M. Baity-Jesi
Class Imbalance in Anomaly Detection: Learning from an Exactly Solvable Model
Jan 20, 2025Abstract:Class imbalance (CI) is a longstanding problem in machine learning, slowing down training and reducing performances. Although empirical remedies exist, it is often unclear which ones work best and when, due to the lack of an overarching theory. We address a common case of imbalance, that of anomaly (or outlier) detection. We provide a theoretical framework to analyze, interpret and address CI. It is based on an exact solution of the teacher-student perceptron model, through replica theory. Within this framework, one can distinguish several sources of CI: either intrinsic, train or test imbalance. Our analysis reveals that the optimal train imbalance is generally different from 50%, with a non trivial dependence on the intrinsic imbalance, the abundance of data and on the noise in the learning. Moreover, there is a crossover between a small noise training regime where results are independent of the noise level to a high noise regime where performances quickly degrade with noise. Our results challenge some of the conventional wisdom on CI and offer practical guidelines to address it.
Ensembles of Vision Transformers as a New Paradigm for Automated Classification in Ecology
Mar 03, 2022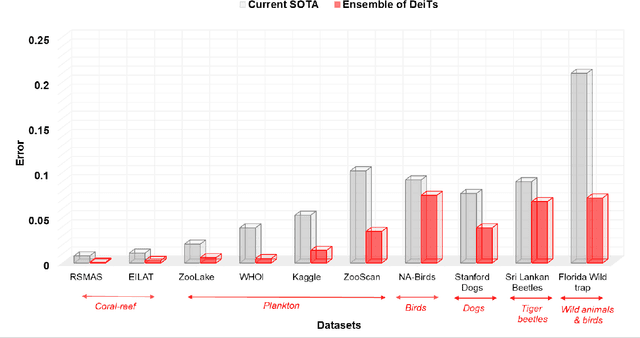
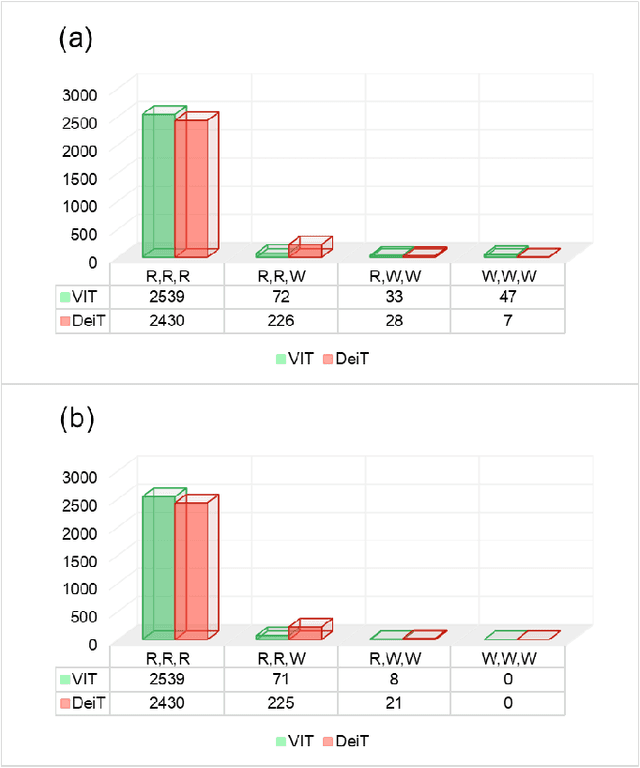
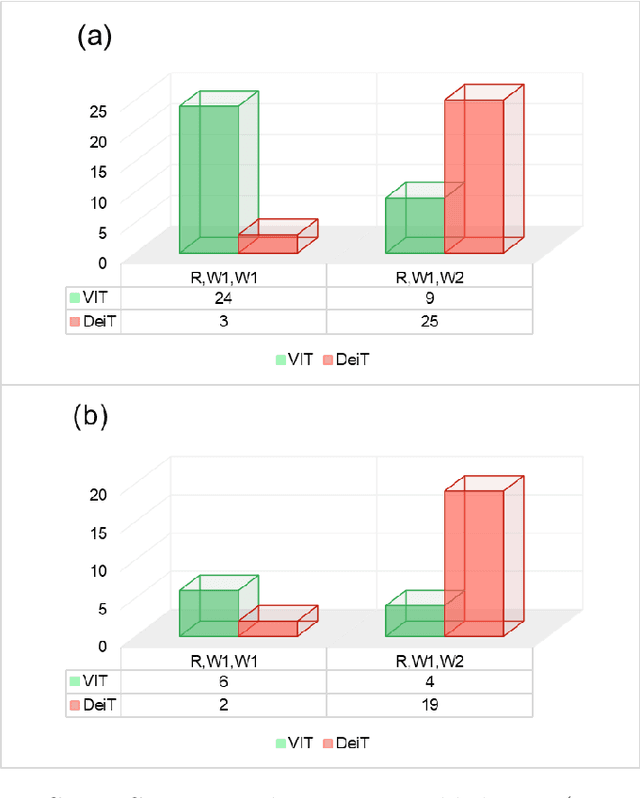
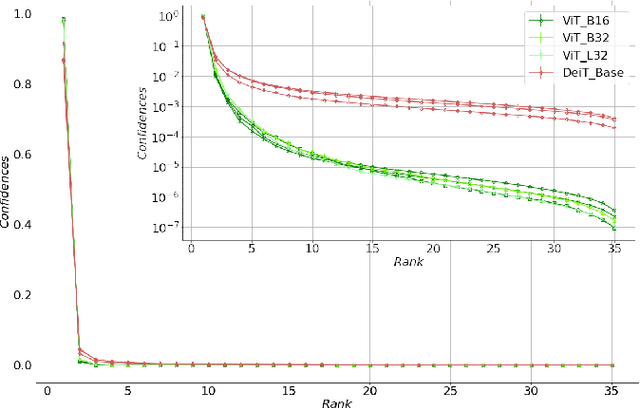
Abstract:Monitoring biodiversity is paramount to manage and protect natural resources, particularly in times of global change. Collecting images of organisms over large temporal or spatial scales is a promising practice to monitor and study biodiversity change of natural ecosystems, providing large amounts of data with minimal interference with the environment. Deep learning models are currently used to automate classification of organisms into taxonomic units. However, imprecision in these classifiers introduce a measurement noise that is difficult to control and can significantly hinder the analysis and interpretation of data. In our study, we show that this limitation can be overcome by ensembles of Data-efficient image Transformers (DeiTs), which significantly outperform the previous state of the art (SOTA). We validate our results on a large number of ecological imaging datasets of diverse origin, and organisms of study ranging from plankton to insects, birds, dog breeds, animals in the wild, and corals. On all the data sets we test, we achieve a new SOTA, with a reduction of the error with respect to the previous SOTA ranging from 18.48% to 87.50%, depending on the data set, and often achieving performances very close to perfect classification. The main reason why ensembles of DeiTs perform better is not due to the single-model performance of DeiTs, but rather to the fact that predictions by independent models have a smaller overlap, and this maximizes the profit gained by ensembling. This positions DeiT ensembles as the best candidate for image classification in biodiversity monitoring.
Deep Learning Classification of Lake Zooplankton
Aug 11, 2021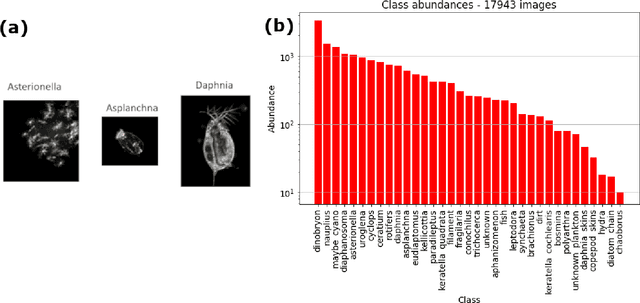
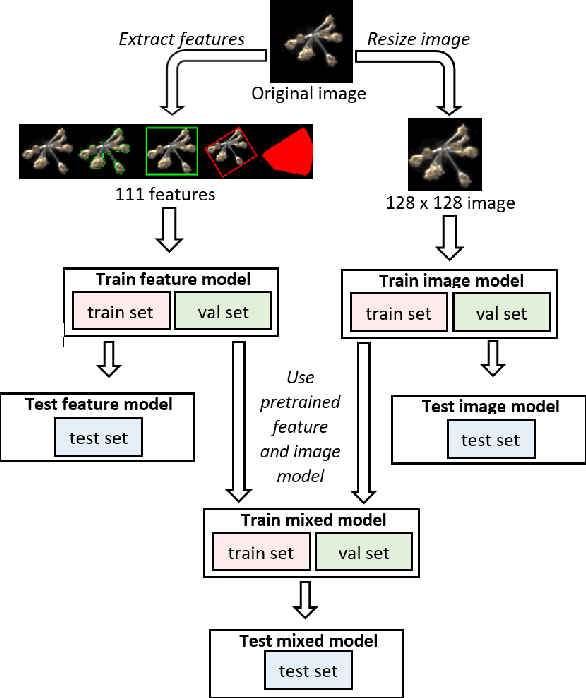
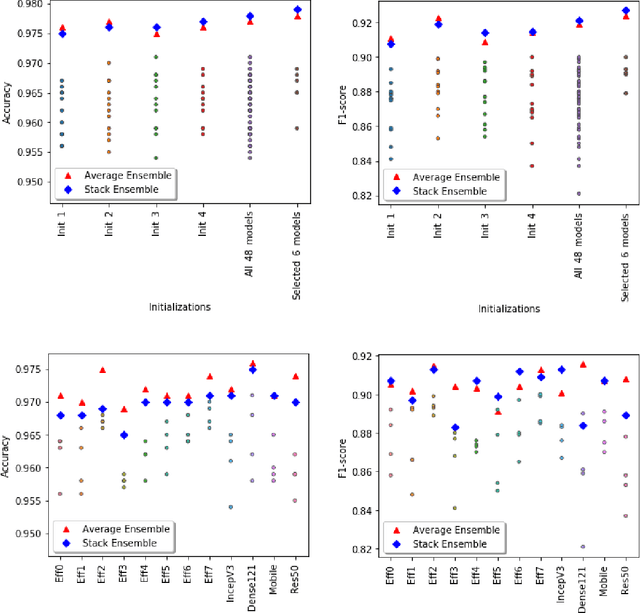
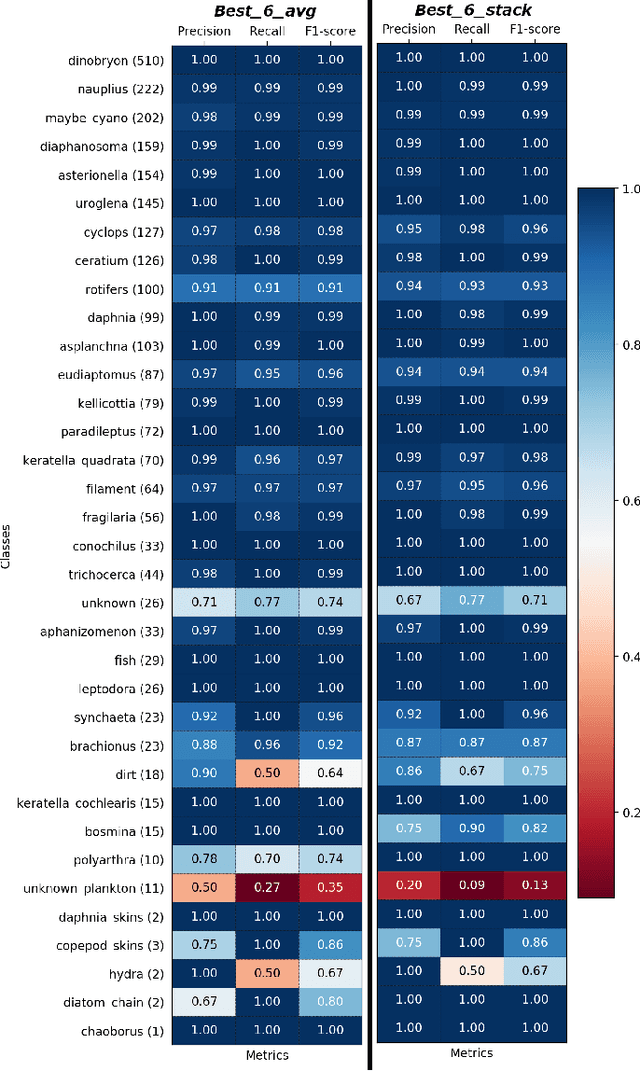
Abstract:Plankton are effective indicators of environmental change and ecosystem health in freshwater habitats, but collection of plankton data using manual microscopic methods is extremely labor-intensive and expensive. Automated plankton imaging offers a promising way forward to monitor plankton communities with high frequency and accuracy in real-time. Yet, manual annotation of millions of images proposes a serious challenge to taxonomists. Deep learning classifiers have been successfully applied in various fields and provided encouraging results when used to categorize marine plankton images. Here, we present a set of deep learning models developed for the identification of lake plankton, and study several strategies to obtain optimal performances,which lead to operational prescriptions for users. To this aim, we annotated into 35 classes over 17900 images of zooplankton and large phytoplankton colonies, detected in Lake Greifensee (Switzerland) with the Dual Scripps Plankton Camera. Our best models were based on transfer learning and ensembling, which classified plankton images with 98% accuracy and 93% F1 score. When tested on freely available plankton datasets produced by other automated imaging tools (ZooScan, FlowCytobot and ISIIS), our models performed better than previously used models. Our annotated data, code and classification models are freely available online.
Comparing Dynamics: Deep Neural Networks versus Glassy Systems
Jun 07, 2018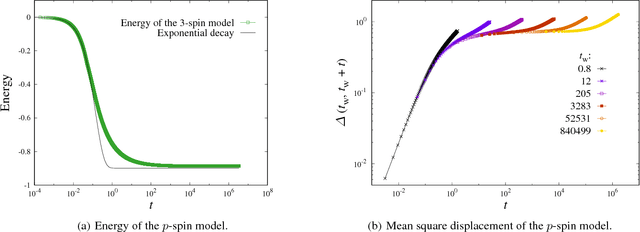
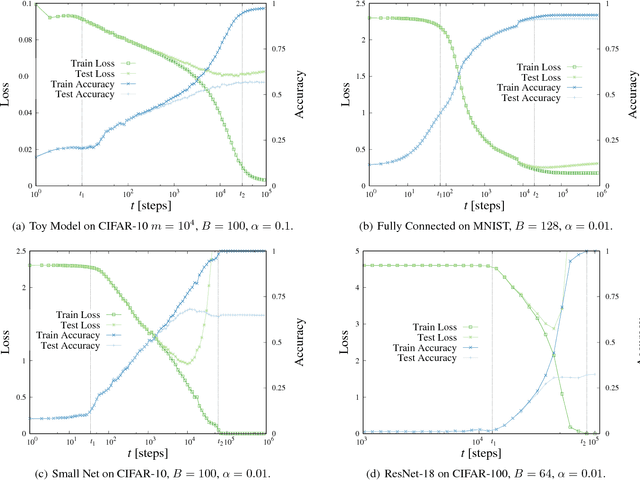
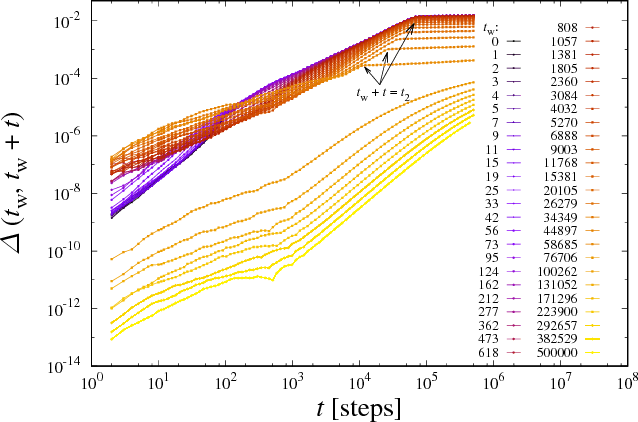
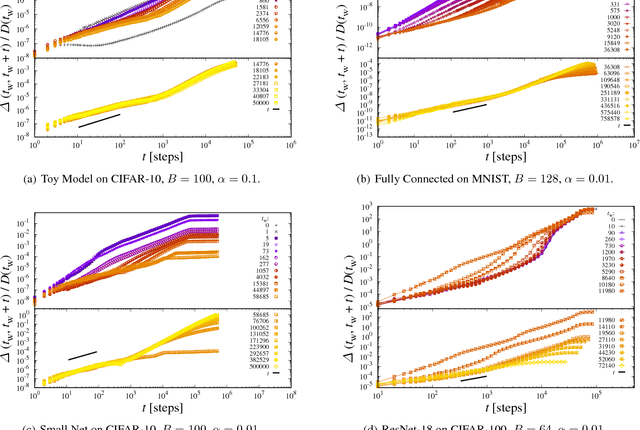
Abstract:We analyze numerically the training dynamics of deep neural networks (DNN) by using methods developed in statistical physics of glassy systems. The two main issues we address are (1) the complexity of the loss landscape and of the dynamics within it, and (2) to what extent DNNs share similarities with glassy systems. Our findings, obtained for different architectures and datasets, suggest that during the training process the dynamics slows down because of an increasingly large number of flat directions. At large times, when the loss is approaching zero, the system diffuses at the bottom of the landscape. Despite some similarities with the dynamics of mean-field glassy systems, in particular, the absence of barrier crossing, we find distinctive dynamical behaviors in the two cases, showing that the statistical properties of the corresponding loss and energy landscapes are different. In contrast, when the network is under-parametrized we observe a typical glassy behavior, thus suggesting the existence of different phases depending on whether the network is under-parametrized or over-parametrized.
* 10 pages, 5 figures. Version accepted at ICML 2018
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge