Redha Ali
Convolutional neural network based on transfer learning for breast cancer screening
Dec 22, 2021
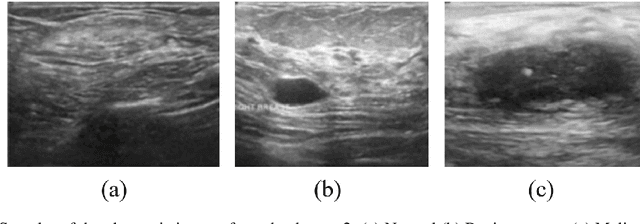
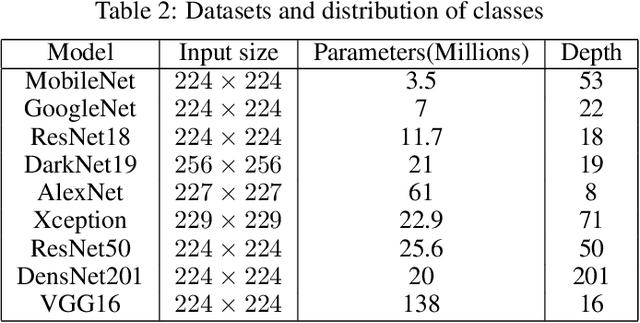
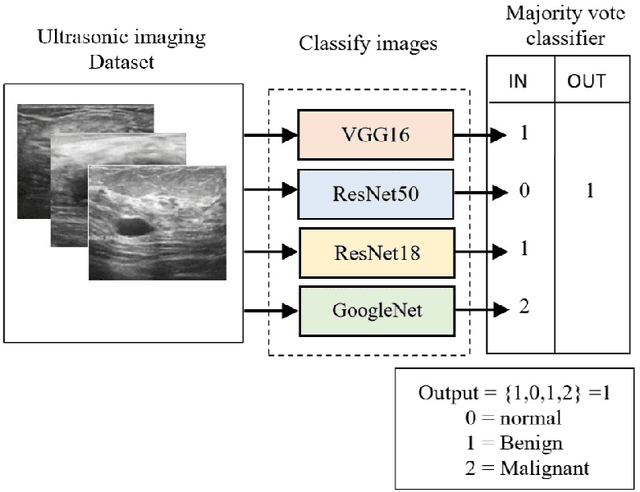
Abstract:Breast cancer is the most common cancer in the world and the most prevalent cause of death among women worldwide. Nevertheless, it is also one of the most treatable malignancies if detected early. In this paper, a deep convolutional neural network-based algorithm is proposed to aid in accurately identifying breast cancer from ultrasonic images. In this algorithm, several neural networks are fused in a parallel architecture to perform the classification process and the voting criteria are applied in the final classification decision between the candidate object classes where the output of each neural network is representing a single vote. Several experiments were conducted on the breast ultrasound dataset consisting of 537 Benign, 360 malignant, and 133 normal images. These experiments show an optimistic result and a capability of the proposed model to outperform many state-of-the-art algorithms on several measures. Using k-fold cross-validation and a bagging classifier ensemble, we achieved an accuracy of 99.5% and a sensitivity of 99.6%.
Skin lesion segmentation and classification using deep learning and handcrafted features
Dec 20, 2021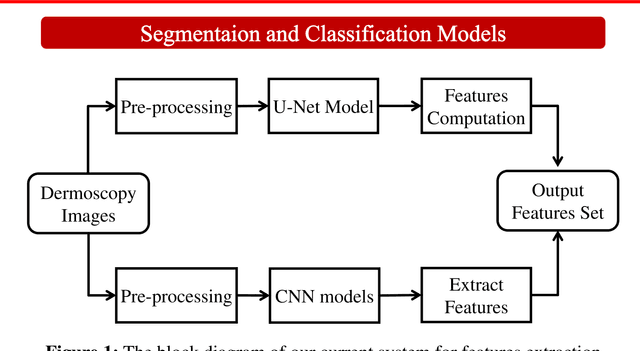
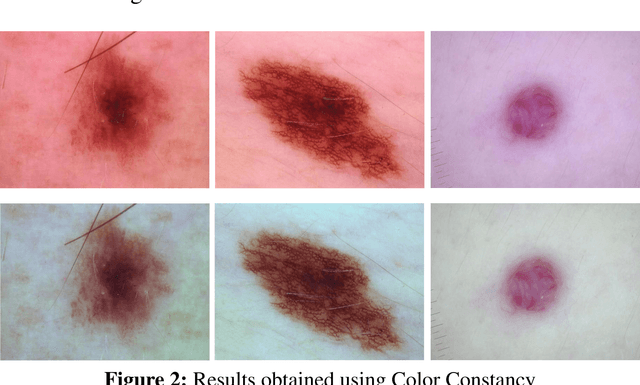

Abstract:Accurate diagnostics of a skin lesion is a critical task in classification dermoscopic images. In this research, we form a new type of image features, called hybrid features, which has stronger discrimination ability than single method features. This study involves a new technique where we inject the handcrafted features or feature transfer into the fully connected layer of Convolutional Neural Network (CNN) model during the training process. Based on our literature review until now, no study has examined or investigated the impact on classification performance by injecting the handcrafted features into the CNN model during the training process. In addition, we also investigated the impact of segmentation mask and its effect on the overall classification performance. Our model achieves an 92.3% balanced multiclass accuracy, which is 6.8% better than the typical single method classifier architecture for deep learning.
Fused Deep Convolutional Neural Network for Precision Diagnosis of COVID-19 Using Chest X-Ray Images
Sep 15, 2020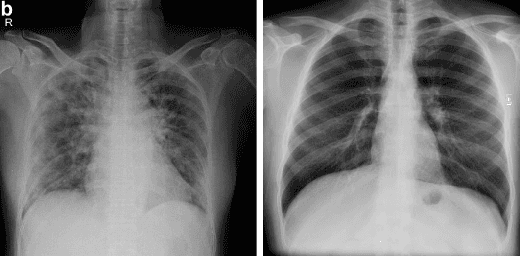
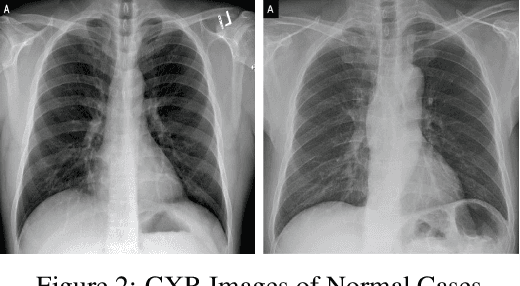
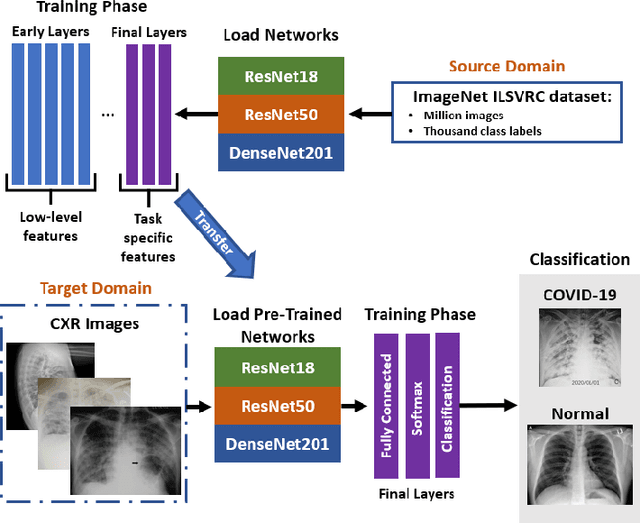
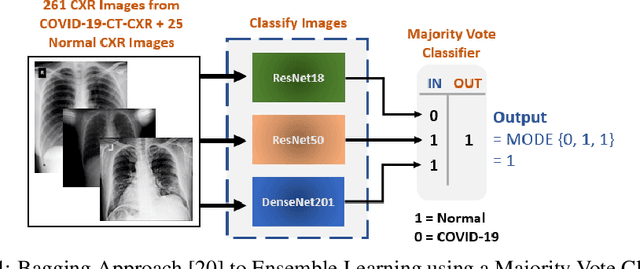
Abstract:With a Coronavirus disease (COVID-19) case count exceeding 10 million worldwide, there is an increased need for a diagnostic capability. The main variables in increasing diagnostic capability are reduced cost, turnaround or diagnosis time, and upfront equipment cost and accessibility. Two candidates for machine learning COVID-19 diagnosis are Computed Tomography (CT) scans and plain chest X-rays. While CT scans score higher in sensitivity, they have a higher cost, maintenance requirement, and turnaround time as compared to plain chest X-rays. The use of portable chest X-radiograph (CXR) is recommended by the American College of Radiology (ACR) since using CT places a massive burden on radiology services. Therefore, X-ray imagery paired with machine learning techniques is proposed a first-line triage tool for COVID-19 diagnostics. In this paper we propose a computer-aided diagnosis (CAD) to accurately classify chest X-ray scans of COVID-19 and normal subjects by fine-tuning several neural networks (ResNet18, ResNet50, DenseNet201) pre-trained on the ImageNet dataset. These neural networks are fused in a parallel architecture and the voting criteria are applied in the final classification decision between the candidate object classes where the output of each neural network is representing a single vote. Several experiments are conducted on the weakly labeled COVID-19-CT-CXR dataset consisting of 263 COVID-19 CXR images extracted from PubMed Central Open Access subsets combined with 25 normal classification CXR images. These experiments show an optimistic result and a capability of the proposed model to outperforming many state-of-the-art algorithms on several measures. Using k-fold cross-validation and a bagging classifier ensemble, we achieve an accuracy of 99.7% and a sensitivity of 100%.
Understanding Deep Neural Network Predictions for Medical Imaging Applications
Dec 20, 2019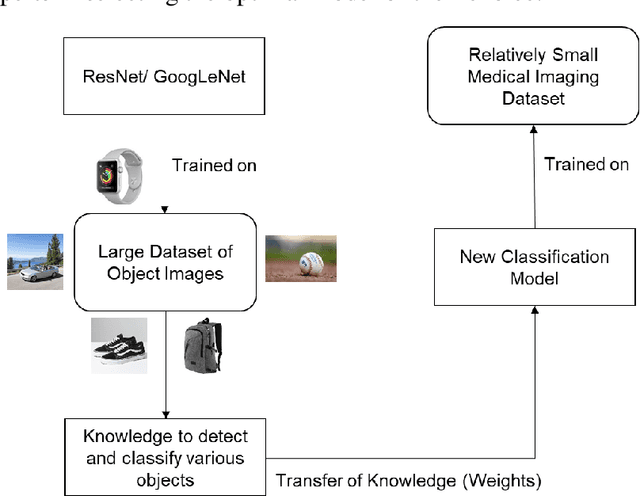
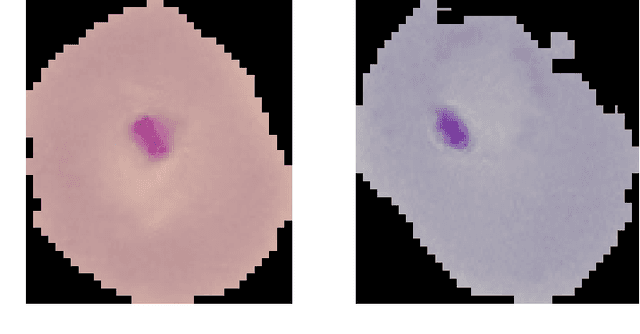
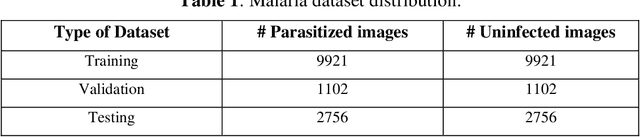

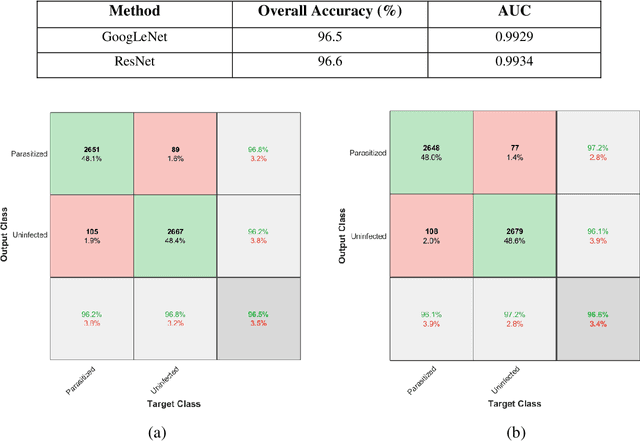
Abstract:Computer-aided detection has been a research area attracting great interest in the past decade. Machine learning algorithms have been utilized extensively for this application as they provide a valuable second opinion to the doctors. Despite several machine learning models being available for medical imaging applications, not many have been implemented in the real-world due to the uninterpretable nature of the decisions made by the network. In this paper, we investigate the results provided by deep neural networks for the detection of malaria, diabetic retinopathy, brain tumor, and tuberculosis in different imaging modalities. We visualize the class activation mappings for all the applications in order to enhance the understanding of these networks. This type of visualization, along with the corresponding network performance metrics, would aid the data science experts in better understanding of their models as well as assisting doctors in their decision-making process.
Skin Lesion Segmentation and Classification for ISIC 2018 by Combining Deep CNN and Handcrafted Features
Aug 14, 2019
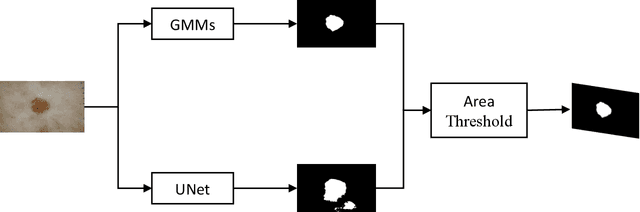
Abstract:This short report describes our submission to the ISIC 2018 Challenge in Skin Lesion Analysis Towards Melanoma Detection for Task1 and Task 3. This work has been accomplished by a team of researchers at the University of Dayton Signal and Image Processing Lab. Our proposed approach is computationally efficient are combines information from both deep learning and handcrafted features. For Task3, we form a new type of image features, called hybrid features, which has stronger discrimination ability than single method features. These features are utilized as inputs to a decision-making model that is based on a multiclass Support Vector Machine (SVM) classifier. The proposed technique is evaluated on online validation databases. Our score was 0.841 with SVM classifier on the validation dataset.
Skin Lesion Segmentation and Classification for ISIC 2018 Using Traditional Classifiers with Hand-Crafted Features
Jul 18, 2018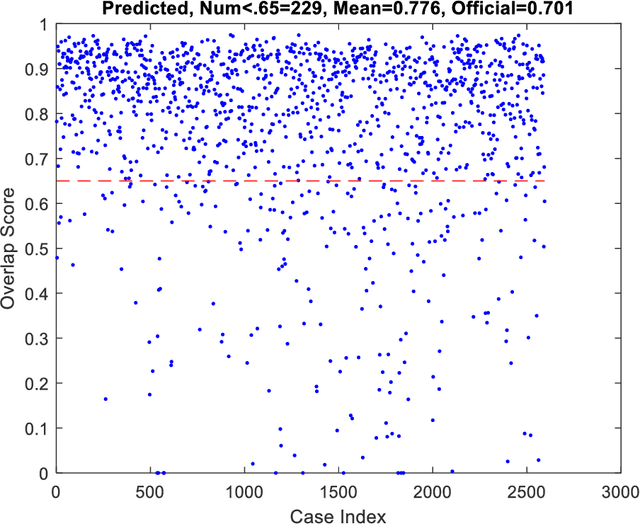
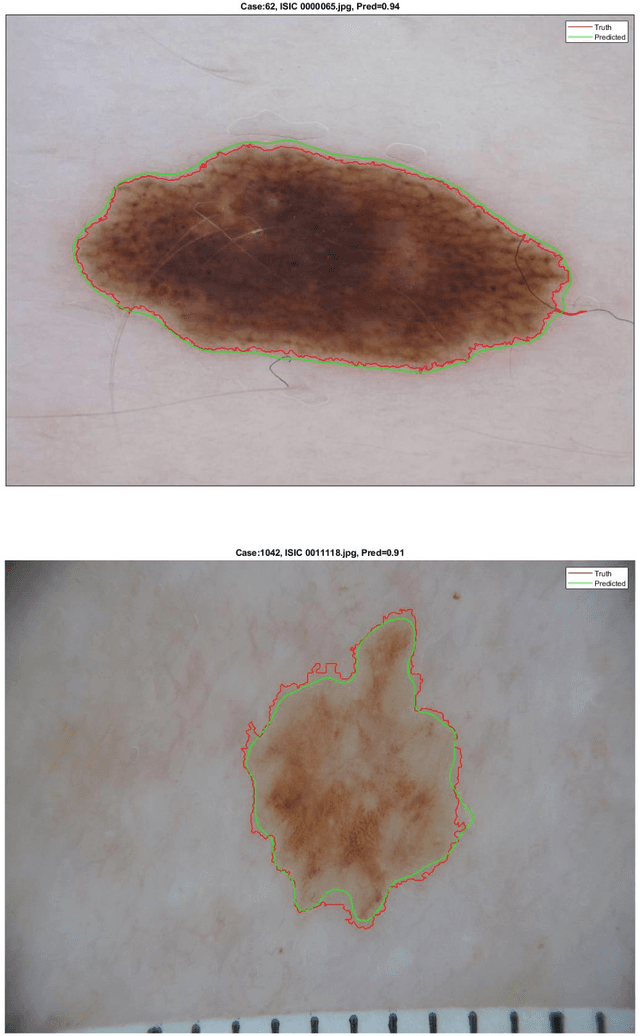
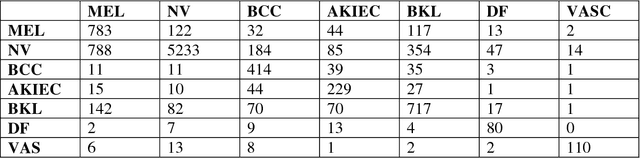

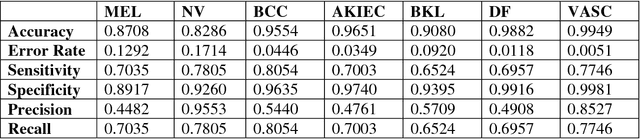
Abstract:This paper provides the required description of the methods used to obtain submitted results for Task1 and Task 3 of ISIC 2018: Skin Lesion Analysis Towards Melanoma Detection. The results have been created by a team of researchers at the University of Dayton Signal and Image Processing Lab. In this submission, traditional classifiers with hand-crafted features are utilized for Task 1 and Task 3. Our team is providing additional separate submissions using deep learning methods for comparison.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge