DXA scans to the MR scans where they are used to train a network to segment anatomical regions without requiring ground-truth MR examples. To aid further research, our code will be made publicly available.
A Convolutional Approach to Vertebrae Detection and Labelling in Whole Spine MRI
Jul 13, 2020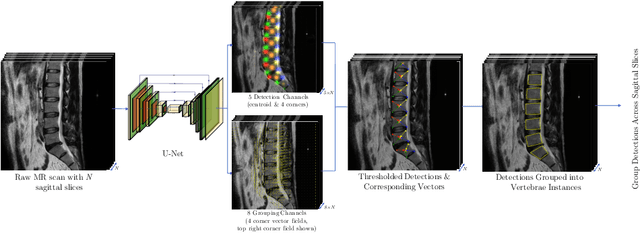
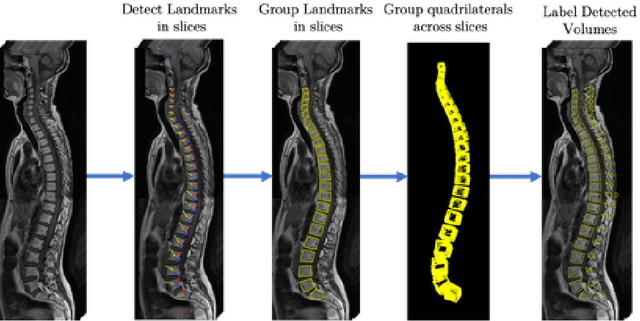

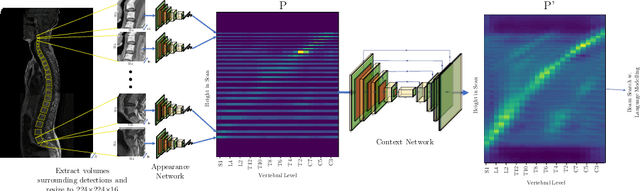
Abstract:We propose a novel convolutional method for the detection and identification of vertebrae in whole spine MRIs. This involves using a learnt vector field to group detected vertebrae corners together into individual vertebral bodies and convolutional image-to-image translation followed by beam search to label vertebral levels in a self-consistent manner. The method can be applied without modification to lumbar, cervical and thoracic-only scans across a range of different MR sequences. The resulting system achieves 98.1% detection rate and 96.5% identification rate on a challenging clinical dataset of whole spine scans and matches or exceeds the performance of previous systems on lumbar-only scans. Finally, we demonstrate the clinical applicability of this method, using it for automated scoliosis detection in both lumbar and whole spine MR scans.
The Ladder Algorithm: Finding Repetitive Structures in Medical Images by Induction
Jan 30, 2020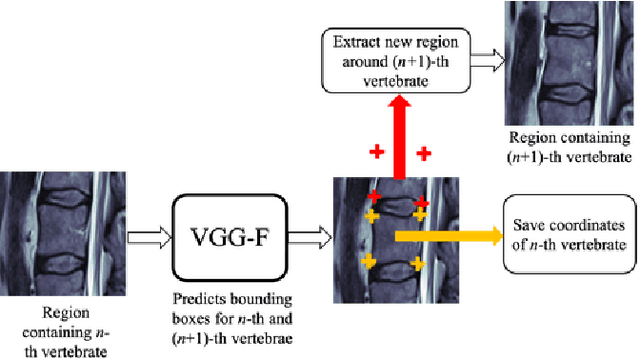
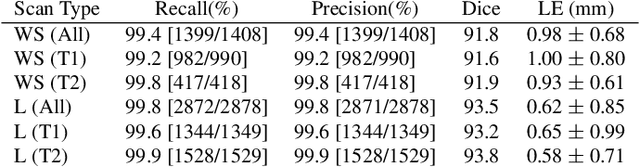
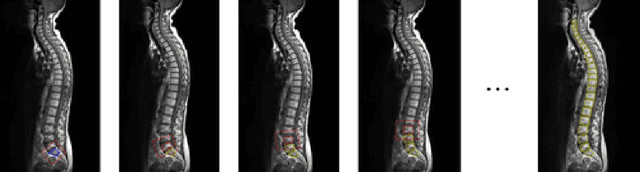
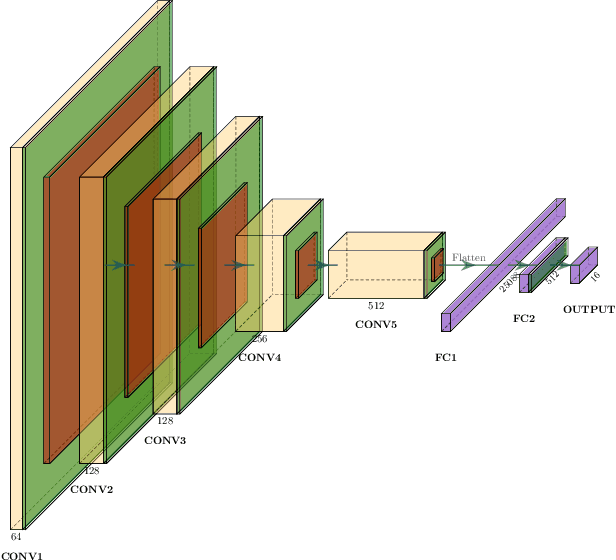
Abstract:In this paper we introduce the Ladder Algorithm; a novel recurrent algorithm to detect repetitive structures in natural images with high accuracy using little training data. We then demonstrate the algorithm on the task of extracting vertebrae from whole spine magnetic resonance scans with only lumbar MR scans for training data. It is shown to achieve high perforamance with 99.8% precision and recall, exceeding current state of the art approaches for lumbar vertebrae detection in T1 and T2 weighted scans. It also generalises without retraining to whole spine images with minimal drop in accuracy, achieving 99.4% detection rate.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge