Lukas Uzolas
MotionDreamer: Zero-Shot 3D Mesh Animation from Video Diffusion Models
May 30, 2024



Abstract:Animation techniques bring digital 3D worlds and characters to life. However, manual animation is tedious and automated techniques are often specialized to narrow shape classes. In our work, we propose a technique for automatic re-animation of arbitrary 3D shapes based on a motion prior extracted from a video diffusion model. Unlike existing 4D generation methods, we focus solely on the motion, and we leverage an explicit mesh-based representation compatible with existing computer-graphics pipelines. Furthermore, our utilization of diffusion features enhances accuracy of our motion fitting. We analyze efficacy of these features for animation fitting and we experimentally validate our approach for two different diffusion models and four animation models. Finally, we demonstrate that our time-efficient zero-shot method achieves a superior performance re-animating a diverse set of 3D shapes when compared to existing techniques in a user study. The project website is located at https://lukas.uzolas.com/MotionDreamer.
Template-free Articulated Neural Point Clouds for Reposable View Synthesis
May 30, 2023



Abstract:Dynamic Neural Radiance Fields (NeRFs) achieve remarkable visual quality when synthesizing novel views of time-evolving 3D scenes. However, the common reliance on backward deformation fields makes reanimation of the captured object poses challenging. Moreover, the state of the art dynamic models are often limited by low visual fidelity, long reconstruction time or specificity to narrow application domains. In this paper, we present a novel method utilizing a point-based representation and Linear Blend Skinning (LBS) to jointly learn a Dynamic NeRF and an associated skeletal model from even sparse multi-view video. Our forward-warping approach achieves state-of-the-art visual fidelity when synthesizing novel views and poses while significantly reducing the necessary learning time when compared to existing work. We demonstrate the versatility of our representation on a variety of articulated objects from common datasets and obtain reposable 3D reconstructions without the need of object-specific skeletal templates. Code will be made available at https://github.com/lukasuz/Articulated-Point-NeRF.
Deep Anomaly Generation: An Image Translation Approach of Synthesizing Abnormal Banded Chromosome Images
Sep 20, 2021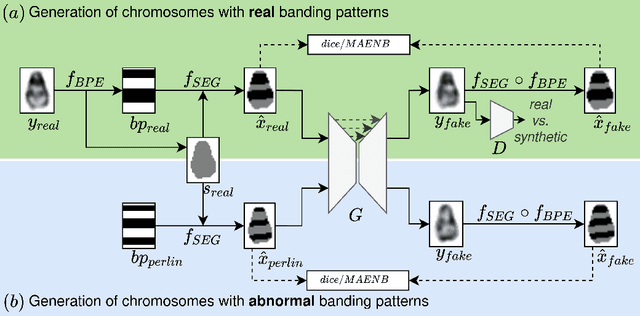
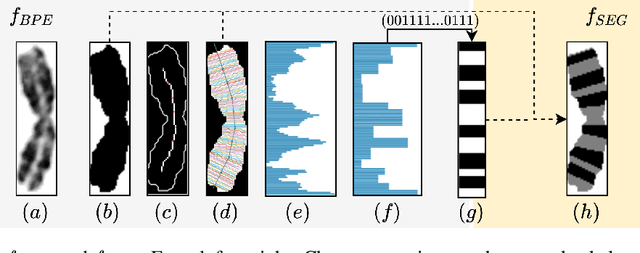
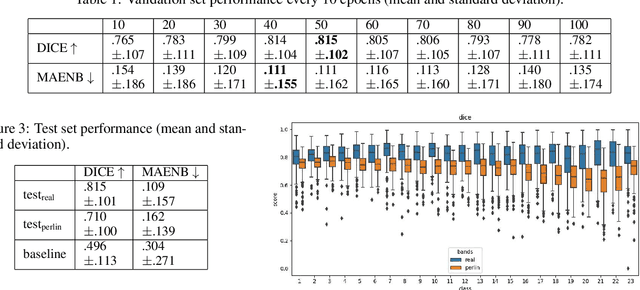
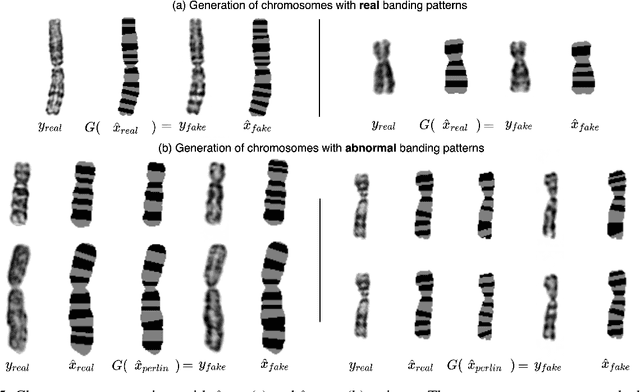
Abstract:Advances in deep-learning-based pipelines have led to breakthroughs in a variety of microscopy image diagnostics. However, a sufficiently big training data set is usually difficult to obtain due to high annotation costs. In the case of banded chromosome images, the creation of big enough libraries is difficult for multiple pathologies due to the rarity of certain genetic disorders. Generative Adversarial Networks (GANs) have proven to be effective in generating synthetic images and extending training data sets. In our work, we implement a conditional adversarial network that allows generation of realistic single chromosome images following user-defined banding patterns. To this end, an image-to-image translation approach based on self-generated 2D chromosome segmentation label maps is used. Our validation shows promising results when synthesizing chromosomes with seen as well as unseen banding patterns. We believe that this approach can be exploited for data augmentation of chromosome data sets with structural abnormalities. Therefore, the proposed method could help to tackle medical image analysis problems such as data simulation, segmentation, detection, or classification in the field of cytogenetics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge