Patrick Virtue
Joint Embedding and Classification for SAR Target Recognition
Dec 16, 2017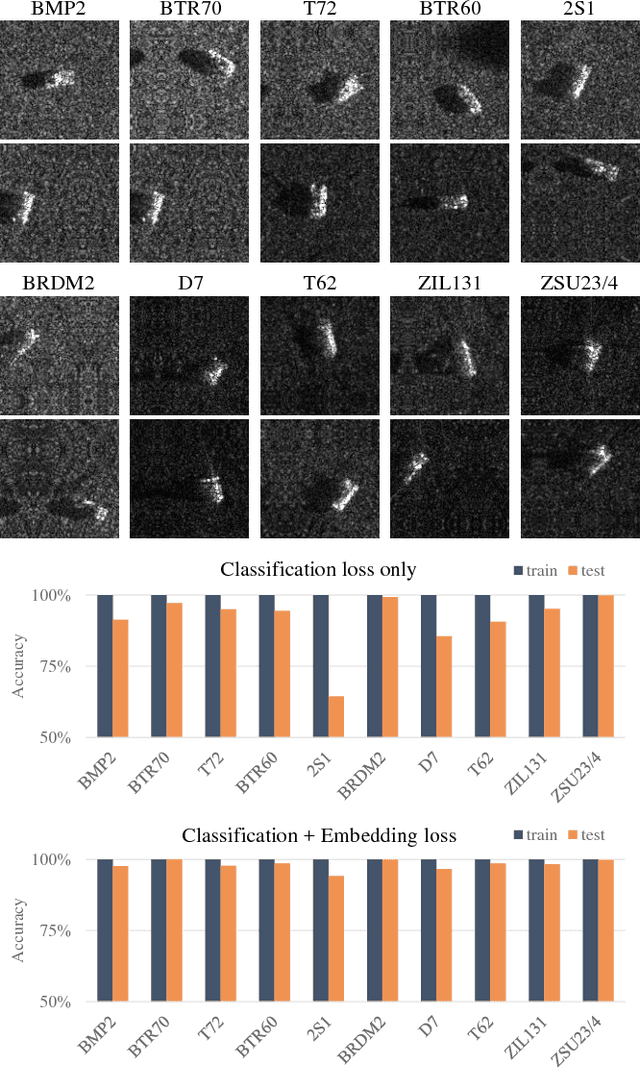
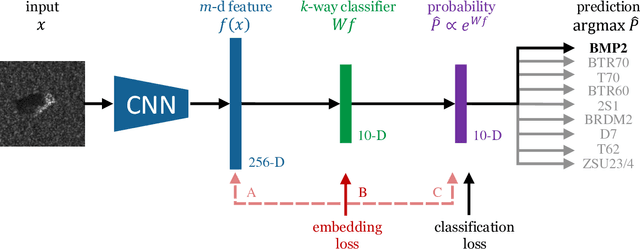
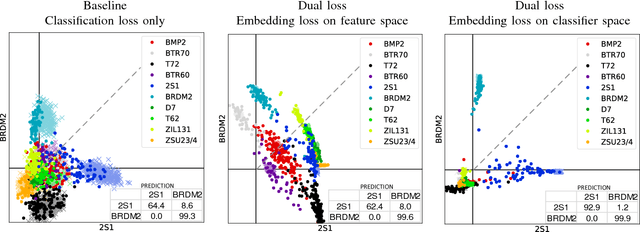
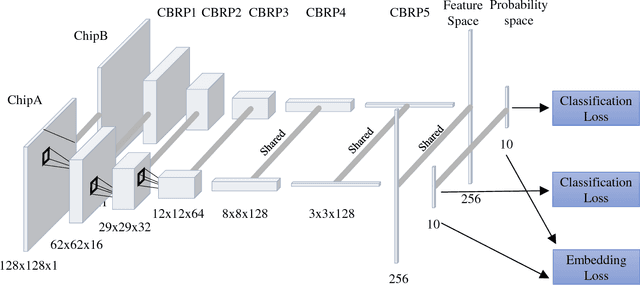
Abstract:Deep learning can be an effective and efficient means to automatically detect and classify targets in synthetic aperture radar (SAR) images, but it is critical for trained neural networks to be robust to variations that exist between training and test environments. The layers in a neural network can be understood as successive transformations of an input image into embedded feature representations and ultimately into a semantic class label. To address the overfitting problem in SAR target classification, we train neural networks to optimize the spatial clustering of points in the embedded space in addition to optimizing the final classification score. We demonstrate that networks trained with this dual embedding and classification loss outperform networks with classification loss only. We study placing the embedding loss after different network layers and find that applying the embedding loss on the classification space results in the best SAR classification performance. Finally, our visualization of the network's ten-dimensional classification space supports our claim that the embedding loss encourages greater separation between target class clusters for both training and testing partitions of the MSTAR dataset.
Better than Real: Complex-valued Neural Nets for MRI Fingerprinting
Jul 01, 2017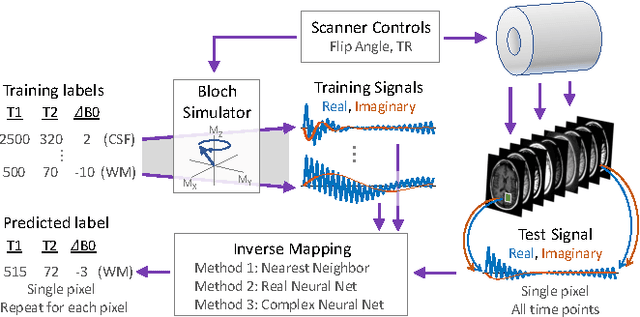
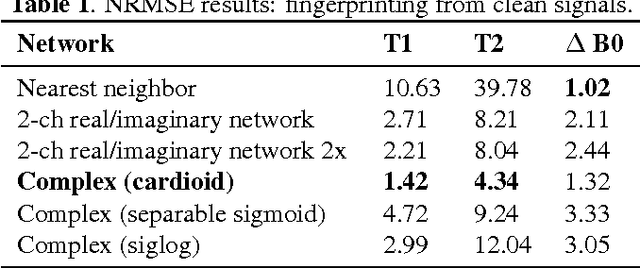
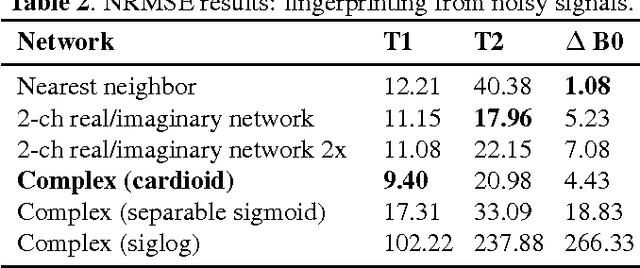

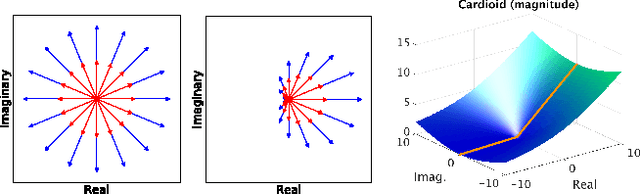

Abstract:The task of MRI fingerprinting is to identify tissue parameters from complex-valued MRI signals. The prevalent approach is dictionary based, where a test MRI signal is compared to stored MRI signals with known tissue parameters and the most similar signals and tissue parameters retrieved. Such an approach does not scale with the number of parameters and is rather slow when the tissue parameter space is large. Our first novel contribution is to use deep learning as an efficient nonlinear inverse mapping approach. We generate synthetic (tissue, MRI) data from an MRI simulator, and use them to train a deep net to map the MRI signal to the tissue parameters directly. Our second novel contribution is to develop a complex-valued neural network with new cardioid activation functions. Our results demonstrate that complex-valued neural nets could be much more accurate than real-valued neural nets at complex-valued MRI fingerprinting.
On the Empirical Effect of Gaussian Noise in Under-sampled MRI Reconstruction
Oct 03, 2016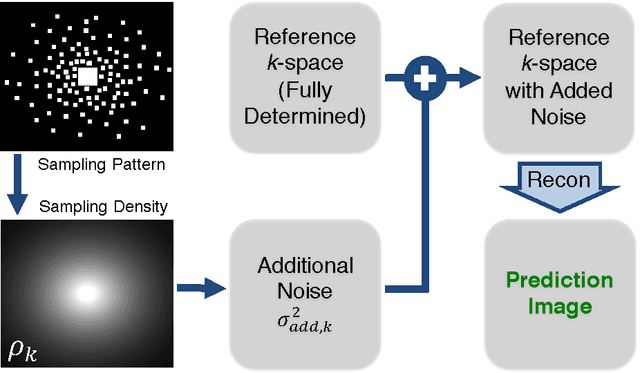
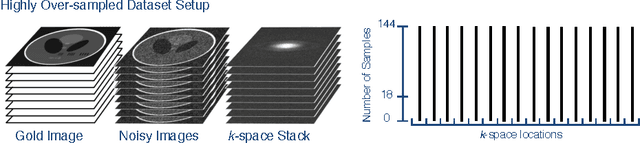
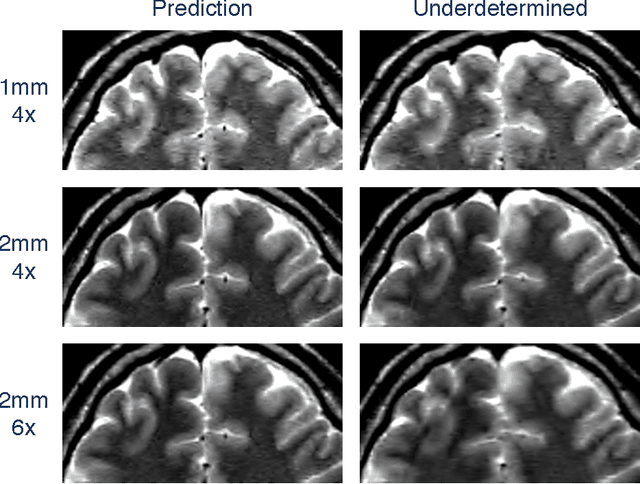
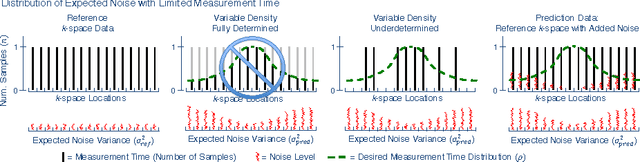

Abstract:In Fourier-based medical imaging, sampling below the Nyquist rate results in an underdetermined system, in which linear reconstructions will exhibit artifacts. Another consequence of under-sampling is lower signal to noise ratio (SNR) due to fewer acquired measurements. Even if an oracle provided the information to perfectly disambiguate the underdetermined system, the reconstructed image could still have lower image quality than a corresponding fully sampled acquisition because of the reduced measurement time. The effects of lower SNR and the underdetermined system are coupled during reconstruction, making it difficult to isolate the impact of lower SNR on image quality. To this end, we present an image quality prediction process that reconstructs fully sampled, fully determined data with noise added to simulate the loss of SNR induced by a given under-sampling pattern. The resulting prediction image empirically shows the effect of noise in under-sampled image reconstruction without any effect from an underdetermined system. We discuss how our image quality prediction process can simulate the distribution of noise for a given under-sampling pattern, including variable density sampling that produces colored noise in the measurement data. An interesting consequence of our prediction model is that we can show that recovery from underdetermined non-uniform sampling is equivalent to a weighted least squares optimization that accounts for heterogeneous noise levels across measurements. Through a series of experiments with synthetic and in vivo datasets, we demonstrate the efficacy of the image quality prediction process and show that it provides a better estimation of reconstruction image quality than the corresponding fully-sampled reference image.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge