Omid Shams Solari
\texttt{BayesBreak}: Generalized Hierarchical Bayesian Segmentation with Irregular Designs, Multi-Sample Hierarchies, and Grouped/Latent-Group Designs
Mar 16, 2026Abstract:Bayesian change-point and segmentation models provide uncertainty-aware piecewise-constant representations of ordered data, but exact inference is often tied to narrow likelihood classes, single-sequence settings, or index-uniform designs. We present \texttt{BayesBreak}, a modular offline Bayesian segmentation framework built around a simple separation: each candidate block contributes a marginal likelihood and any required moment numerators, and a global dynamic program combines those block scores into posterior quantities over segment counts, boundary locations, and latent signals. For weighted exponential-family likelihoods with conjugate priors, block evidences and posterior moments are available in closed form from cumulative sufficient statistics, yielding exact sum-product inference for $P(y\mid k)$, $P(k\mid y)$, boundary marginals, and Bayes regression curves. We also distinguish these quantities from the \emph{joint} MAP segmentation, which is recovered by a separate max-sum backtracking recursion.
BLOCCS: Block Sparse Canonical Correlation Analysis With Application To Interpretable Omics Integration
Sep 17, 2019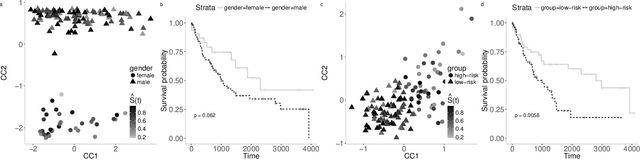
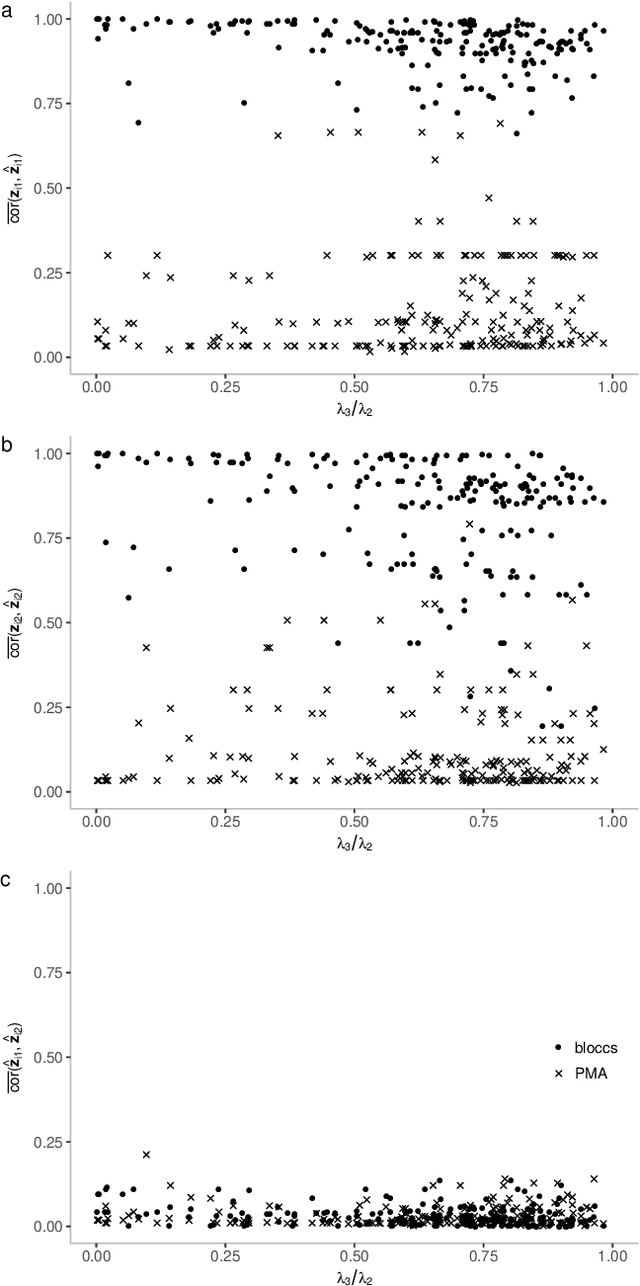
Abstract:We introduce Block Sparse Canonical Correlation Analysis which estimates multiple pairs of canonical directions (together a "block") at once, resulting in significantly improved orthogonality of the sparse directions which, we demonstrate, translates to more interpretable solutions. Our approach builds on the sparse CCA method of (Solari, Brown, and Bickel 2019) in that we also express the bi-convex objective of our block formulation as a concave minimization problem over an orthogonal k-frame in a unit Euclidean ball, which in turn, due to concavity of the objective, is shrunk to a Stiefel manifold, which is optimized via gradient descent algorithm. Our simulations show that our method outperforms existing sCCA algorithms and implementations in terms of computational cost and stability, mainly due to the drastic shrinkage of our search space, and the correlation within and orthogonality between pairs of estimated canonical covariates. Finally, we apply our method, available as an R-package called BLOCCS, to multi-omic data on Lung Squamous Cell Carcinoma(LUSC) obtained via The Cancer Genome Atlas, and demonstrate its capability in capturing meaningful biological associations relevant to the hypothesis under study rather than spurious dominant variations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge