Milo Roucairol
Solving the HP model with Nested Monte Carlo Search
Jan 25, 2023Abstract:In this paper we present a new Monte Carlo Search (MCS) algorithm for finding the ground state energy of proteins in the HP-model. We also compare it briefly to other MCS algorithms not usually used on the HP-model and provide an overview of the algorithms used on HP-model. The algorithm presented in this paper does not beat state of the art algorithms, see PERM (Hsu and Grassberger 2011), REMC (Thachuk, Shmygelska, and Hoos 2007) or WLRE (W\"ust and Landau 2012) for better results. Hsu, H.-P.; and Grassberger, P. 2011. A review of Monte Carlo simulations of polymers with PERM. Journal of Statistical Physics, 144 (3): 597 to 637. Thachuk, C.; Shmygelska, A.; and Hoos, H. H. 2007. A replica exchange Monte Carlo algorithm for protein folding in the HP model. BMC Bioinformatics, 8(1): 342. W\"ust, T.; and Landau, D. P. 2012. Optimized Wang-Landau sampling of lattice polymers: Ground state search and folding thermodynamics of HP model proteins. The Journal of Chemical Physics, 137(6): 064903.
Refutation of Spectral Graph Theory Conjectures with Monte Carlo Search
Jul 04, 2022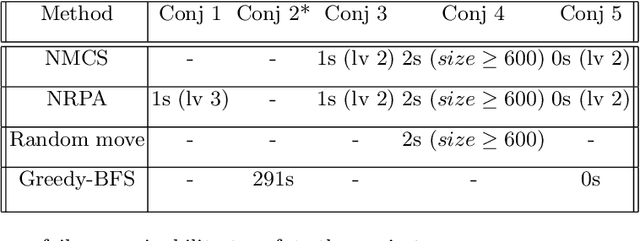
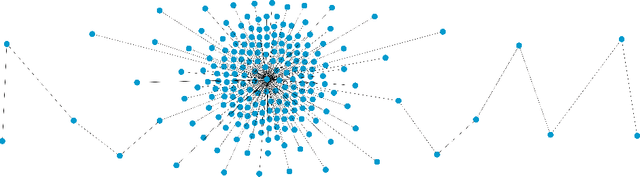
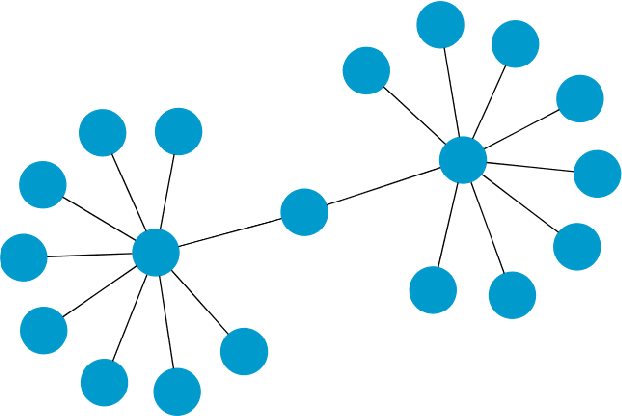
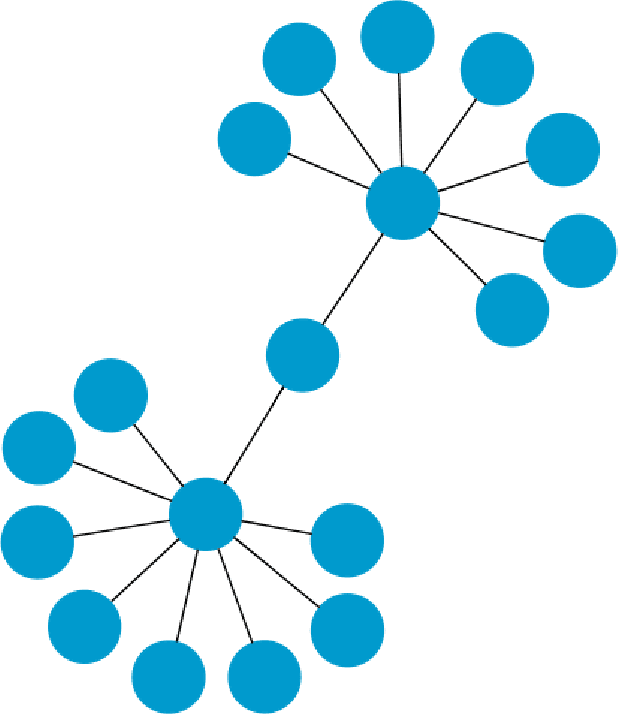

Abstract:We demonstrate how Monte Carlo Search (MCS) algorithms, namely Nested Monte Carlo Search (NMCS) and Nested Rollout Policy Adaptation (NRPA), can be used to build graphs and find counter-examples to spectral graph theory conjectures in minutes. We also refute a conjecture attributed to Peter Shor that was left open.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge