Nicolas Papadakis
IMB
GraphXCOVID: Explainable Deep Graph Diffusion Pseudo-Labelling for Identifying COVID-19 on Chest X-rays
Sep 30, 2020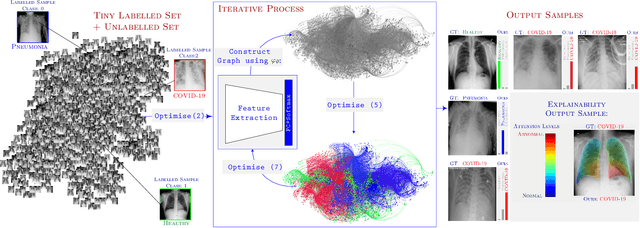
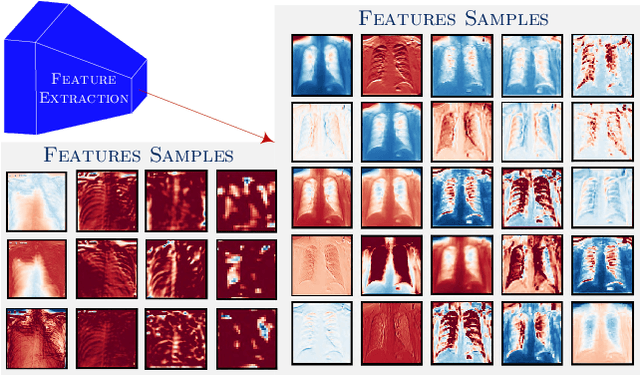
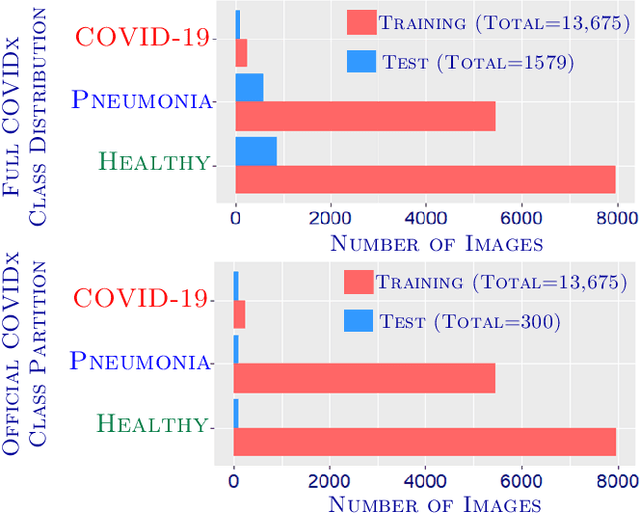
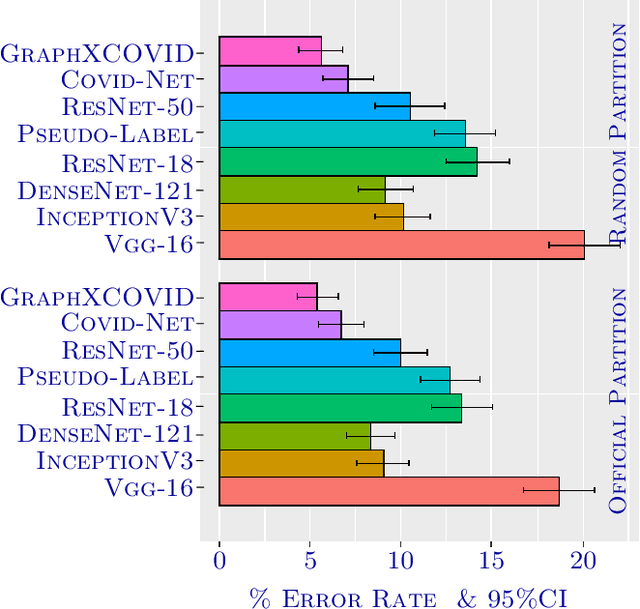
Abstract:Can one learn to diagnose COVID-19 under extreme minimal supervision? Since the outbreak of the novel COVID-19 there has been a rush for developing Artificial Intelligence techniques for expert-level disease identification on Chest X-ray data. In particular, the use of deep supervised learning has become the go-to paradigm. However, the performance of such models is heavily dependent on the availability of a large and representative labelled dataset. The creation of which is a heavily expensive and time consuming task, and especially imposes a great challenge for a novel disease. Semi-supervised learning has shown the ability to match the incredible performance of supervised models whilst requiring a small fraction of the labelled examples. This makes the semi-supervised paradigm an attractive option for identifying COVID-19. In this work, we introduce a graph based deep semi-supervised framework for classifying COVID-19 from chest X-rays. Our framework introduces an optimisation model for graph diffusion that reinforces the natural relation among the tiny labelled set and the vast unlabelled data. We then connect the diffusion prediction output as pseudo-labels that are used in an iterative scheme in a deep net. We demonstrate, through our experiments, that our model is able to outperform the current leading supervised model with a tiny fraction of the labelled examples. Finally, we provide attention maps to accommodate the radiologist's mental model, better fitting their perceptual and cognitive abilities. These visualisation aims to assist the radiologist in judging whether the diagnostic is correct or not, and in consequence to accelerate the decision.
Wasserstein Generative Models for Patch-based Texture Synthesis
Jun 19, 2020
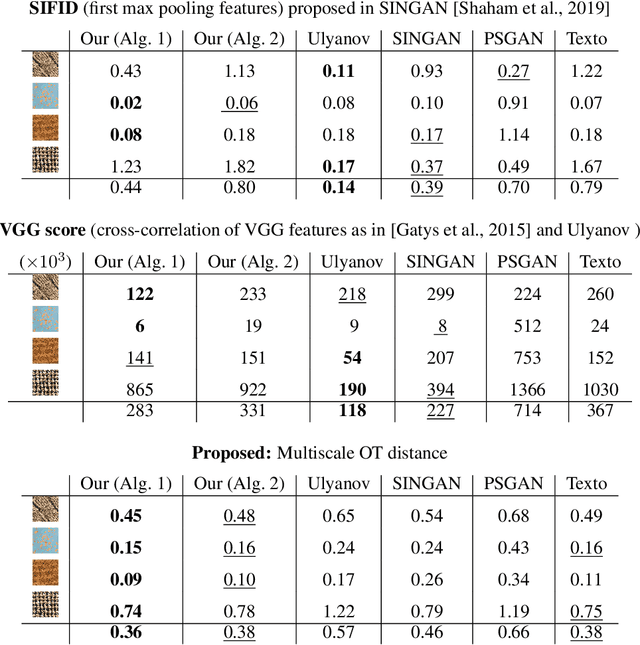

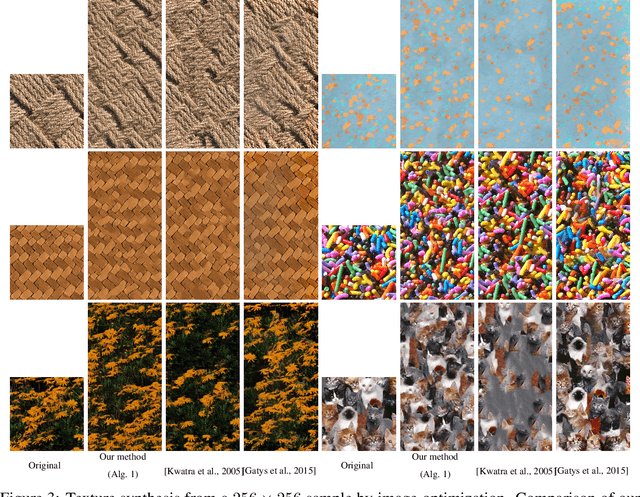
Abstract:In this paper, we propose a framework to train a generative model for texture image synthesis from a single example. To do so, we exploit the local representation of images via the space of patches, that is, square sub-images of fixed size (e.g. $4\times 4$). Our main contribution is to consider optimal transport to enforce the multiscale patch distribution of generated images, which leads to two different formulations. First, a pixel-based optimization method is proposed, relying on discrete optimal transport. We show that it is related to a well-known texture optimization framework based on iterated patch nearest-neighbor projections, while avoiding some of its shortcomings. Second, in a semi-discrete setting, we exploit the differential properties of Wasserstein distances to learn a fully convolutional network for texture generation. Once estimated, this network produces realistic and arbitrarily large texture samples in real time. The two formulations result in non-convex concave problems that can be optimized efficiently with convergence properties and improved stability compared to adversarial approaches, without relying on any regularization. By directly dealing with the patch distribution of synthesized images, we also overcome limitations of state-of-the art techniques, such as patch aggregation issues that usually lead to low frequency artifacts (e.g. blurring) in traditional patch-based approaches, or statistical inconsistencies (e.g. color or patterns) in learning approaches.
Multi-task deep learning for image segmentation using recursive approximation tasks
May 26, 2020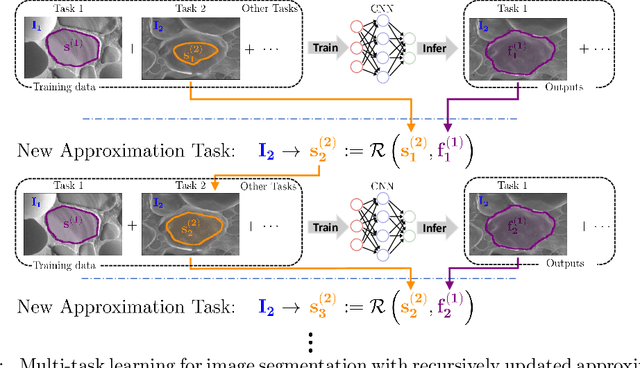
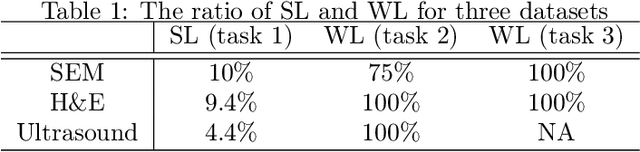
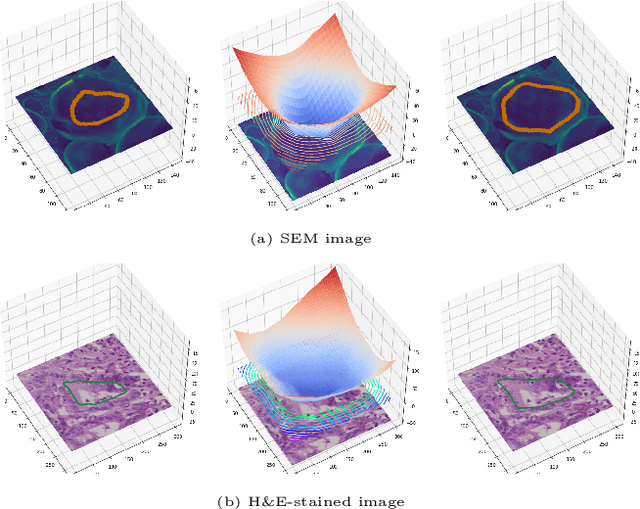

Abstract:Fully supervised deep neural networks for segmentation usually require a massive amount of pixel-level labels which are manually expensive to create. In this work, we develop a multi-task learning method to relax this constraint. We regard the segmentation problem as a sequence of approximation subproblems that are recursively defined and in increasing levels of approximation accuracy. The subproblems are handled by a framework that consists of 1) a segmentation task that learns from pixel-level ground truth segmentation masks of a small fraction of the images, 2) a recursive approximation task that conducts partial object regions learning and data-driven mask evolution starting from partial masks of each object instance, and 3) other problem oriented auxiliary tasks that are trained with sparse annotations and promote the learning of dedicated features. Most training images are only labeled by (rough) partial masks, which do not contain exact object boundaries, rather than by their full segmentation masks. During the training phase, the approximation task learns the statistics of these partial masks, and the partial regions are recursively increased towards object boundaries aided by the learned information from the segmentation task in a fully data-driven fashion. The network is trained on an extremely small amount of precisely segmented images and a large set of coarse labels. Annotations can thus be obtained in a cheap way. We demonstrate the efficiency of our approach in three applications with microscopy images and ultrasound images.
Variational Osmosis for Non-linear Image Fusion
Oct 04, 2019



Abstract:We propose a new variational model for nonlinear image fusion. Our approach incorporates the osmosis model proposed in Vogel et al. (2013) and Weickert et al. (2013) as an energy term in a variational model. The osmosis energy is known to realize visually plausible image data fusion. As a consequence, our method is invariant to multiplicative brightness changes. On the practical side, it requires minimal supervision and parameter tuning and can encode prior information on the structure of the images to be fused. We develop a primal-dual algorithm for solving this new image fusion model and we apply the resulting minimisation scheme to multi-modal image fusion for face fusion, colour transfer and some cultural heritage conservation challenges. Visual comparison to state-of-the-art proves the quality and flexibility of our method.
GraphX$^{NET}-$ Chest X-Ray Classification Under Extreme Minimal Supervision
Jul 25, 2019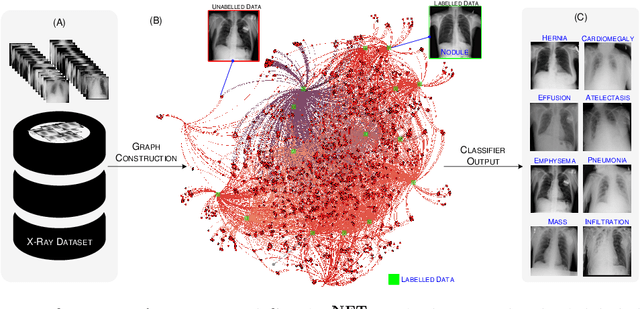
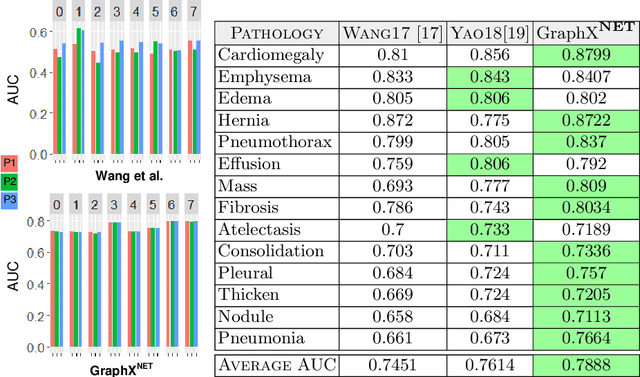
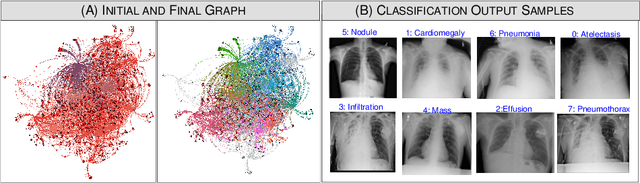
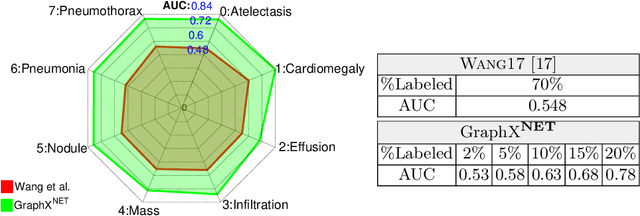
Abstract:The task of classifying X-ray data is a problem of both theoretical and clinical interest. Whilst supervised deep learning methods rely upon huge amounts of labelled data, the critical problem of achieving a good classification accuracy when an extremely small amount of labelled data is available has yet to be tackled. In this work, we introduce a novel semi-supervised framework for X-ray classification which is based on a graph-based optimisation model. To the best of our knowledge, this is the first method that exploits graph-based semi-supervised learning for X-ray data classification. Furthermore, we introduce a new multi-class classification functional with carefully selected class priors which allows for a smooth solution that strengthens the synergy between the limited number of labels and the huge amount of unlabelled data. We demonstrate, through a set of numerical and visual experiments, that our method produces highly competitive results on the ChestX-ray14 data set whilst drastically reducing the need for annotated data.
A multi-task U-net for segmentation with lazy labels
Jun 20, 2019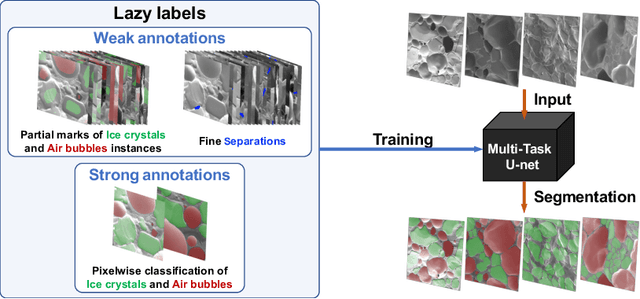
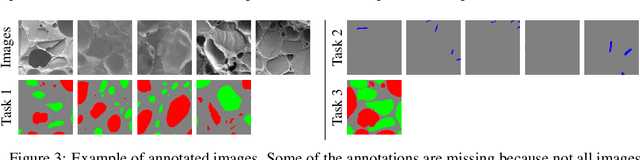
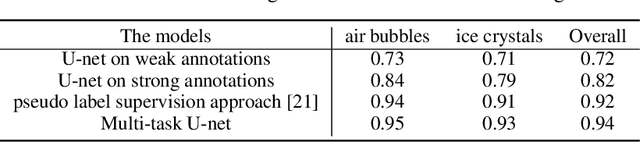
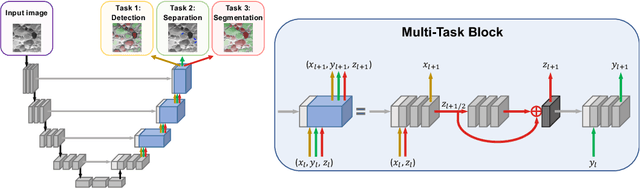

Abstract:The need for labour intensive pixel-wise annotation is a major limitation of many fully supervised learning methods for image segmentation. In this paper, we propose a deep convolutional neural network for multi-class segmentation that circumvents this problem by being trainable on coarse data labels combined with only a very small number of images with pixel-wise annotations. We call this new labelling strategy 'lazy' labels. Image segmentation is then stratified into three connected tasks: rough detection of class instances, separation of wrongly connected objects without a clear boundary, and pixel-wise segmentation to find the accurate boundaries of each object. These problems are integrated into a multitask learning framework and the model is trained end-to-end in a semi-supervised fashion. The method is applied on a dataset of food microscopy images. We show that the model gives accurate segmentation results even if exact boundary labels are missing for a majority of the annotated data. This allows more flexibility and efficiency for training deep neural networks that are data hungry in a practical setting where manual annotation is expensive, by collecting more lazy (rough) annotations than precisely segmented images.
Beyond Supervised Classification: Extreme Minimal Supervision with the Graph 1-Laplacian
Jun 20, 2019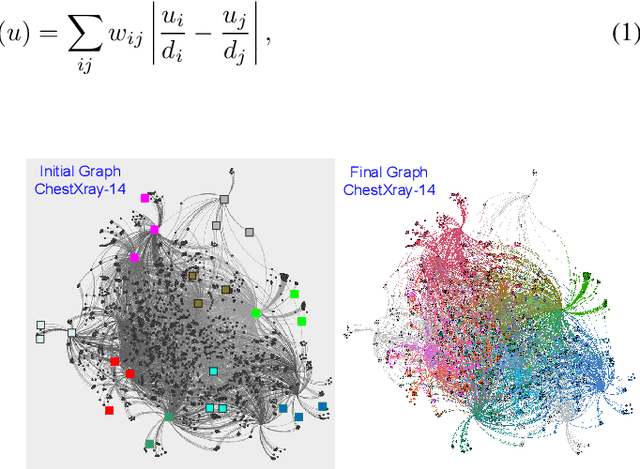
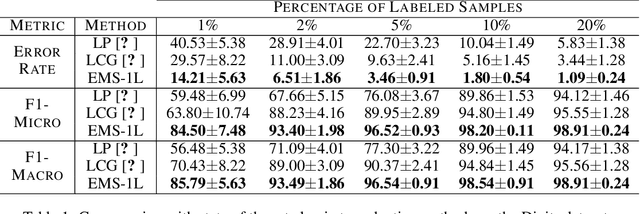
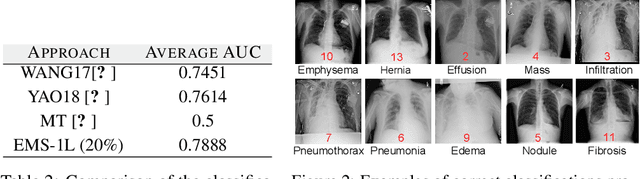
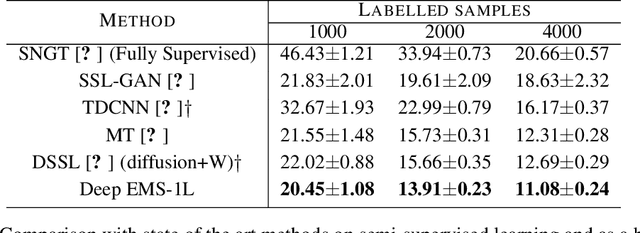
Abstract:We consider the task of classifying when an extremely reduced amount of labelled data is available. This problem is of a great interest, in several real-world problems, as obtaining large amounts of labelled data is expensive and time consuming. We present a novel semi-supervised framework for multi-class classification that is based on the normalised and non-smooth graph 1-Laplacian. Our transductive framework is framed under a novel functional with carefully selected class priors - that enforces a sufficiently smooth solution that strengthens the intrinsic relation between the labelled and unlabelled data. We demonstrate through extensive experimental results on large datasets CIFAR-10 and ChestX-ray14, that our method outperforms classic methods and readily competes with recent deep-learning approaches.
Robust superpixels using color and contour features along linear path
Mar 17, 2019
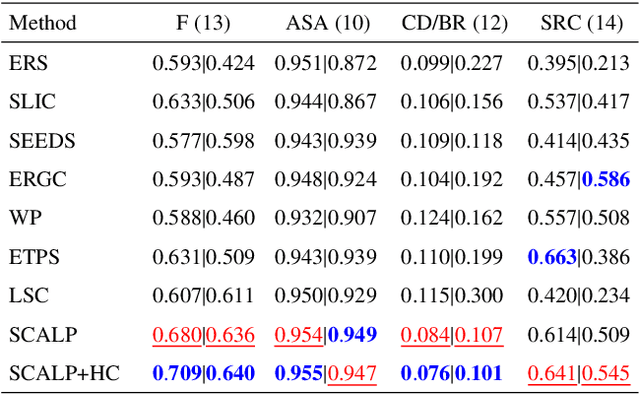
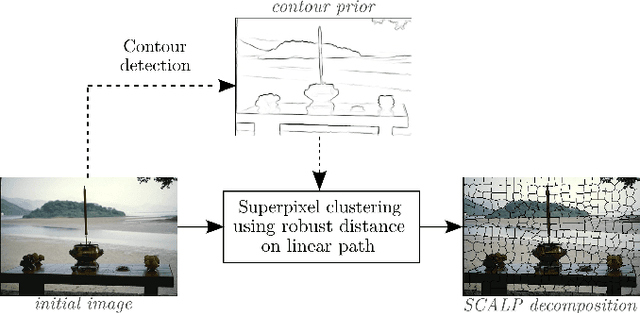
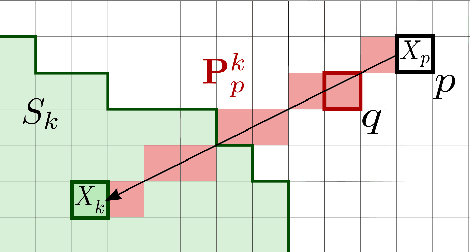
Abstract:Superpixel decomposition methods are widely used in computer vision and image processing applications. By grouping homogeneous pixels, the accuracy can be increased and the decrease of the number of elements to process can drastically reduce the computational burden. For most superpixel methods, a trade-off is computed between 1) color homogeneity, 2) adherence to the image contours and 3) shape regularity of the decomposition. In this paper, we propose a framework that jointly enforces all these aspects and provides accurate and regular Superpixels with Contour Adherence using Linear Path (SCALP). During the decomposition, we propose to consider color features along the linear path between the pixel and the corresponding superpixel barycenter. A contour prior is also used to prevent the crossing of image boundaries when associating a pixel to a superpixel. Finally, in order to improve the decomposition accuracy and the robustness to noise, we propose to integrate the pixel neighborhood information, while preserving the same computational complexity. SCALP is extensively evaluated on standard segmentation dataset, and the obtained results outperform the ones of the state-of-the-art methods. SCALP is also extended for supervoxel decomposition on MRI images.
SuperPatchMatch: an Algorithm for Robust Correspondences using Superpixel Patches
Mar 17, 2019
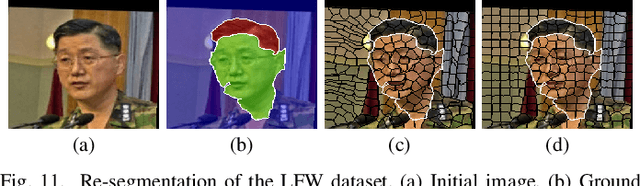
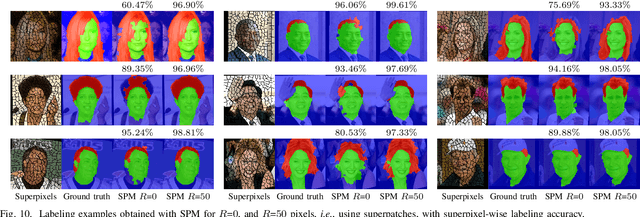

Abstract:Superpixels have become very popular in many computer vision applications. Nevertheless, they remain underexploited since the superpixel decomposition may produce irregular and non stable segmentation results due to the dependency to the image content. In this paper, we first introduce a novel structure, a superpixel-based patch, called SuperPatch. The proposed structure, based on superpixel neighborhood, leads to a robust descriptor since spatial information is naturally included. The generalization of the PatchMatch method to SuperPatches, named SuperPatchMatch, is introduced. Finally, we propose a framework to perform fast segmentation and labeling from an image database, and demonstrate the potential of our approach since we outperform, in terms of computational cost and accuracy, the results of state-of-the-art methods on both face labeling and medical image segmentation.
An Optimized PatchMatch for Multi-scale and Multi-feature Label Fusion
Mar 17, 2019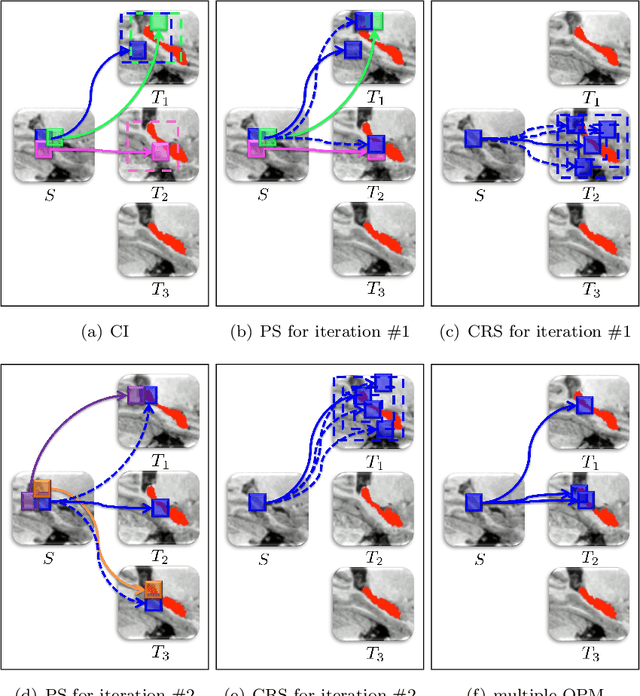
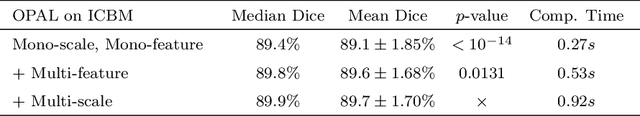
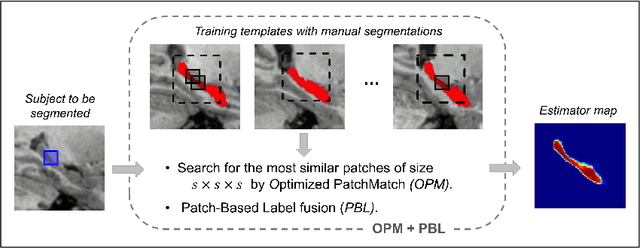
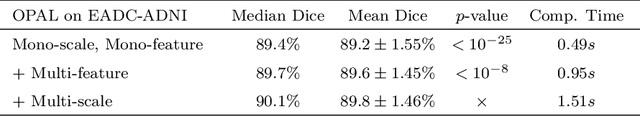
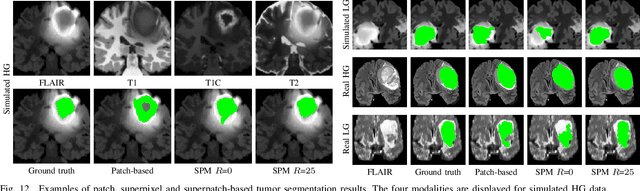
Abstract:Automatic segmentation methods are important tools for quantitative analysis of Magnetic Resonance Images (MRI). Recently, patch-based label fusion approaches have demonstrated state-of-the-art segmentation accuracy. In this paper, we introduce a new patch-based label fusion framework to perform segmentation of anatomical structures. The proposed approach uses an Optimized PAtchMatch Label fusion (OPAL) strategy that drastically reduces the computation time required for the search of similar patches. The reduced computation time of OPAL opens the way for new strategies and facilitates processing on large databases. In this paper, we investigate new perspectives offered by OPAL, by introducing a new multi-scale and multi-feature framework. During our validation on hippocampus segmentation we use two datasets: young adults in the ICBM cohort and elderly adults in the EADC-ADNI dataset. For both, OPAL is compared to state-of-the-art methods. Results show that OPAL obtained the highest median Dice coefficient (89.9% for ICBM and 90.1% for EADC-ADNI). Moreover, in both cases, OPAL produced a segmentation accuracy similar to inter-expert variability. On the EADC-ADNI dataset, we compare the hippocampal volumes obtained by manual and automatic segmentation. The volumes appear to be highly correlated that enables to perform more accurate separation of pathological populations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge