Mihaela van der Schaar
Identifiable Energy-based Representations: An Application to Estimating Heterogeneous Causal Effects
Aug 06, 2021
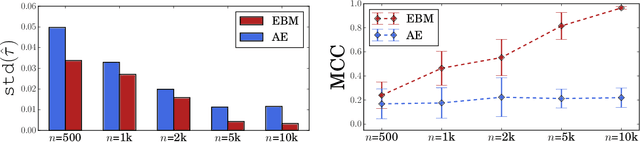
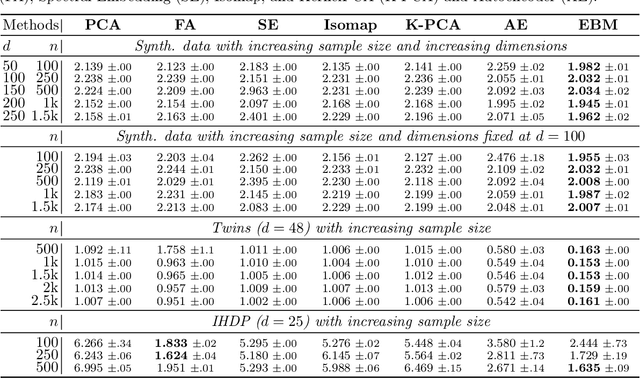
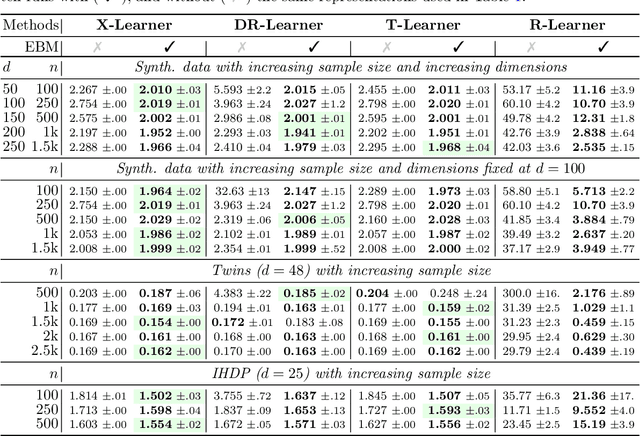
Abstract:Conditional average treatment effects (CATEs) allow us to understand the effect heterogeneity across a large population of individuals. However, typical CATE learners assume all confounding variables are measured in order for the CATE to be identifiable. Often, this requirement is satisfied by simply collecting many variables, at the expense of increased sample complexity for estimating CATEs. To combat this, we propose an energy-based model (EBM) that learns a low-dimensional representation of the variables by employing a noise contrastive loss function. With our EBM we introduce a preprocessing step that alleviates the dimensionality curse for any existing model and learner developed for estimating CATE. We prove that our EBM keeps the representations partially identifiable up to some universal constant, as well as having universal approximation capability to avoid excessive information loss from model misspecification; these properties combined with our loss function, enable the representations to converge and keep the CATE estimation consistent. Experiments demonstrate the convergence of the representations, as well as show that estimating CATEs on our representations performs better than on the variables or the representations obtained via various benchmark dimensionality reduction methods.
Doing Great at Estimating CATE? On the Neglected Assumptions in Benchmark Comparisons of Treatment Effect Estimators
Jul 28, 2021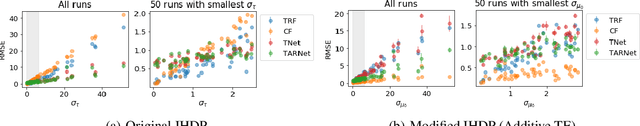
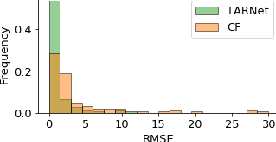
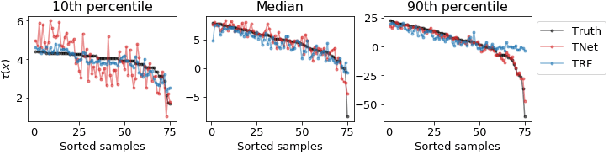
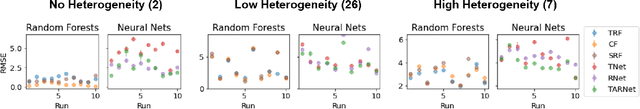
Abstract:The machine learning toolbox for estimation of heterogeneous treatment effects from observational data is expanding rapidly, yet many of its algorithms have been evaluated only on a very limited set of semi-synthetic benchmark datasets. In this paper, we show that even in arguably the simplest setting -- estimation under ignorability assumptions -- the results of such empirical evaluations can be misleading if (i) the assumptions underlying the data-generating mechanisms in benchmark datasets and (ii) their interplay with baseline algorithms are inadequately discussed. We consider two popular machine learning benchmark datasets for evaluation of heterogeneous treatment effect estimators -- the IHDP and ACIC2016 datasets -- in detail. We identify problems with their current use and highlight that the inherent characteristics of the benchmark datasets favor some algorithms over others -- a fact that is rarely acknowledged but of immense relevance for interpretation of empirical results. We close by discussing implications and possible next steps.
Inverse Contextual Bandits: Learning How Behavior Evolves over Time
Jul 13, 2021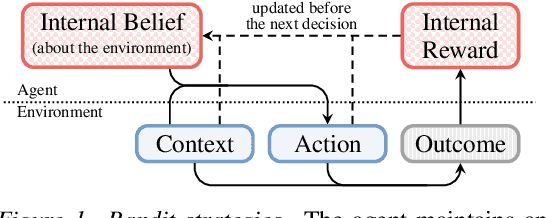
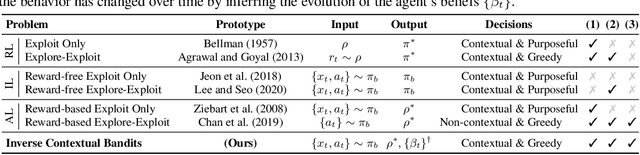
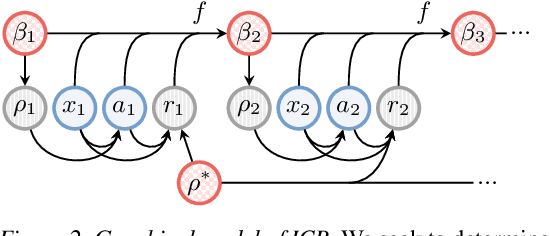
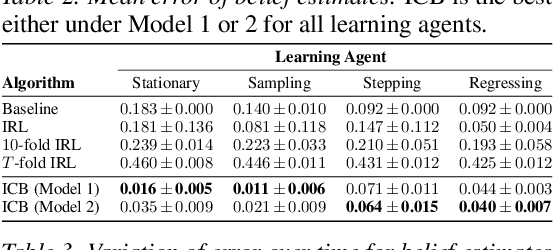
Abstract:Understanding an agent's priorities by observing their behavior is critical for transparency and accountability in decision processes, such as in healthcare. While conventional approaches to policy learning almost invariably assume stationarity in behavior, this is hardly true in practice: Medical practice is constantly evolving, and clinical professionals are constantly fine-tuning their priorities. We desire an approach to policy learning that provides (1) interpretable representations of decision-making, accounts for (2) non-stationarity in behavior, as well as operating in an (3) offline manner. First, we model the behavior of learning agents in terms of contextual bandits, and formalize the problem of inverse contextual bandits (ICB). Second, we propose two algorithms to tackle ICB, each making varying degrees of assumptions regarding the agent's learning strategy. Finally, through both real and simulated data for liver transplantations, we illustrate the applicability and explainability of our method, as well as validating its accuracy.
Integrating Expert ODEs into Neural ODEs: Pharmacology and Disease Progression
Jun 17, 2021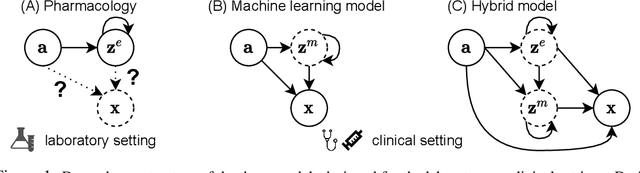
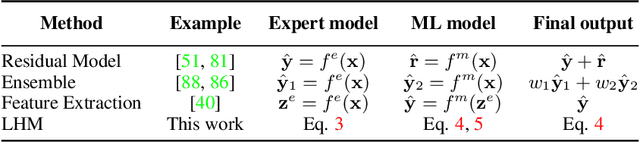
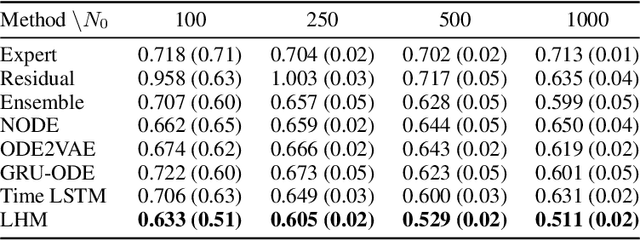
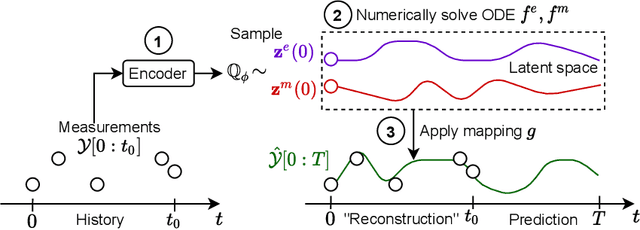

Abstract:Modeling a system's temporal behaviour in reaction to external stimuli is a fundamental problem in many areas. Pure Machine Learning (ML) approaches often fail in the small sample regime and cannot provide actionable insights beyond predictions. A promising modification has been to incorporate expert domain knowledge into ML models. The application we consider is predicting the progression of disease under medications, where a plethora of domain knowledge is available from pharmacology. Pharmacological models describe the dynamics of carefully-chosen medically meaningful variables in terms of systems of Ordinary Differential Equations (ODEs). However, these models only describe a limited collection of variables, and these variables are often not observable in clinical environments. To close this gap, we propose the latent hybridisation model (LHM) that integrates a system of expert-designed ODEs with machine-learned Neural ODEs to fully describe the dynamics of the system and to link the expert and latent variables to observable quantities. We evaluated LHM on synthetic data as well as real-world intensive care data of COVID-19 patients. LHM consistently outperforms previous works, especially when few training samples are available such as at the beginning of the pandemic.
Explaining Time Series Predictions with Dynamic Masks
Jun 09, 2021
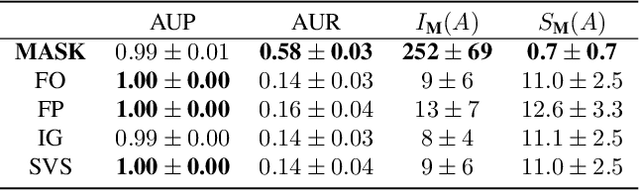

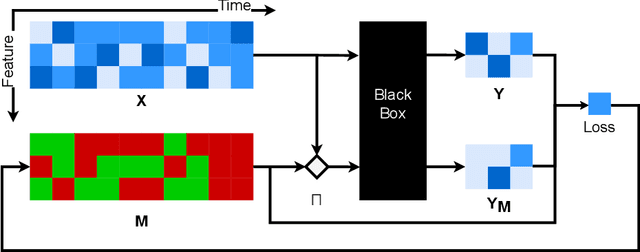
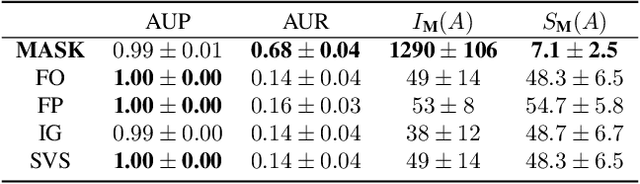
Abstract:How can we explain the predictions of a machine learning model? When the data is structured as a multivariate time series, this question induces additional difficulties such as the necessity for the explanation to embody the time dependency and the large number of inputs. To address these challenges, we propose dynamic masks (Dynamask). This method produces instance-wise importance scores for each feature at each time step by fitting a perturbation mask to the input sequence. In order to incorporate the time dependency of the data, Dynamask studies the effects of dynamic perturbation operators. In order to tackle the large number of inputs, we propose a scheme to make the feature selection parsimonious (to select no more feature than necessary) and legible (a notion that we detail by making a parallel with information theory). With synthetic and real-world data, we demonstrate that the dynamic underpinning of Dynamask, together with its parsimony, offer a neat improvement in the identification of feature importance over time. The modularity of Dynamask makes it ideal as a plug-in to increase the transparency of a wide range of machine learning models in areas such as medicine and finance, where time series are abundant.
The Medkit-Learn(ing) Environment: Medical Decision Modelling through Simulation
Jun 08, 2021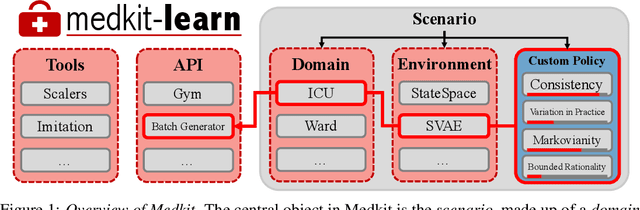
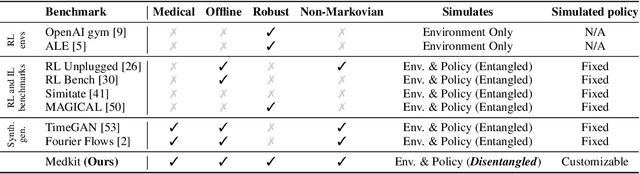
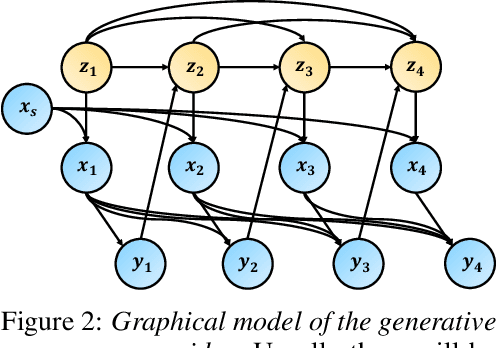
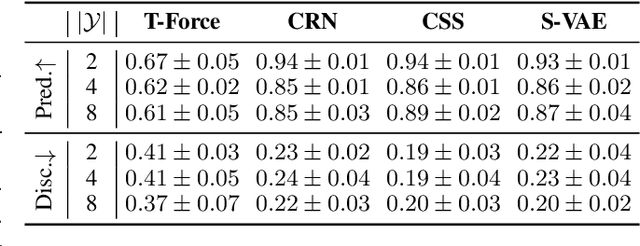
Abstract:Understanding decision-making in clinical environments is of paramount importance if we are to bring the strengths of machine learning to ultimately improve patient outcomes. Several factors including the availability of public data, the intrinsically offline nature of the problem, and the complexity of human decision making, has meant that the mainstream development of algorithms is often geared towards optimal performance in tasks that do not necessarily translate well into the medical regime; often overlooking more niche issues commonly associated with the area. We therefore present a new benchmarking suite designed specifically for medical sequential decision making: the Medkit-Learn(ing) Environment, a publicly available Python package providing simple and easy access to high-fidelity synthetic medical data. While providing a standardised way to compare algorithms in a realistic medical setting we employ a generating process that disentangles the policy and environment dynamics to allow for a range of customisations, thus enabling systematic evaluation of algorithms' robustness against specific challenges prevalent in healthcare.
On Inductive Biases for Heterogeneous Treatment Effect Estimation
Jun 07, 2021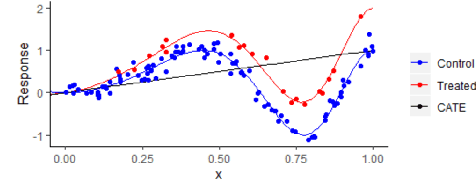
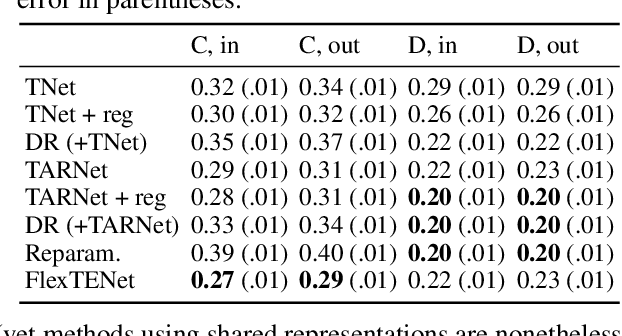
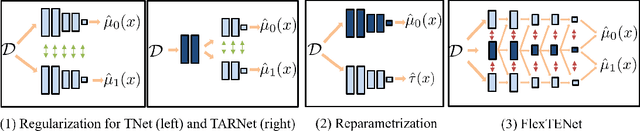
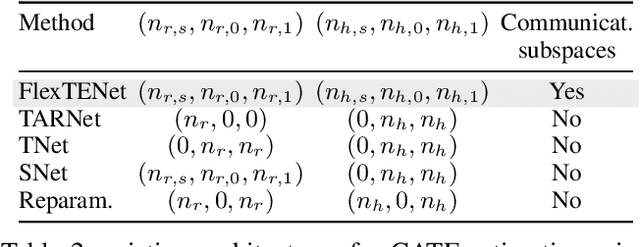
Abstract:We investigate how to exploit structural similarities of an individual's potential outcomes (POs) under different treatments to obtain better estimates of conditional average treatment effects in finite samples. Especially when it is unknown whether a treatment has an effect at all, it is natural to hypothesize that the POs are similar - yet, some existing strategies for treatment effect estimation employ regularization schemes that implicitly encourage heterogeneity even when it does not exist and fail to fully make use of shared structure. In this paper, we investigate and compare three end-to-end learning strategies to overcome this problem - based on regularization, reparametrization and a flexible multi-task architecture - each encoding inductive bias favoring shared behavior across POs. To build understanding of their relative strengths, we implement all strategies using neural networks and conduct a wide range of semi-synthetic experiments. We observe that all three approaches can lead to substantial improvements upon numerous baselines and gain insight into performance differences across various experimental settings.
Consistency of mechanistic causal discovery in continuous-time using Neural ODEs
May 06, 2021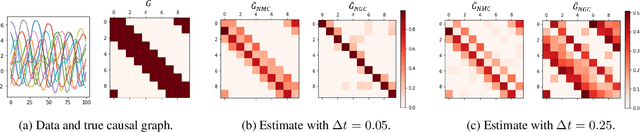
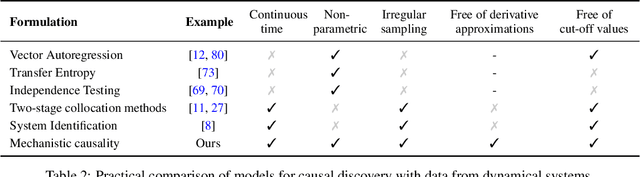
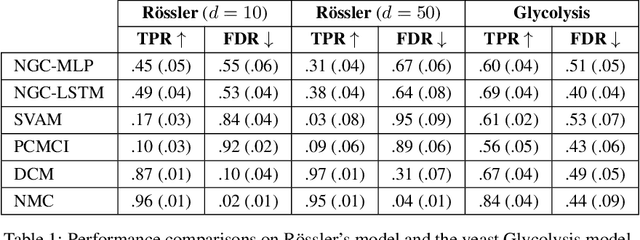
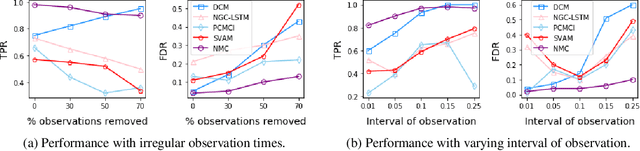

Abstract:The discovery of causal mechanisms from time series data is a key problem in fields working with complex systems. Most identifiability results and learning algorithms assume the underlying dynamics to be discrete in time. Comparatively few, in contrast, explicitly define causal associations in infinitesimal intervals of time, independently of the scale of observation and of the regularity of sampling. In this paper, we consider causal discovery in continuous-time for the study of dynamical systems. We prove that for vector fields parameterized in a large class of neural networks, adaptive regularization schemes consistently recover causal graphs in systems of ordinary differential equations (ODEs). Using this insight, we propose a causal discovery algorithm based on penalized Neural ODEs that we show to be applicable to the general setting of irregularly-sampled multivariate time series and to strongly outperform the state of the art.
Deconfounded Score Method: Scoring DAGs with Dense Unobserved Confounding
Mar 28, 2021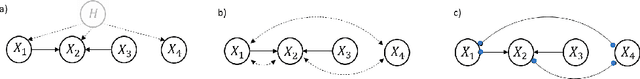
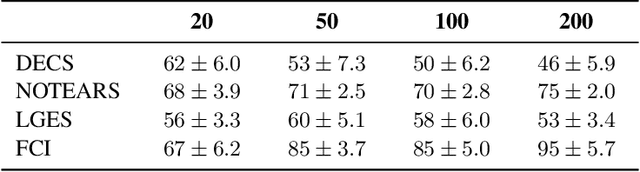
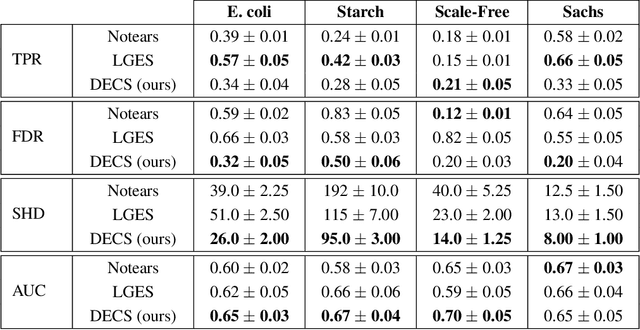
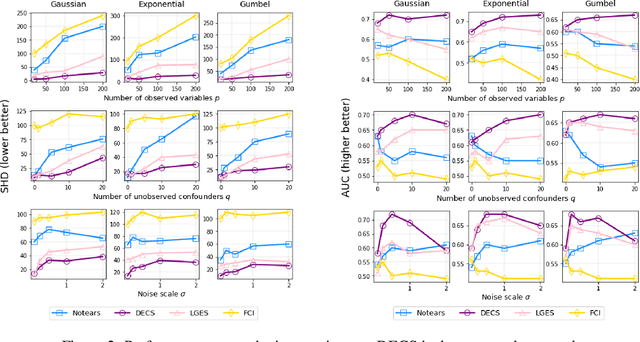

Abstract:Unobserved confounding is one of the greatest challenges for causal discovery. The case in which unobserved variables have a potentially widespread effect on many of the observed ones is particularly difficult because most pairs of variables are conditionally dependent given any other subset. In this paper, we show that beyond conditional independencies, unobserved confounding in this setting leaves a characteristic footprint in the observed data distribution that allows for disentangling spurious and causal effects. Using this insight, we demonstrate that a sparse linear Gaussian directed acyclic graph among observed variables may be recovered approximately and propose an adjusted score-based causal discovery algorithm that may be implemented with general-purpose solvers and scales to high-dimensional problems. We find, in addition, that despite the conditions we pose to guarantee causal recovery, performance in practice is robust to large deviations in model assumptions.
Scalable Bayesian Inverse Reinforcement Learning
Mar 11, 2021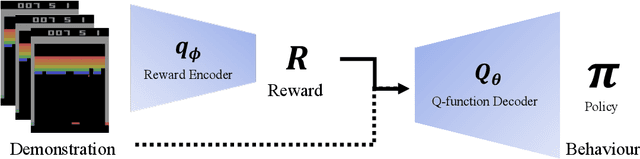
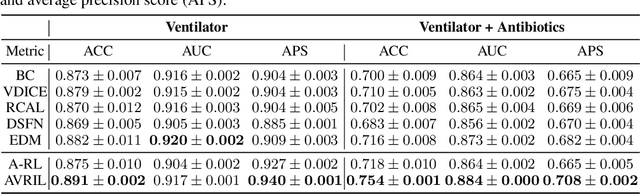
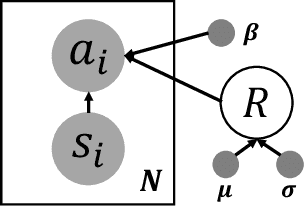
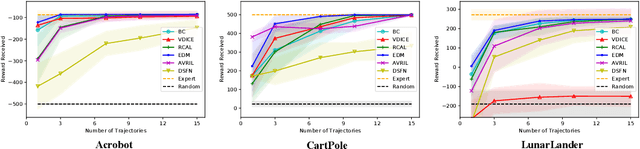
Abstract:Bayesian inference over the reward presents an ideal solution to the ill-posed nature of the inverse reinforcement learning problem. Unfortunately current methods generally do not scale well beyond the small tabular setting due to the need for an inner-loop MDP solver, and even non-Bayesian methods that do themselves scale often require extensive interaction with the environment to perform well, being inappropriate for high stakes or costly applications such as healthcare. In this paper we introduce our method, Approximate Variational Reward Imitation Learning (AVRIL), that addresses both of these issues by jointly learning an approximate posterior distribution over the reward that scales to arbitrarily complicated state spaces alongside an appropriate policy in a completely offline manner through a variational approach to said latent reward. Applying our method to real medical data alongside classic control simulations, we demonstrate Bayesian reward inference in environments beyond the scope of current methods, as well as task performance competitive with focused offline imitation learning algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge