Mrutyunjaya Panda
Utkal University, Vani Vihar, Bhubaneswar, India
Explainable artificial intelligence for Healthcare applications using Random Forest Classifier with LIME and SHAP
Nov 09, 2023Abstract:With the advances in computationally efficient artificial Intelligence (AI) techniques and their numerous applications in our everyday life, there is a pressing need to understand the computational details hidden in black box AI techniques such as most popular machine learning and deep learning techniques; through more detailed explanations. The origin of explainable AI (xAI) is coined from these challenges and recently gained more attention by the researchers by adding explainability comprehensively in traditional AI systems. This leads to develop an appropriate framework for successful applications of xAI in real life scenarios with respect to innovations, risk mitigation, ethical issues and logical values to the users. In this book chapter, an in-depth analysis of several xAI frameworks and methods including LIME (Local Interpretable Model-agnostic Explanations) and SHAP (SHapley Additive exPlanations) are provided. Random Forest Classifier as black box AI is used on a publicly available Diabetes symptoms dataset with LIME and SHAP for better interpretations. The results obtained are interesting in terms of transparency, valid and trustworthiness in diabetes disease prediction.
Elephant Search with Deep Learning for Microarray Data Analysis
Jul 12, 2017



Abstract:Even though there is a plethora of research in Microarray gene expression data analysis, still, it poses challenges for researchers to effectively and efficiently analyze the large yet complex expression of genes. The feature (gene) selection method is of paramount importance for understanding the differences in biological and non-biological variation between samples. In order to address this problem, a novel elephant search (ES) based optimization is proposed to select best gene expressions from the large volume of microarray data. Further, a promising machine learning method is envisioned to leverage such high dimensional and complex microarray dataset for extracting hidden patterns inside to make a meaningful prediction and most accurate classification. In particular, stochastic gradient descent based Deep learning (DL) with softmax activation function is then used on the reduced features (genes) for better classification of different samples according to their gene expression levels. The experiments are carried out on nine most popular Cancer microarray gene selection datasets, obtained from UCI machine learning repository. The empirical results obtained by the proposed elephant search based deep learning (ESDL) approach are compared with most recent published article for its suitability in future Bioinformatics research.
Big Models for Big Data using Multi objective averaged one dependence estimators
Oct 25, 2016



Abstract:Even though, many researchers tried to explore the various possibilities on multi objective feature selection, still it is yet to be explored with best of its capabilities in data mining applications rather than going for developing new ones. In this paper, multi-objective evolutionary algorithm ENORA is used to select the features in a multi-class classification problem. The fusion of AnDE (averaged n-dependence estimators) with n=1, a variant of naive Bayes with efficient feature selection by ENORA is performed in order to obtain a fast hybrid classifier which can effectively learn from big data. This method aims at solving the problem of finding optimal feature subset from full data which at present still remains to be a difficult problem. The efficacy of the obtained classifier is extensively evaluated with a range of most popular 21 real world dataset, ranging from small to big. The results obtained are encouraging in terms of time, Root mean square error, zero-one loss and classification accuracy.
Towards the effectiveness of Deep Convolutional Neural Network based Fast Random Forest Classifier
Sep 28, 2016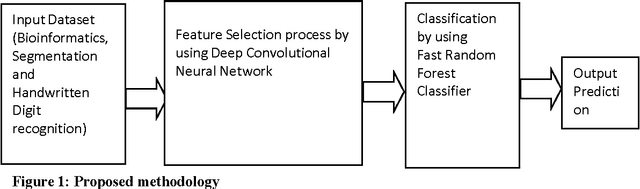
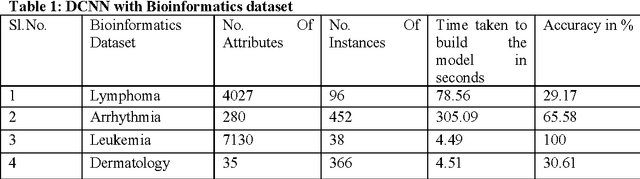
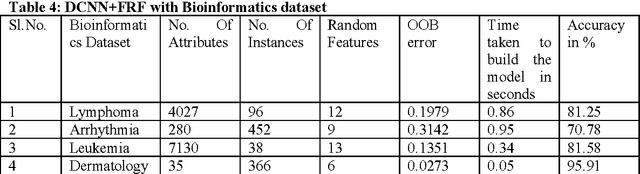
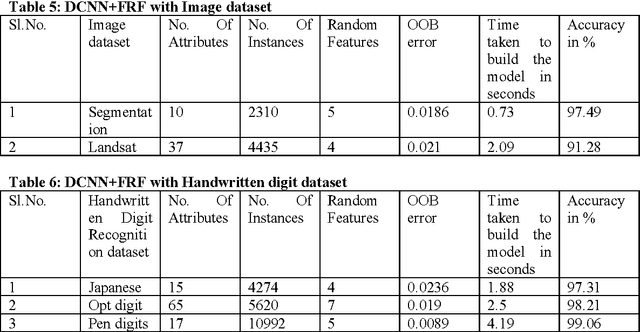
Abstract:Deep Learning is considered to be a quite young in the area of machine learning research, found its effectiveness in dealing complex yet high dimensional dataset that includes but limited to images, text and speech etc. with multiple levels of representation and abstraction. As there are a plethora of research on these datasets by various researchers , a win over them needs lots of attention. Careful setting of Deep learning parameters is of paramount importance in order to avoid the overfitting unlike conventional methods with limited parameter settings. Deep Convolutional neural network (DCNN) with multiple layers of compositions and appropriate settings might be is an efficient machine learning method that can outperform the conventional methods in a great way. However, due to its slow adoption in learning, there are also always a chance of overfitting during feature selection process, which can be addressed by employing a regularization method called dropout. Fast Random Forest (FRF) is a powerful ensemble classifier especially when the datasets are noisy and when the number of attributes is large in comparison to the number of instances, as is the case of Bioinformatics datasets. Several publicly available Bioinformatics dataset, Handwritten digits recognition and Image segmentation dataset are considered for evaluation of the proposed approach. The excellent performance obtained by the proposed DCNN based feature selection with FRF classifier on high dimensional datasets makes it a fast and accurate classifier in comparison the state-of-the-art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge