Rainer Heintzmann
PtyLab.m/py/jl: a cross-platform, open-source inverse modeling toolbox for conventional and Fourier ptychography
Jan 16, 2023



Abstract:Conventional (CP) and Fourier (FP) ptychography have emerged as versatile quantitative phase imaging techniques. While the main application cases for each technique are different, namely lens-less short wavelength imaging for CP and lens-based visible light imaging for FP, both methods share a common algorithmic ground. CP and FP have in part independently evolved to include experimentally robust forward models and inversion techniques. This separation has resulted in a plethora of algorithmic extensions, some of which have not crossed the boundary from one modality to the other. Here, we present an open source, cross-platform software, called PtyLab, enabling both CP and FP data analysis in a unified framework. With this framework, we aim to facilitate and accelerate cross-pollination between the two techniques. Moreover, the availability in Matlab, Python, and Julia will set a low barrier to enter each field.
Guided-deconvolution for Correlative Light and Electron Microscopy
Aug 19, 2022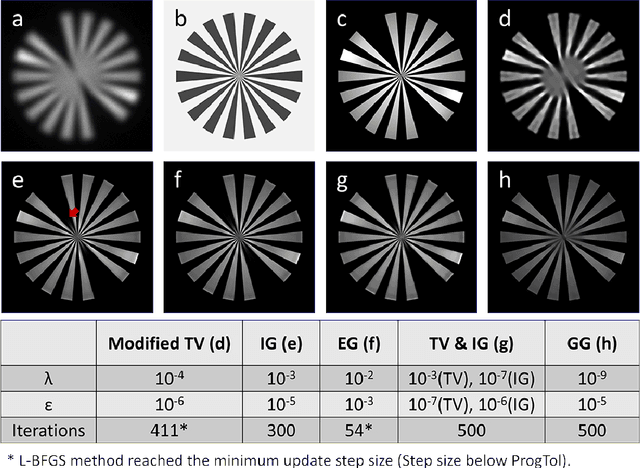
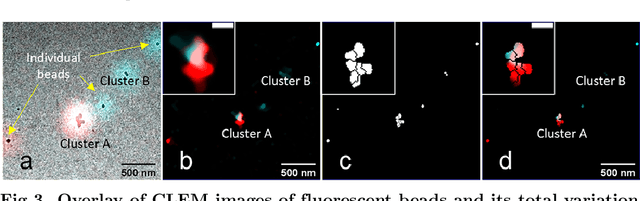
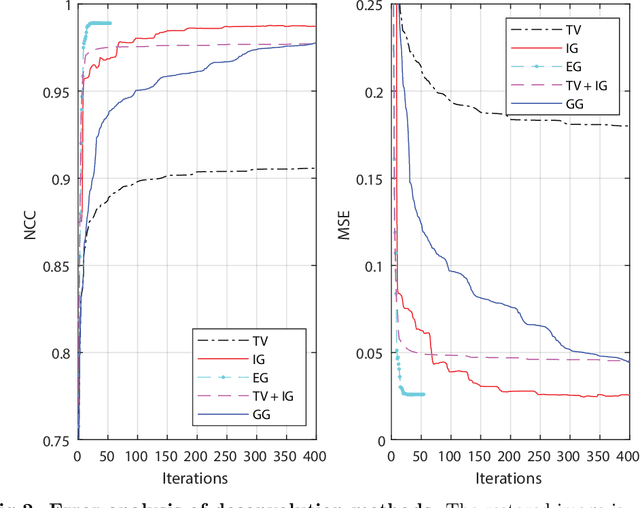

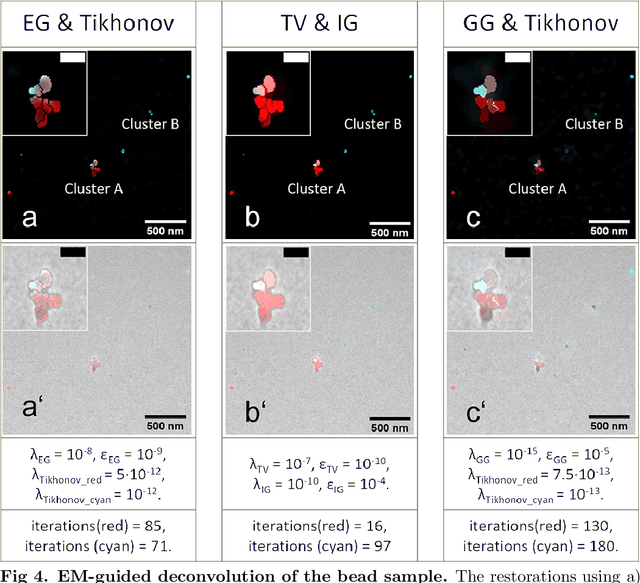
Abstract:Correlative light and electron microscopy is a powerful tool to study the internal structure of cells. It combines the mutual benefit of correlating light (LM) and electron (EM) microscopy information. However, the classical approach of overlaying LM onto EM images to assign functional to structural information is hampered by the large discrepancy in structural detail visible in the LM images. This paper aims at investigating an optimized approach which we call EM-guided deconvolution. It attempts to automatically assign fluorescence-labelled structures to details visible in the EM image to bridge the gaps in both resolution and specificity between the two imaging modes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge