Paulo Afonso Ferreira
NemaNet: A convolutional neural network model for identification of nematodes soybean crop in brazil
Mar 05, 2021
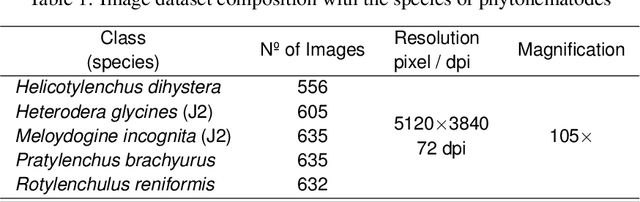
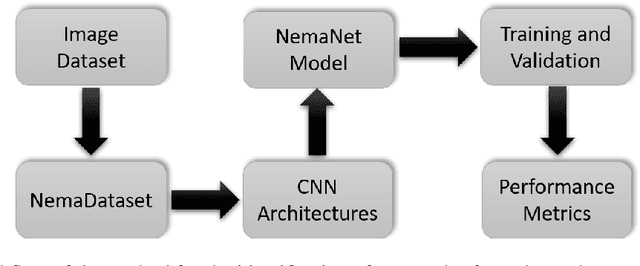
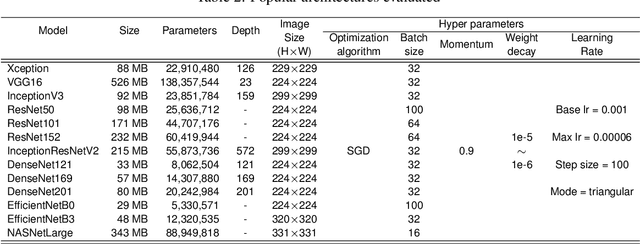
Abstract:Phytoparasitic nematodes (or phytonematodes) are causing severe damage to crops and generating large-scale economic losses worldwide. In soybean crops, annual losses are estimated at 10.6% of world production. Besides, identifying these species through microscopic analysis by an expert with taxonomy knowledge is often laborious, time-consuming, and susceptible to failure. In this perspective, robust and automatic approaches are necessary for identifying phytonematodes capable of providing correct diagnoses for the classification of species and subsidizing the taking of all control and prevention measures. This work presents a new public data set called NemaDataset containing 3,063 microscopic images from five nematode species with the most significant damage relevance for the soybean crop. Additionally, we propose a new Convolutional Neural Network (CNN) model defined as NemaNet and a comparative assessment with thirteen popular models of CNNs, all of them representing the state of the art classification and recognition. The general average calculated for each model, on a from-scratch training, the NemaNet model reached 96.99% accuracy, while the best evaluation fold reached 98.03%. In training with transfer learning, the average accuracy reached 98.88\%. The best evaluation fold reached 99.34% and achieve an overall accuracy improvement over 6.83% and 4.1%, for from-scratch and transfer learning training, respectively, when compared to other popular models.
Plant Diseases recognition on images using Convolutional Neural Networks: A Systematic Review
Sep 09, 2020
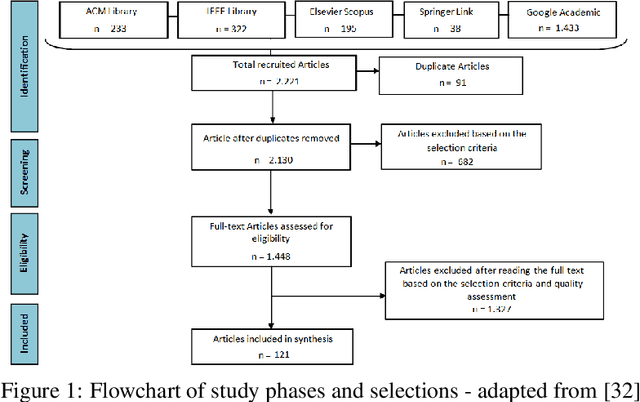
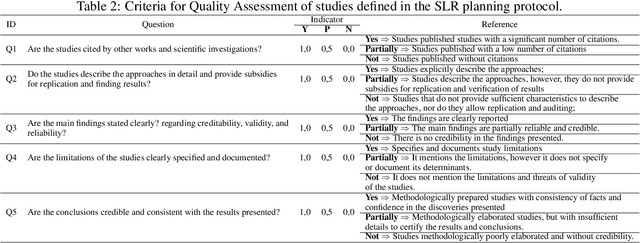
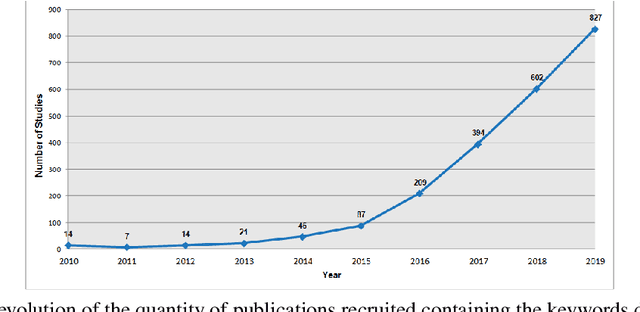
Abstract:Plant diseases are considered one of the main factors influencing food production and minimize losses in production, and it is essential that crop diseases have fast detection and recognition. The recent expansion of deep learning methods has found its application in plant disease detection, offering a robust tool with highly accurate results. In this context, this work presents a systematic review of the literature that aims to identify the state of the art of the use of convolutional neural networks(CNN) in the process of identification and classification of plant diseases, delimiting trends, and indicating gaps. In this sense, we present 121 papers selected in the last ten years with different approaches to treat aspects related to disease detection, characteristics of the data set, the crops and pathogens investigated. From the results of the systematic review, it is possible to understand the innovative trends regarding the use of CNNs in the identification of plant diseases and to identify the gaps that need the attention of the research community.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge