Marcel van Herk
An anatomically-informed correspondence initialisation method to improve learning-based registration for radiotherapy
Feb 26, 2025


Abstract:We propose an anatomically-informed initialisation method for interpatient CT non-rigid registration (NRR), using a learning-based model to estimate correspondences between organ structures. A thin plate spline (TPS) deformation, set up using the correspondence predictions, is used to initialise the scans before a second NRR step. We compare two established NRR methods for the second step: a B-spline iterative optimisation-based algorithm and a deep learning-based approach. Registration performance is evaluated with and without the initialisation by assessing the similarity of propagated structures. Our proposed initialisation improved the registration performance of the learning-based method to more closely match the traditional iterative algorithm, with the mean distance-to-agreement reduced by 1.8mm for structures included in the TPS and 0.6mm for structures not included, while maintaining a substantial speed advantage (5 vs. 72 seconds).
Unsupervised correspondence with combined geometric learning and imaging for radiotherapy applications
Sep 25, 2023



Abstract:The aim of this study was to develop a model to accurately identify corresponding points between organ segmentations of different patients for radiotherapy applications. A model for simultaneous correspondence and interpolation estimation in 3D shapes was trained with head and neck organ segmentations from planning CT scans. We then extended the original model to incorporate imaging information using two approaches: 1) extracting features directly from image patches, and 2) including the mean square error between patches as part of the loss function. The correspondence and interpolation performance were evaluated using the geodesic error, chamfer distance and conformal distortion metrics, as well as distances between anatomical landmarks. Each of the models produced significantly better correspondences than the baseline non-rigid registration approach. The original model performed similarly to the model with direct inclusion of image features. The best performing model configuration incorporated imaging information as part of the loss function which produced more anatomically plausible correspondences. We will use the best performing model to identify corresponding anatomical points on organs to improve spatial normalisation, an important step in outcome modelling, or as an initialisation for anatomically informed registrations. All our code is publicly available at https://github.com/rrr-uom-projects/Unsup-RT-Corr-Net
The impact of training dataset size and ensemble inference strategies on head and neck auto-segmentation
Mar 30, 2023



Abstract:Convolutional neural networks (CNNs) are increasingly being used to automate segmentation of organs-at-risk in radiotherapy. Since large sets of highly curated data are scarce, we investigated how much data is required to train accurate and robust head and neck auto-segmentation models. For this, an established 3D CNN was trained from scratch with different sized datasets (25-1000 scans) to segment the brainstem, parotid glands and spinal cord in CTs. Additionally, we evaluated multiple ensemble techniques to improve the performance of these models. The segmentations improved with training set size up to 250 scans and the ensemble methods significantly improved performance for all organs. The impact of the ensemble methods was most notable in the smallest datasets, demonstrating their potential for use in cases where large training datasets are difficult to obtain.
Generalised Automatic Anatomy Finder (GAAF): A general framework for 3D location-finding in CT scans
Sep 13, 2022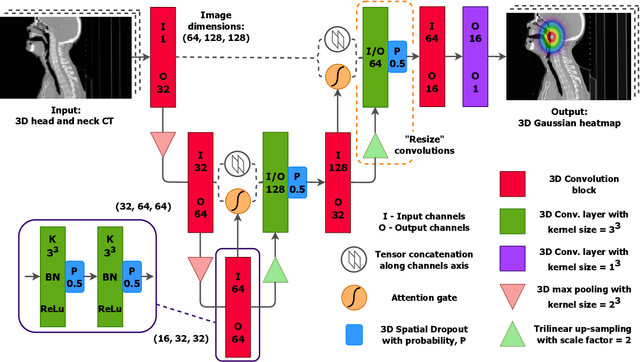
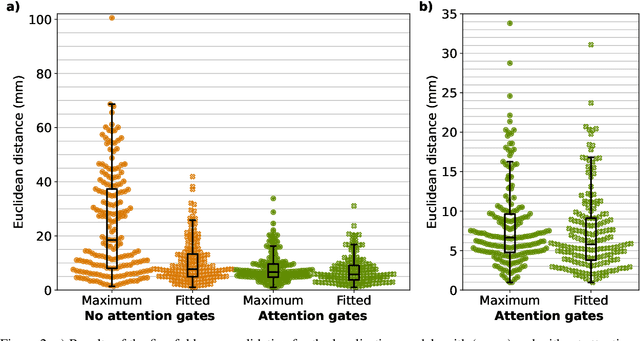
Abstract:We present GAAF, a Generalised Automatic Anatomy Finder, for the identification of generic anatomical locations in 3D CT scans. GAAF is an end-to-end pipeline, with dedicated modules for data pre-processing, model training, and inference. At it's core, GAAF uses a custom a localisation convolutional neural network (CNN). The CNN model is small, lightweight and can be adjusted to suit the particular application. The GAAF framework has so far been tested in the head and neck, and is able to find anatomical locations such as the centre-of-mass of the brainstem. GAAF was evaluated in an open-access dataset and is capable of accurate and robust localisation performance. All our code is open source and available at https://github.com/rrr-uom-projects/GAAF.
Automatic identification of segmentation errors for radiotherapy using geometric learning
Jun 27, 2022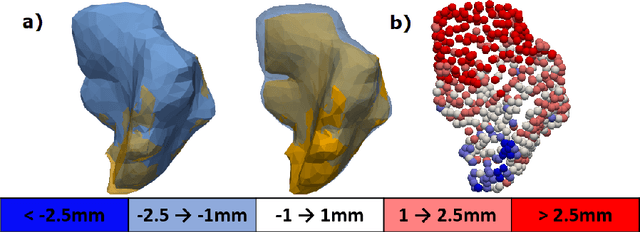
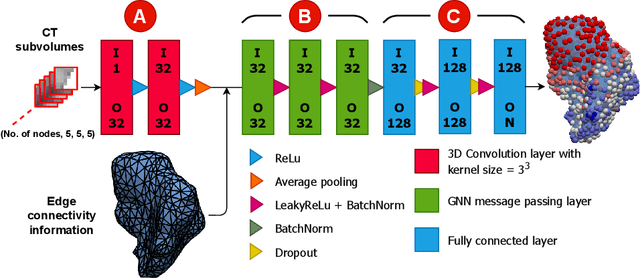
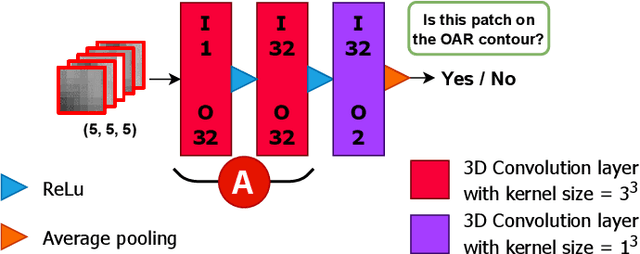
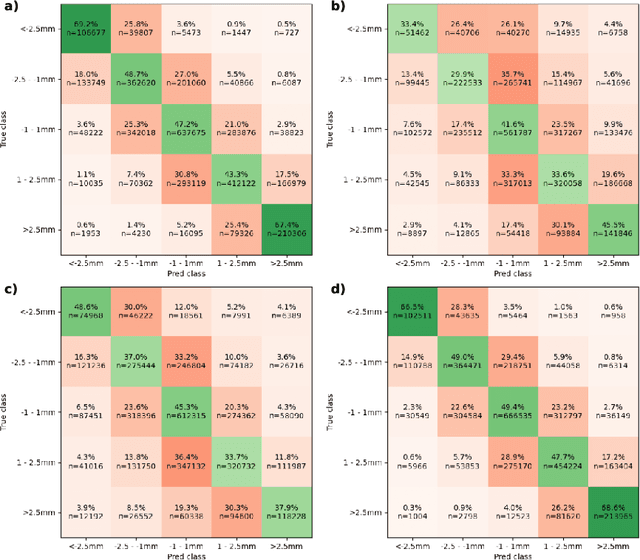
Abstract:Automatic segmentation of organs-at-risk (OARs) in CT scans using convolutional neural networks (CNNs) is being introduced into the radiotherapy workflow. However, these segmentations still require manual editing and approval by clinicians prior to clinical use, which can be time consuming. The aim of this work was to develop a tool to automatically identify errors in 3D OAR segmentations without a ground truth. Our tool uses a novel architecture combining a CNN and graph neural network (GNN) to leverage the segmentation's appearance and shape. The proposed model is trained using self-supervised learning using a synthetically-generated dataset of segmentations of the parotid and with realistic contouring errors. The effectiveness of our model is assessed with ablation tests, evaluating the efficacy of different portions of the architecture as well as the use of transfer learning from an unsupervised pretext task. Our best performing model predicted errors on the parotid gland with a precision of 85.0% & 89.7% for internal and external errors respectively, and recall of 66.5% & 68.6%. This offline QA tool could be used in the clinical pathway, potentially decreasing the time clinicians spend correcting contours by detecting regions which require their attention. All our code is publicly available at https://github.com/rrr-uom-projects/contour_auto_QATool.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge