Piotr Sokół
Back to the Continuous Attractor
Jul 31, 2024



Abstract:Continuous attractors offer a unique class of solutions for storing continuous-valued variables in recurrent system states for indefinitely long time intervals. Unfortunately, continuous attractors suffer from severe structural instability in general--they are destroyed by most infinitesimal changes of the dynamical law that defines them. This fragility limits their utility especially in biological systems as their recurrent dynamics are subject to constant perturbations. We observe that the bifurcations from continuous attractors in theoretical neuroscience models display various structurally stable forms. Although their asymptotic behaviors to maintain memory are categorically distinct, their finite-time behaviors are similar. We build on the persistent manifold theory to explain the commonalities between bifurcations from and approximations of continuous attractors. Fast-slow decomposition analysis uncovers the persistent manifold that survives the seemingly destructive bifurcation. Moreover, recurrent neural networks trained on analog memory tasks display approximate continuous attractors with predicted slow manifold structures. Therefore, continuous attractors are functionally robust and remain useful as a universal analogy for understanding analog memory.
Graph Vertex Embeddings: Distance, Regularization and Community Detection
Apr 09, 2024



Abstract:Graph embeddings have emerged as a powerful tool for representing complex network structures in a low-dimensional space, enabling the use of efficient methods that employ the metric structure in the embedding space as a proxy for the topological structure of the data. In this paper, we explore several aspects that affect the quality of a vertex embedding of graph-structured data. To this effect, we first present a family of flexible distance functions that faithfully capture the topological distance between different vertices. Secondly, we analyze vertex embeddings as resulting from a fitted transformation of the distance matrix rather than as a direct result of optimization. Finally, we evaluate the effectiveness of our proposed embedding constructions by performing community detection on a host of benchmark datasets. The reported results are competitive with classical algorithms that operate on the entire graph while benefitting from a substantially reduced computational complexity due to the reduced dimensionality of the representations.
Hida-Matérn Kernel
Jul 15, 2021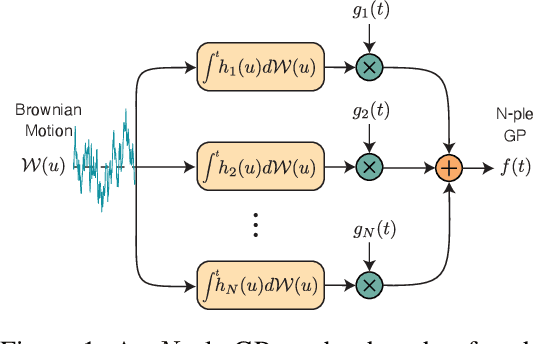
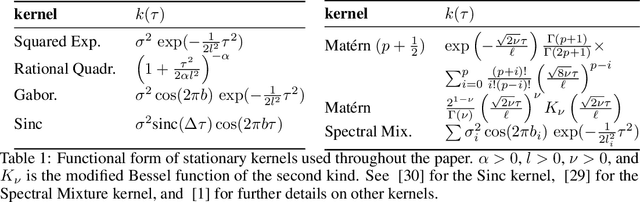

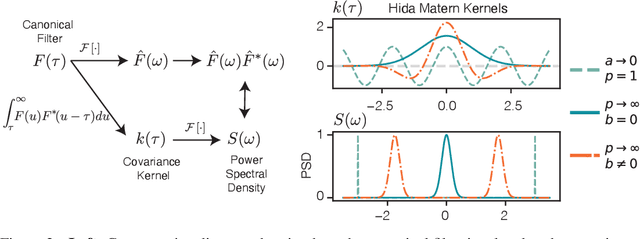
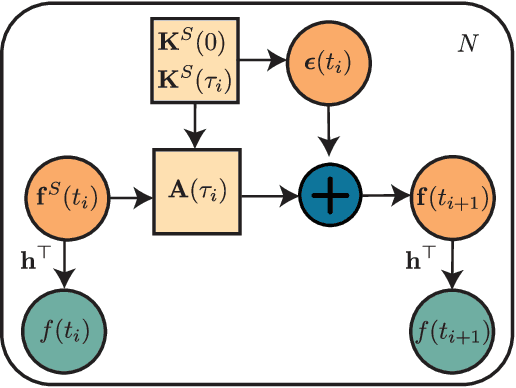
Abstract:We present the class of Hida-Mat\'ern kernels, which is the canonical family of covariance functions over the entire space of stationary Gauss-Markov Processes. It extends upon Mat\'ern kernels, by allowing for flexible construction of priors over processes with oscillatory components. Any stationary kernel, including the widely used squared-exponential and spectral mixture kernels, are either directly within this class or are appropriate asymptotic limits, demonstrating the generality of this class. Taking advantage of its Markovian nature we show how to represent such processes as state space models using only the kernel and its derivatives. In turn this allows us to perform Gaussian Process inference more efficiently and side step the usual computational burdens. We also show how exploiting special properties of the state space representation enables improved numerical stability in addition to further reductions of computational complexity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge