Laurent Lejeune
A Positive/Unlabeled Approach for the Segmentation of Medical Sequences using Point-Wise Supervision
Jul 18, 2021
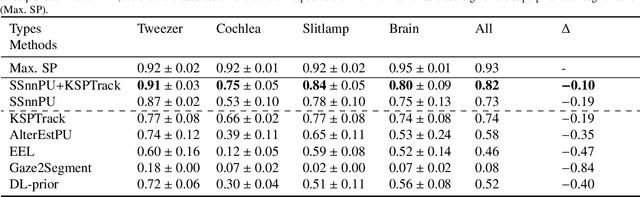
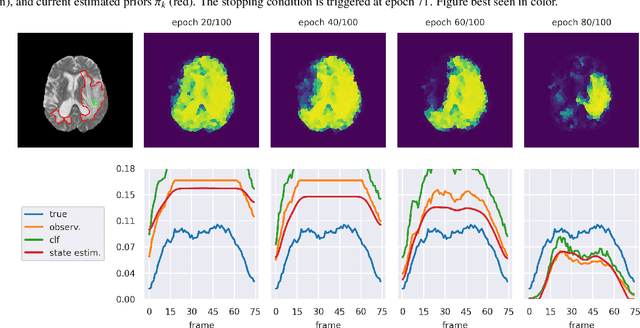
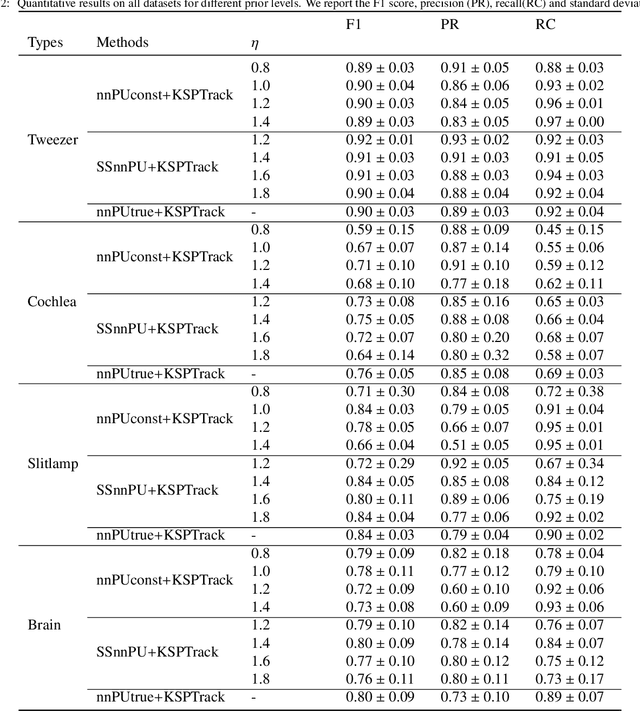
Abstract:The ability to quickly annotate medical imaging data plays a critical role in training deep learning frameworks for segmentation. Doing so for image volumes or video sequences is even more pressing as annotating these is particularly burdensome. To alleviate this problem, this work proposes a new method to efficiently segment medical imaging volumes or videos using point-wise annotations only. This allows annotations to be collected extremely quickly and remains applicable to numerous segmentation tasks. Our approach trains a deep learning model using an appropriate Positive/Unlabeled objective function using sparse point-wise annotations. While most methods of this kind assume that the proportion of positive samples in the data is known a-priori, we introduce a novel self-supervised method to estimate this prior efficiently by combining a Bayesian estimation framework and new stopping criteria. Our method iteratively estimates appropriate class priors and yields high segmentation quality for a variety of object types and imaging modalities. In addition, by leveraging a spatio-temporal tracking framework, we regularize our predictions by leveraging the complete data volume. We show experimentally that our approach outperforms state-of-the-art methods tailored to the same problem.
Iterative multi-path tracking for video and volume segmentation with sparse point supervision
Aug 27, 2018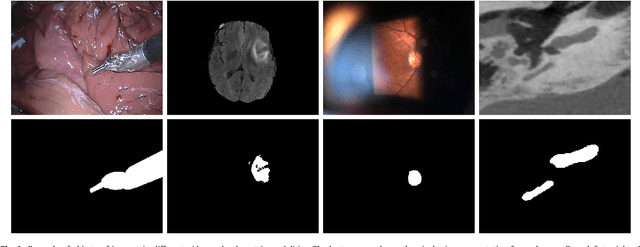
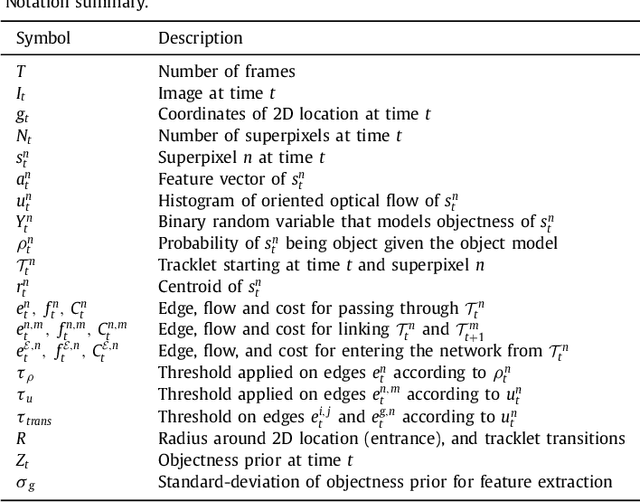
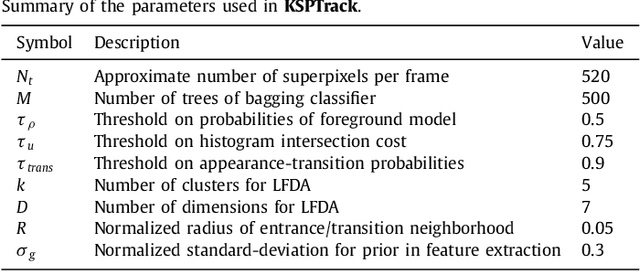

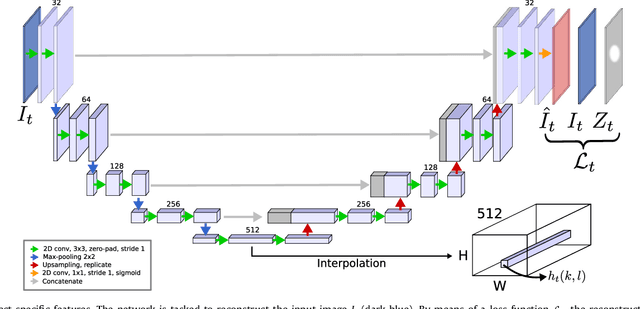

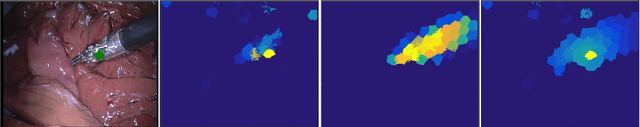
Abstract:Recent machine learning strategies for segmentation tasks have shown great ability when trained on large pixel-wise annotated image datasets. It remains a major challenge however to aggregate such datasets, as the time and monetary cost associated with collecting extensive annotations is extremely high. This is particularly the case for generating precise pixel-wise annotations in video and volumetric image data. To this end, this work presents a novel framework to produce pixel-wise segmentations using minimal supervision. Our method relies on 2D point supervision, whereby a single 2D location within an object of interest is provided on each image of the data. Our method then estimates the object appearance in a semi-supervised fashion by learning object-image-specific features and by using these in a semi-supervised learning framework. Our object model is then used in a graph-based optimization problem that takes into account all provided locations and the image data in order to infer the complete pixel-wise segmentation. In practice, we solve this optimally as a tracking problem using a K-shortest path approach. Both the object model and segmentation are then refined iteratively to further improve the final segmentation. We show that by collecting 2D locations using a gaze tracker, our approach can provide state-of-the-art segmentations on a range of objects and image modalities (video and 3D volumes), and that these can then be used to train supervised machine learning classifiers.
Expected exponential loss for gaze-based video and volume ground truth annotation
Jul 16, 2017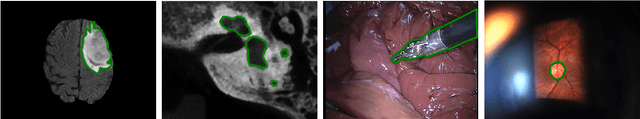
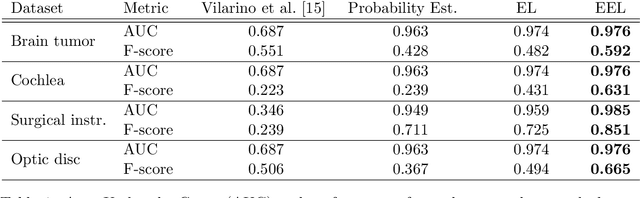
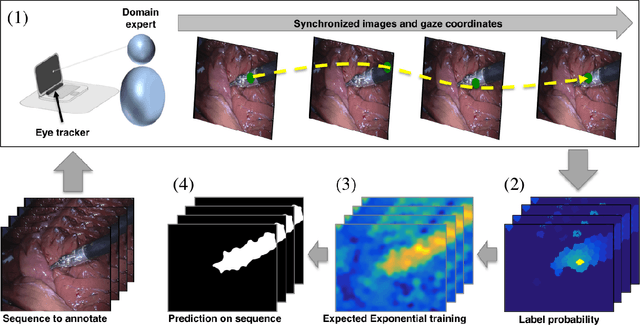

Abstract:Many recent machine learning approaches used in medical imaging are highly reliant on large amounts of image and ground truth data. In the context of object segmentation, pixel-wise annotations are extremely expensive to collect, especially in video and 3D volumes. To reduce this annotation burden, we propose a novel framework to allow annotators to simply observe the object to segment and record where they have looked at with a \$200 eye gaze tracker. Our method then estimates pixel-wise probabilities for the presence of the object throughout the sequence from which we train a classifier in semi-supervised setting using a novel Expected Exponential loss function. We show that our framework provides superior performances on a wide range of medical image settings compared to existing strategies and that our method can be combined with current crowd-sourcing paradigms as well.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge