Maurice HT Ling
Resistance Maintained in Digital Organisms despite Guanine/Cytosine-Based Fitness Cost and Extended De-Selection: Implications to Microbial Antibiotics Resistance
Feb 19, 2023Abstract:Antibiotics resistance has caused much complication in the treatment of diseases, where the pathogen is no longer susceptible to specific antibiotics and the use of such antibiotics are no longer effective for treatment. A recent study that utilizes digital organisms suggests that complete elimination of specific antibiotic resistance is unlikely after the disuse of antibiotics, assuming that there are no fitness costs for maintaining resistance once resistance are established. Fitness cost are referred to as reaction to change in environment, where organism improves its' abilities in one area at the expense of the other. Our goal in this study is to use digital organisms to examine the rate of gain and loss of resistance where fitness costs have incurred in maintaining resistance. Our results showed that GC-content based fitness cost during de-selection by removal of antibiotic-induced selective pressure portrayed similar trends in resistance compared to that of no fitness cost, at all stages of initial selection, repeated de-selection and re-introduction of selective pressure. Paired t-test suggested that prolonged stabilization of resistance after initial loss is not statistically significant for its difference to that of no fitness cost. This suggests that complete elimination of specific antibiotics resistance is unlikely after the disuse of antibiotics despite presence of fitness cost in maintaining antibiotic resistance during the disuse of antibiotics, once a resistant pool of micro-organism has been established.
On the Liveliness of Artificial Life
Feb 19, 2023Abstract:There has been on-going philosophical debate on whether artificial life models, also known as digital organisms, are truly alive. The main difficulty appears to be finding an encompassing and definite definition of life. By examining similarities and differences in recent definitions of life, we define life as "any system with a boundary to confine the system within a definite volume and protect the system from external effects, consisting of a program that is capable of improvisation, able to react and adapt to the environment, able to regenerate parts of it-self or its entirety, with energy system comprises of non-interference sets of secluded reactions for self-sustenance, is considered alive or a living system. Any incomplete system containing a program and can be re-assembled into a living system; thereby, converting the reassembled system for the purpose of the incomplete system, are also considered alive." Using this definition, we argue that digital organisms may not be the boundary case of life even though some digital organisms are not considered alive; thereby, taking the view that some form of digital organisms can be considered alive. In addition, we present an experimental framework based on continuity of the overall system and potential discontinuity of elements within the system for testing future definitions of life.
Parts-of-Speech Tagger Errors Do Not Necessarily Degrade Accuracy in Extracting Information from Biomedical Text
Apr 02, 2008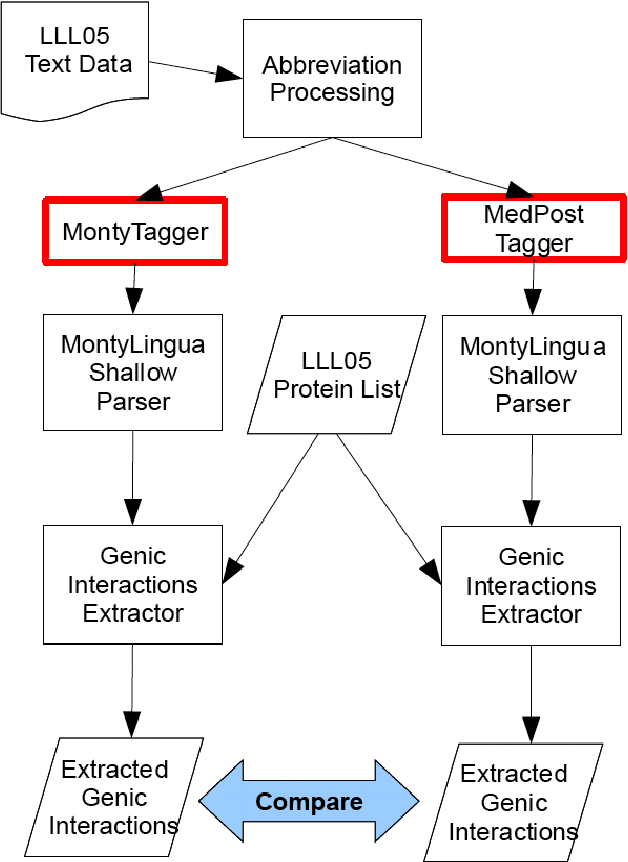
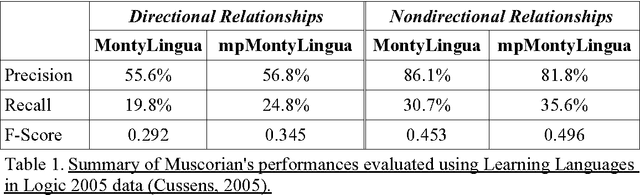
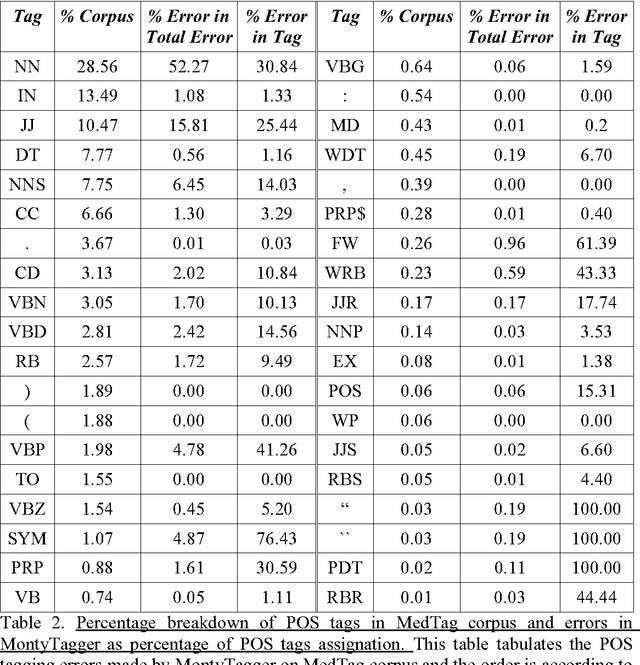
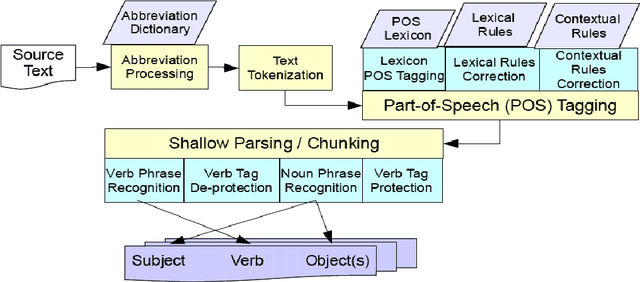
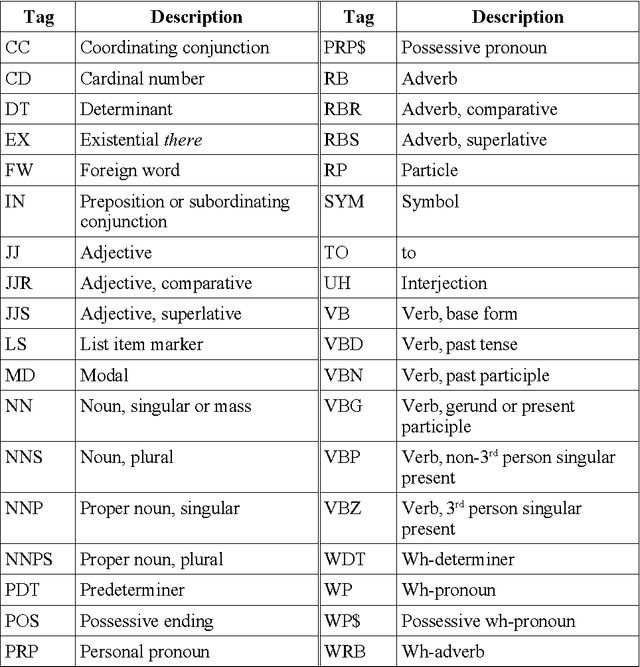
Abstract:A recent study reported development of Muscorian, a generic text processing tool for extracting protein-protein interactions from text that achieved comparable performance to biomedical-specific text processing tools. This result was unexpected since potential errors from a series of text analysis processes is likely to adversely affect the outcome of the entire process. Most biomedical entity relationship extraction tools have used biomedical-specific parts-of-speech (POS) tagger as errors in POS tagging and are likely to affect subsequent semantic analysis of the text, such as shallow parsing. This study aims to evaluate the parts-of-speech (POS) tagging accuracy and attempts to explore whether a comparable performance is obtained when a generic POS tagger, MontyTagger, was used in place of MedPost, a tagger trained in biomedical text. Our results demonstrated that MontyTagger, Muscorian's POS tagger, has a POS tagging accuracy of 83.1% when tested on biomedical text. Replacing MontyTagger with MedPost did not result in a significant improvement in entity relationship extraction from text; precision of 55.6% from MontyTagger versus 56.8% from MedPost on directional relationships and 86.1% from MontyTagger compared to 81.8% from MedPost on nondirectional relationships. This is unexpected as the potential for poor POS tagging by MontyTagger is likely to affect the outcome of the information extraction. An analysis of POS tagging errors demonstrated that 78.5% of tagging errors are being compensated by shallow parsing. Thus, despite 83.1% tagging accuracy, MontyTagger has a functional tagging accuracy of 94.6%.
Reconstruction of Protein-Protein Interaction Pathways by Mining Subject-Verb-Objects Intermediates
Aug 06, 2007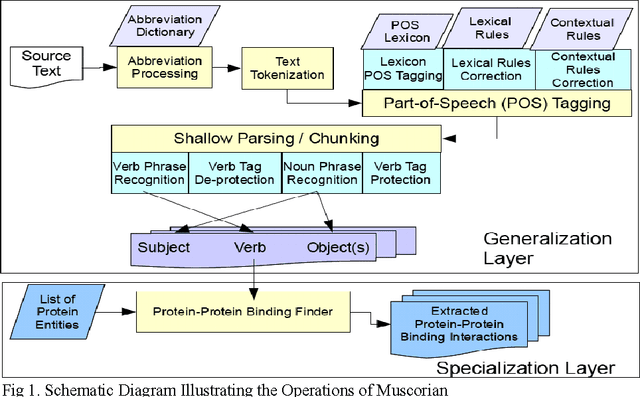
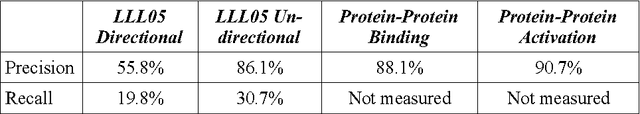
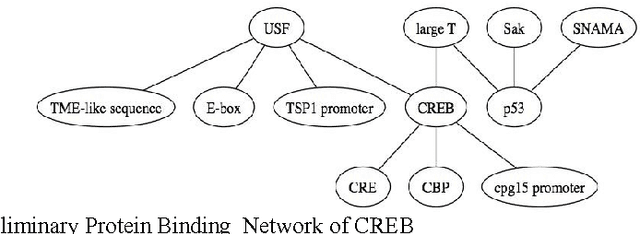
Abstract:The exponential increase in publication rate of new articles is limiting access of researchers to relevant literature. This has prompted the use of text mining tools to extract key biological information. Previous studies have reported extensive modification of existing generic text processors to process biological text. However, this requirement for modification had not been examined. In this study, we have constructed Muscorian, using MontyLingua, a generic text processor. It uses a two-layered generalization-specialization paradigm previously proposed where text was generically processed to a suitable intermediate format before domain-specific data extraction techniques are applied at the specialization layer. Evaluation using a corpus and experts indicated 86-90% precision and approximately 30% recall in extracting protein-protein interactions, which was comparable to previous studies using either specialized biological text processing tools or modified existing tools. Our study had also demonstrated the flexibility of the two-layered generalization-specialization paradigm by using the same generalization layer for two specialized information extraction tasks.
* 2nd IAPR Workshop on Pattern Recognition in Bioinformatics (PRIB 2007). 14 pages, 4 figures
An Anthological Review of Research Utilizing MontyLingua, a Python-Based End-to-End Text Processor
Nov 22, 2006Abstract:MontyLingua, an integral part of ConceptNet which is currently the largest commonsense knowledge base, is an English text processor developed using Python programming language in MIT Media Lab. The main feature of MontyLingua is the coverage for all aspects of English text processing from raw input text to semantic meanings and summary generation, yet each component in MontyLingua is loosely-coupled to each other at the architectural and code level, which enabled individual components to be used independently or substituted. However, there has been no review exploring the role of MontyLingua in recent research work utilizing it. This paper aims to review the use of and roles played by MontyLingua and its components in research work published in 19 articles between October 2004 and August 2006. We had observed a diversified use of MontyLingua in many different areas, both generic and domain-specific. Although the use of text summarizing component had not been observe, we are optimistic that it will have a crucial role in managing the current trend of information overload in future research.
* 9 pages
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge