R. M. Farouk
Sparse Representation and Non-Negative Matrix Factorization for image denoise
Jul 05, 2018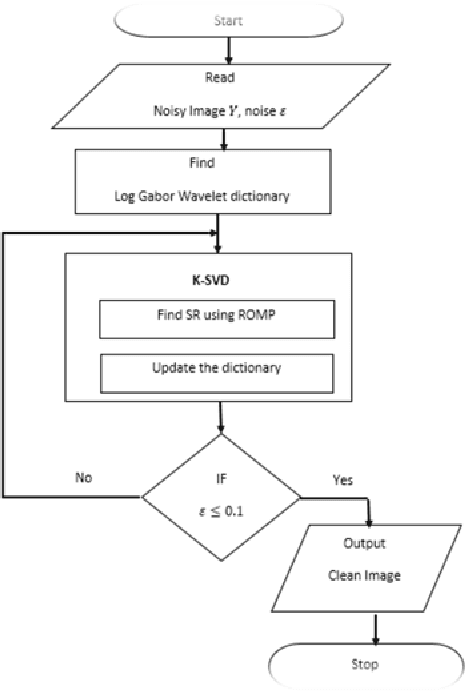
Abstract:Recently, the problem of blind image separation has been widely investigated, especially the medical image denoise which is the main step in medical diag-nosis. Removing the noise without affecting relevant features of the image is the main goal. Sparse decomposition over redundant dictionaries become of the most used approaches to solve this problem. NMF codes naturally favor sparse, parts-based representations. In sparse representation, signals represented as a linear combination of a redundant dictionary atoms. In this paper, we propose an algorithm based on sparse representation over the redundant dictionary and Non-Negative Matrix Factorization (N-NMF). The algorithm initializes a dic-tionary based on training samples constructed from noised image, then it searches for the best representation for the source by using the approximate matching pursuit (AMP). The proposed N-NMF gives a better reconstruction of an image from denoised one. We have compared our numerical results with different image denoising techniques and we have found the performance of the proposed technique is promising. Keywords: Image denoising, sparse representation, dictionary learning, matching pursuit, non-negative matrix factorization.
Blind image separation based on exponentiated transmuted Weibull distribution
May 11, 2016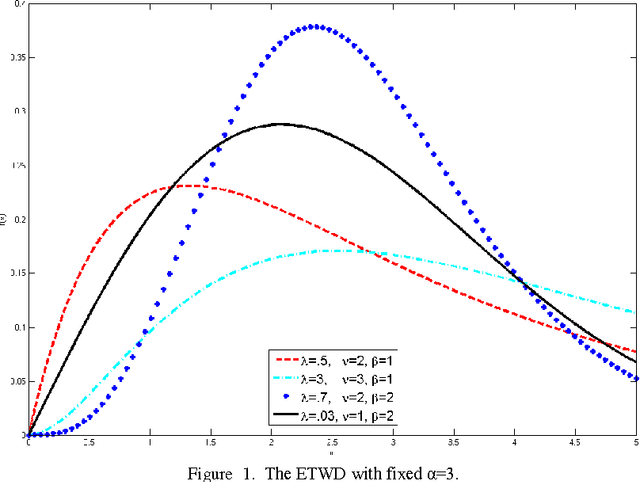
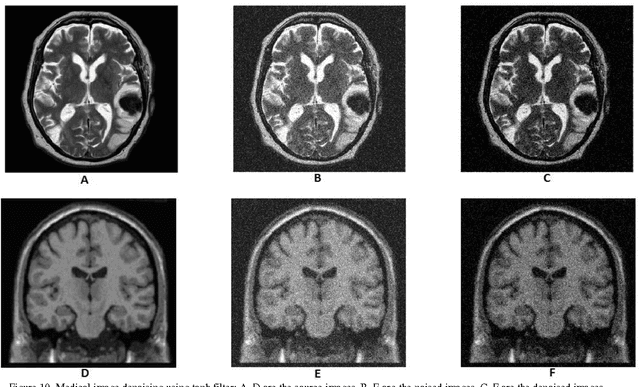
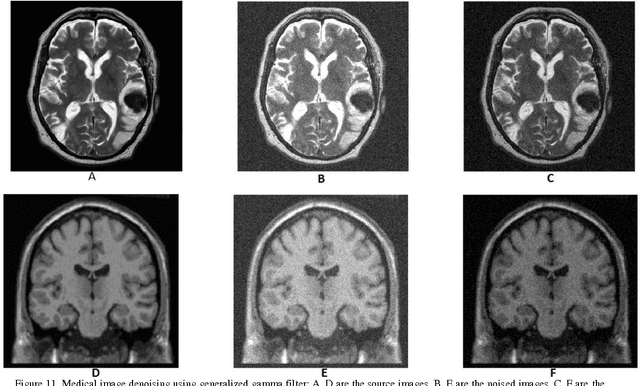
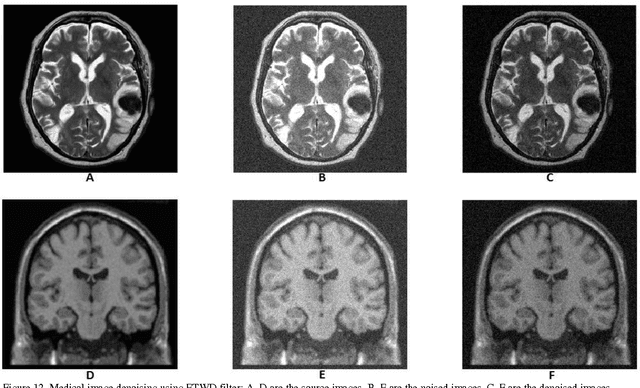
Abstract:In recent years the processing of blind image separation has been investigated. As a result, a number of feature extraction algorithms for direct application of such image structures have been developed. For example, separation of mixed fingerprints found in any crime scene, in which a mixture of two or more fingerprints may be obtained, for identification, we have to separate them. In this paper, we have proposed a new technique for separating a multiple mixed images based on exponentiated transmuted Weibull distribution. To adaptively estimate the parameters of such score functions, an efficient method based on maximum likelihood and genetic algorithm will be used. We also calculate the accuracy of this proposed distribution and compare the algorithmic performance using the efficient approach with other previous generalized distributions. We find from the numerical results that the proposed distribution has flexibility and an efficient result
Robust cDNA microarray image segmentation and analysis technique based on Hough circle transform
Mar 23, 2016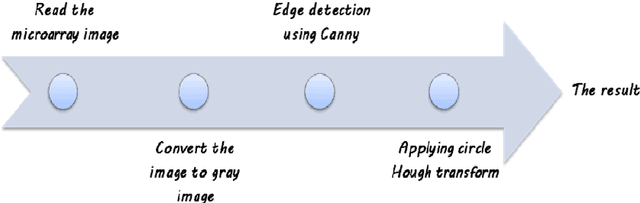
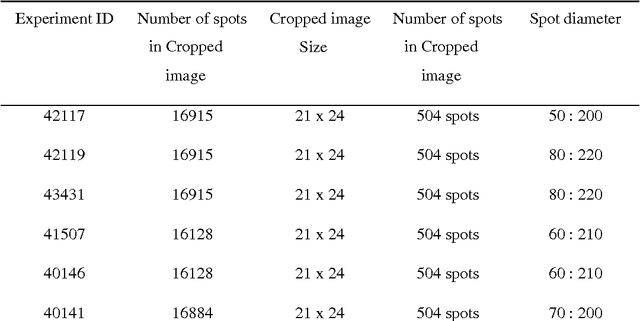
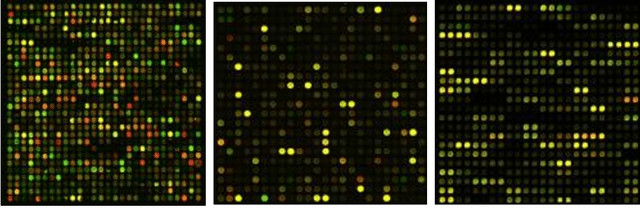
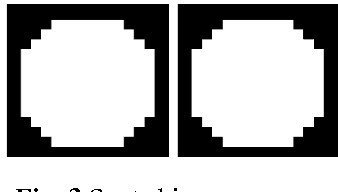
Abstract:One of the most challenging tasks in microarray image analysis is spot segmentation. A solution to this problem is to provide an algorithm than can be used to find any spot within the microarray image. Circular Hough Transformation (CHT) is a powerful feature extraction technique used in image analysis, computer vision, and digital image processing. CHT algorithm is applied on the cDNA microarray images to develop the accuracy and the efficiency of the spots localization, addressing and segmentation process. The purpose of the applied technique is to find imperfect instances of spots within a certain class of circles by applying a voting procedure on the cDNA microarray images for spots localization, addressing and characterizing the pixels of each spot into foreground pixels and background simultaneously. Intensive experiments on the University of North Carolina (UNC) microarray database indicate that the proposed method is superior to the K-means method and the Support vector machine (SVM). Keywords: Hough circle transformation, cDNA microarray image analysis, cDNA microarray image segmentation, spots localization and addressing, spots segmentation
* 13 Pages,12 figures,FSP JOURNAL ISSN:1955-2068,Vol.9, Issue.7, part.1
Recognition of cDNA microarray image Using Feedforward artificial neural network
Oct 09, 2014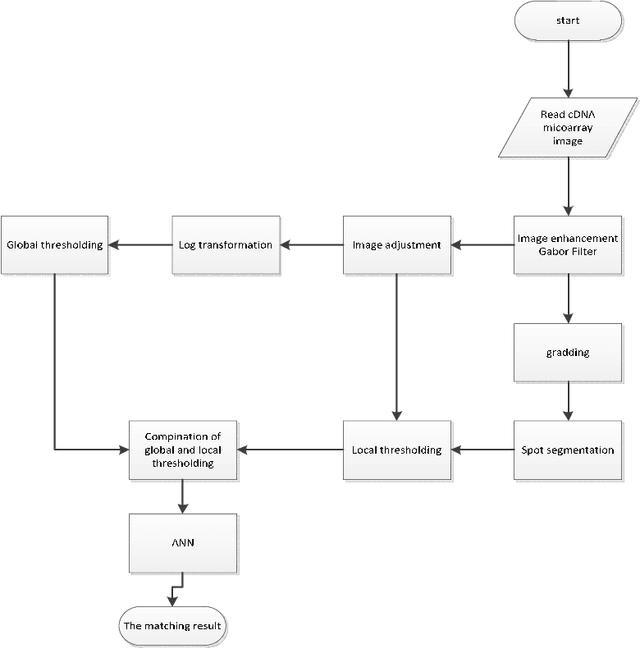
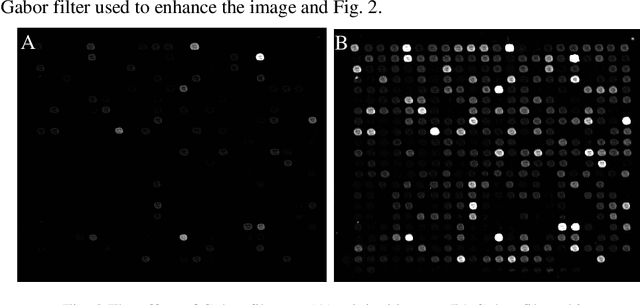
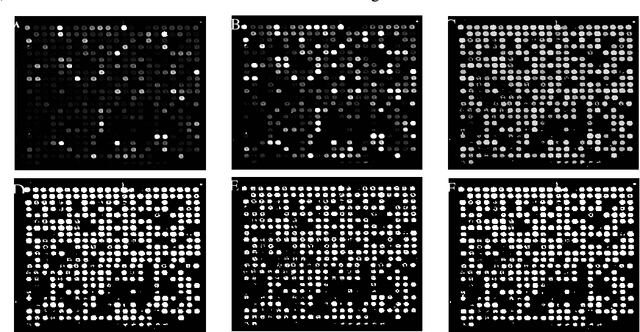

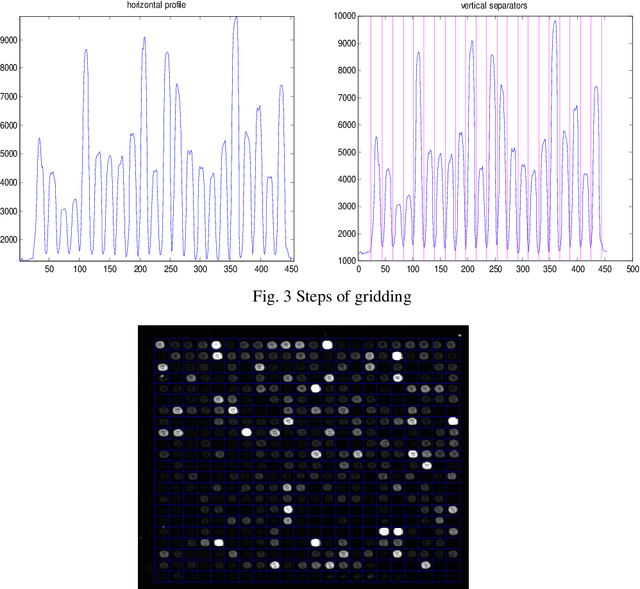

Abstract:The complementary DNA (cDNA) sequence is considered to be the magic biometric technique for personal identification. In this paper, we present a new method for cDNA recognition based on the artificial neural network (ANN). Microarray imaging is used for the concurrent identification of thousands of genes. We have segmented the location of the spots in a cDNA microarray. Thus, a precise localization and segmenting of a spot are essential to obtain a more accurate intensity measurement, leading to a more precise expression measurement of a gene. The segmented cDNA microarray image is resized and it is used as an input for the proposed artificial neural network. For matching and recognition, we have trained the artificial neural network. Recognition results are given for the galleries of cDNA sequences . The numerical results show that, the proposed matching technique is an effective in the cDNA sequences process. We also compare our results with previous results and find out that, the proposed technique is an effective matching performance.
* 17 pages, 7 figures and 23 References
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge