Radhakrishnan Nagarajan
Deciphering Dynamical Nonlinearities in Short Time Series Using Recurrent Neural Networks
Jul 15, 2019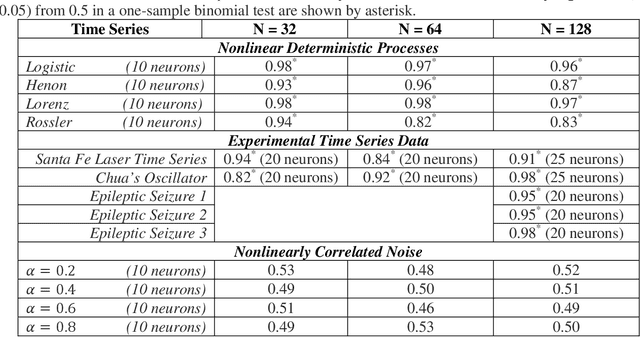
Abstract:Surrogate testing techniques have been used widely to investigate the presence of dynamical nonlinearities, an essential ingredient of deterministic chaotic processes. Traditional surrogate testing subscribes to statistical hypothesis testing and investigates potential differences in discriminant statistics between the given empirical sample and its surrogate counterparts. The choice and estimation of the discriminant statistics can be challenging across short time series. Also, conclusion based on a single empirical sample is an inherent limitation. The present study proposes a recurrent neural network classification framework that uses the raw time series obviating the need for discriminant statistic while accommodating multiple time series realizations for enhanced generalizability of the findings. The results are demonstrated on short time series with lengths (L = 32, 64, 128) from continuous and discrete dynamical systems in chaotic regimes, nonlinear transform of linearly correlated noise and experimental data. Accuracy of the classifier is shown to be markedly higher than >> 50% for the processes in chaotic regimes whereas those of nonlinearly correlated noise were around ~50% similar to that of random guess from a one-sample binomial test. These results are promising and elucidate the usefulness of the proposed framework in identifying potential dynamical nonlinearities from short experimental time series.
On Identifying Significant Edges in Graphical Models of Molecular Networks
Apr 23, 2013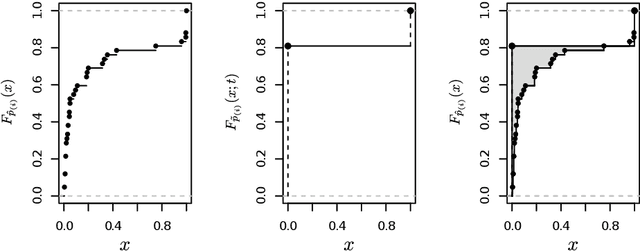
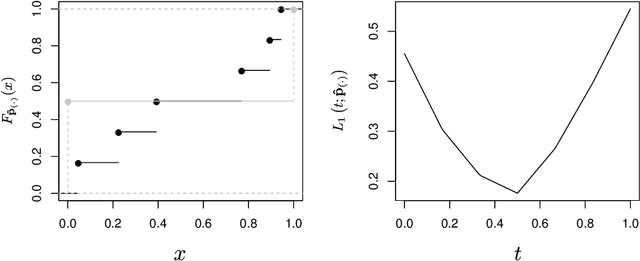
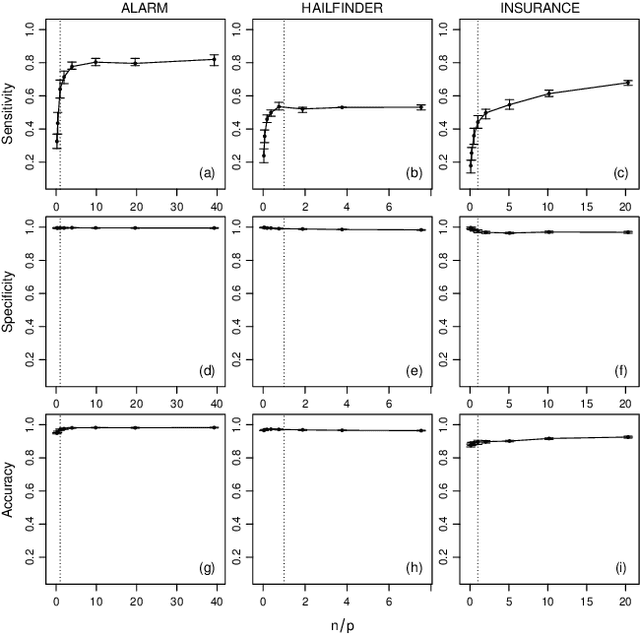
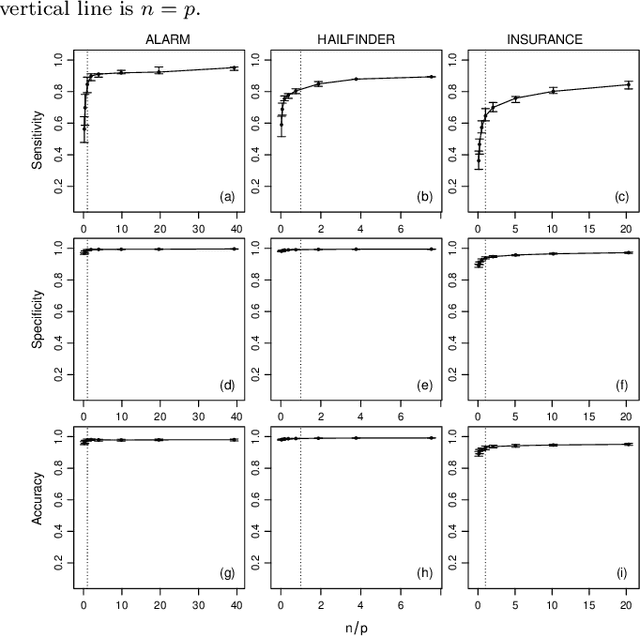
Abstract:Objective: Modelling the associations from high-throughput experimental molecular data has provided unprecedented insights into biological pathways and signalling mechanisms. Graphical models and networks have especially proven to be useful abstractions in this regard. Ad-hoc thresholds are often used in conjunction with structure learning algorithms to determine significant associations. The present study overcomes this limitation by proposing a statistically-motivated approach for identifying significant associations in a network. Methods and Materials: A new method that identifies significant associations in graphical models by estimating the threshold minimising the $L_{\mathrm{1}}$ norm between the cumulative distribution function (CDF) of the observed edge confidences and those of its asymptotic counterpart is proposed. The effectiveness of the proposed method is demonstrated on popular synthetic data sets as well as publicly available experimental molecular data corresponding to gene and protein expression profiles. Results: The improved performance of the proposed approach is demonstrated across the synthetic data sets using sensitivity, specificity and accuracy as performance metrics. The results are also demonstrated across varying sample sizes and three different structure learning algorithms with widely varying assumptions. In all cases, the proposed approach has specificity and accuracy close to 1, while sensitivity increases linearly in the logarithm of the sample size. The estimated threshold systematically outperforms common ad-hoc ones in terms of sensitivity while maintaining comparable levels of specificity and accuracy. Networks from experimental data sets are reconstructed accurately with respect to the results from the original papers.
* 21 pages, 9 figures. Presented at the Conference for Artificial Intelligence in Medicine (AIME '11), Workshop on Probabilistic Problem Solving in Biomedicine
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge