Lars Ailo Bongo
Department of Computer Science, UiT The Arctic University of Norway, Tromsø, Norway
Automating tumor-infiltrating lymphocyte assessment in breast cancer histopathology images using QuPath: a transparent and accessible machine learning pipeline
Apr 23, 2025Abstract:In this study, we built an end-to-end tumor-infiltrating lymphocytes (TILs) assessment pipeline within QuPath, demonstrating the potential of easily accessible tools to perform complex tasks in a fully automatic fashion. First, we trained a pixel classifier to segment tumor, tumor-associated stroma, and other tissue compartments in breast cancer H&E-stained whole-slide images (WSI) to isolate tumor-associated stroma for subsequent analysis. Next, we applied a pre-trained StarDist deep learning model in QuPath for cell detection and used the extracted cell features to train a binary classifier distinguishing TILs from other cells. To evaluate our TILs assessment pipeline, we calculated the TIL density in each WSI and categorized them as low, medium, or high TIL levels. Our pipeline was evaluated against pathologist-assigned TIL scores, achieving a Cohen's kappa of 0.71 on the external test set, corroborating previous research findings. These results confirm that existing software can offer a practical solution for the assessment of TILs in H&E-stained WSIs of breast cancer.
Open-source framework for detecting bias and overfitting for large pathology images
Mar 03, 2025Abstract:Even foundational models that are trained on datasets with billions of data samples may develop shortcuts that lead to overfitting and bias. Shortcuts are non-relevant patterns in data, such as the background color or color intensity. So, to ensure the robustness of deep learning applications, there is a need for methods to detect and remove such shortcuts. Today's model debugging methods are time consuming since they often require customization to fit for a given model architecture in a specific domain. We propose a generalized, model-agnostic framework to debug deep learning models. We focus on the domain of histopathology, which has very large images that require large models - and therefore large computation resources. It can be run on a workstation with a commodity GPU. We demonstrate that our framework can replicate non-image shortcuts that have been found in previous work for self-supervised learning models, and we also identify possible shortcuts in a foundation model. Our easy to use tests contribute to the development of more reliable, accurate, and generalizable models for WSI analysis. Our framework is available as an open-source tool available on github.
A Lightweight and Extensible Cell Segmentation and Classification Model for Whole Slide Images
Feb 26, 2025



Abstract:Developing clinically useful cell-level analysis tools in digital pathology remains challenging due to limitations in dataset granularity, inconsistent annotations, high computational demands, and difficulties integrating new technologies into workflows. To address these issues, we propose a solution that enhances data quality, model performance, and usability by creating a lightweight, extensible cell segmentation and classification model. First, we update data labels through cross-relabeling to refine annotations of PanNuke and MoNuSAC, producing a unified dataset with seven distinct cell types. Second, we leverage the H-Optimus foundation model as a fixed encoder to improve feature representation for simultaneous segmentation and classification tasks. Third, to address foundation models' computational demands, we distill knowledge to reduce model size and complexity while maintaining comparable performance. Finally, we integrate the distilled model into QuPath, a widely used open-source digital pathology platform. Results demonstrate improved segmentation and classification performance using the H-Optimus-based model compared to a CNN-based model. Specifically, average $R^2$ improved from 0.575 to 0.871, and average $PQ$ score improved from 0.450 to 0.492, indicating better alignment with actual cell counts and enhanced segmentation quality. The distilled model maintains comparable performance while reducing parameter count by a factor of 48. By reducing computational complexity and integrating into workflows, this approach may significantly impact diagnostics, reduce pathologist workload, and improve outcomes. Although the method shows promise, extensive validation is necessary prior to clinical deployment.
Prompt Engineering a Schizophrenia Chatbot: Utilizing a Multi-Agent Approach for Enhanced Compliance with Prompt Instructions
Oct 10, 2024

Abstract:Patients with schizophrenia often present with cognitive impairments that may hinder their ability to learn about their condition. These individuals could benefit greatly from education platforms that leverage the adaptability of Large Language Models (LLMs) such as GPT-4. While LLMs have the potential to make topical mental health information more accessible and engaging, their black-box nature raises concerns about ethics and safety. Prompting offers a way to produce semi-scripted chatbots with responses anchored in instructions and validated information, but prompt-engineered chatbots may drift from their intended identity as the conversation progresses. We propose a Critical Analysis Filter for achieving better control over chatbot behavior. In this system, a team of prompted LLM agents are prompt-engineered to critically analyze and refine the chatbot's response and deliver real-time feedback to the chatbot. To test this approach, we develop an informational schizophrenia chatbot and converse with it (with the filter deactivated) until it oversteps its scope. Once drift has been observed, AI-agents are used to automatically generate sample conversations in which the chatbot is being enticed to talk about out-of-bounds topics. We manually assign to each response a compliance score that quantifies the chatbot's compliance to its instructions; specifically the rules about accurately conveying sources and being transparent about limitations. Activating the Critical Analysis Filter resulted in an acceptable compliance score (>=2) in 67.0% of responses, compared to only 8.7% when the filter was deactivated. These results suggest that a self-reflection layer could enable LLMs to be used effectively and safely in mental health platforms, maintaining adaptability while reliably limiting their scope to appropriate use cases.
Coding historical causes of death data with Large Language Models
May 13, 2024


Abstract:This paper investigates the feasibility of using pre-trained generative Large Language Models (LLMs) to automate the assignment of ICD-10 codes to historical causes of death. Due to the complex narratives often found in historical causes of death, this task has traditionally been manually performed by coding experts. We evaluate the ability of GPT-3.5, GPT-4, and Llama 2 LLMs to accurately assign ICD-10 codes on the HiCaD dataset that contains causes of death recorded in the civil death register entries of 19,361 individuals from Ipswich, Kilmarnock, and the Isle of Skye from the UK between 1861-1901. Our findings show that GPT-3.5, GPT-4, and Llama 2 assign the correct code for 69%, 83%, and 40% of causes, respectively. However, we achieve a maximum accuracy of 89% by standard machine learning techniques. All LLMs performed better for causes of death that contained terms still in use today, compared to archaic terms. Also they perform better for short causes (1-2 words) compared to longer causes. LLMs therefore do not currently perform well enough for historical ICD-10 code assignment tasks. We suggest further fine-tuning or alternative frameworks to achieve adequate performance.
Fast TILs estimation in lung cancer WSIs based on semi-stochastic patch sampling
May 05, 2024



Abstract:Addressing the critical need for accurate prognostic biomarkers in cancer treatment, quantifying tumor-infiltrating lymphocytes (TILs) in non-small cell lung cancer (NSCLC) presents considerable challenges. Manual TIL quantification in whole slide images (WSIs) is laborious and subject to variability, potentially undermining patient outcomes. Our study introduces an automated pipeline that utilizes semi-stochastic patch sampling, patch classification to retain prognostically relevant patches, and cell quantification using the HoVer-Net model to streamline the TIL evaluation process. This pipeline efficiently excludes approximately 70% of areas not relevant for prognosis and requires only 5% of the remaining patches to maintain prognostic accuracy (c-index 0.65 +- 0.01). The computational efficiency achieved does not sacrifice prognostic accuracy, as demonstrated by the TILs score's strong correlation with patient survival, which surpasses traditional CD8 IHC scoring methods. While the pipeline demonstrates potential for enhancing NSCLC prognostication and personalization of treatment, comprehensive clinical validation is still required. Future research should focus on verifying its broader clinical utility and investigating additional biomarkers to improve NSCLC prognosis.
Interoperable synthetic health data with SyntHIR to enable the development of CDSS tools
Aug 04, 2023



Abstract:There is a great opportunity to use high-quality patient journals and health registers to develop machine learning-based Clinical Decision Support Systems (CDSS). To implement a CDSS tool in a clinical workflow, there is a need to integrate, validate and test this tool on the Electronic Health Record (EHR) systems used to store and manage patient data. However, it is often not possible to get the necessary access to an EHR system due to legal compliance. We propose an architecture for generating and using synthetic EHR data for CDSS tool development. The architecture is implemented in a system called SyntHIR. The SyntHIR system uses the Fast Healthcare Interoperability Resources (FHIR) standards for data interoperability, the Gretel framework for generating synthetic data, the Microsoft Azure FHIR server as the FHIR-based EHR system and SMART on FHIR framework for tool transportability. We demonstrate the usefulness of SyntHIR by developing a machine learning-based CDSS tool using data from the Norwegian Patient Register (NPR) and Norwegian Patient Prescriptions (NorPD). We demonstrate the development of the tool on the SyntHIR system and then lift it to the Open DIPS environment. In conclusion, SyntHIR provides a generic architecture for CDSS tool development using synthetic FHIR data and a testing environment before implementing it in a clinical setting. However, there is scope for improvement in terms of the quality of the synthetic data generated. The code is open source and available at https://github.com/potter-coder89/SyntHIR.git.
More efficient manual review of automatically transcribed tabular data
Jun 28, 2023Abstract:Machine learning methods have proven useful in transcribing historical data. However, results from even highly accurate methods require manual verification and correction. Such manual review can be time-consuming and expensive, therefore the objective of this paper was to make it more efficient. Previously, we used machine learning to transcribe 2.3 million handwritten occupation codes from the Norwegian 1950 census with high accuracy (97%). We manually reviewed the 90,000 (3%) codes with the lowest model confidence. We allocated those 90,000 codes to human reviewers, who used our annotation tool to review the codes. To assess reviewer agreement, some codes were assigned to multiple reviewers. We then analyzed the review results to understand the relationship between accuracy improvements and effort. Additionally, we interviewed the reviewers to improve the workflow. The reviewers corrected 62.8% of the labels and agreed with the model label in 31.9% of cases. About 0.2% of the images could not be assigned a label, while for 5.1% the reviewers were uncertain, or they assigned an invalid label. 9,000 images were independently reviewed by multiple reviewers, resulting in an agreement of 86.43% and disagreement of 8.96%. We learned that our automatic transcription is biased towards the most frequent codes, with a higher degree of misclassification for the lowest frequency codes. Our interview findings show that the reviewers did internal quality control and found our custom tool well-suited. So, only one reviewer is needed, but they should report uncertainty.
Publicly available datasets of breast histopathology H&E whole-slide images: A systematic review
Jun 02, 2023Abstract:Advancements in digital pathology and computing resources have made a significant impact in the field of computational pathology for breast cancer diagnosis and treatment. However, access to high-quality labeled histopathological images of breast cancer is a big challenge that limits the development of accurate and robust deep learning models. In this systematic review, we identified the publicly available datasets of breast H&E stained whole-slide images (WSI) that can be used to develop deep learning algorithms. We systematically searched nine scientific literature databases and nine research data repositories. We found twelve publicly available datasets, containing 5153 H&E WSIs of breast cancer. Moreover, we reported image metadata and characteristics for each dataset to assist researchers in selecting proper datasets for specific tasks in breast cancer computational pathology. In addition, we compiled a list of patch and private datasets that were used in the included articles as a supplementary resource for researchers. Notably, 22% of the included articles utilized multiple datasets, and only 12% of the articles used an external validation set, suggesting that the performance of other developed models may be susceptible to overestimation. The TCGA-BRCA was used in 47.4% of the selected studies. This dataset has a considerable selection bias that can impact the robustness and generalizability of the trained algorithms. There is also a lack of consistent metadata reporting of breast WSI datasets that can be an issue in developing accurate deep learning models, indicating the necessity of establishing explicit guidelines for documenting breast WSI dataset characteristics and metadata.
A Pragmatic Machine Learning Approach to Quantify Tumor Infiltrating Lymphocytes in Whole Slide Images
Feb 14, 2022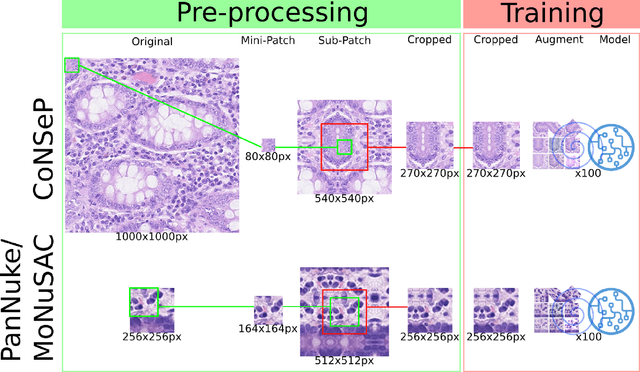
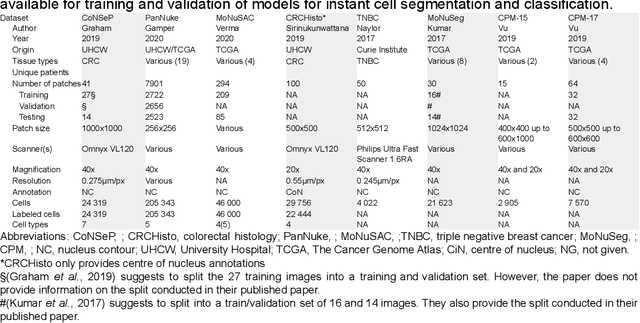
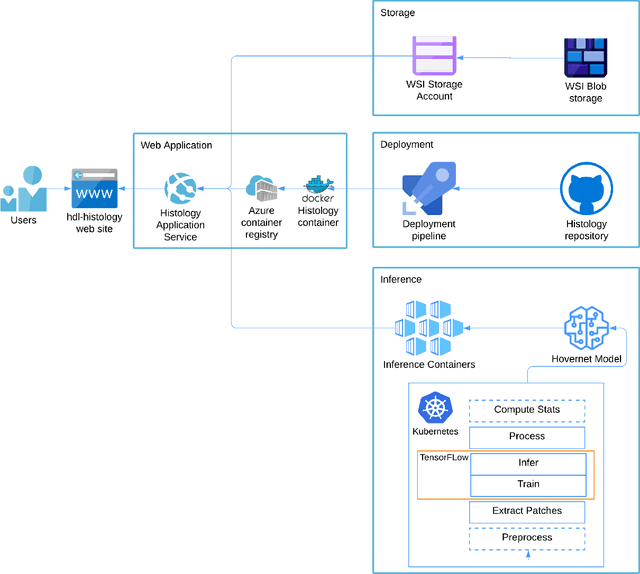
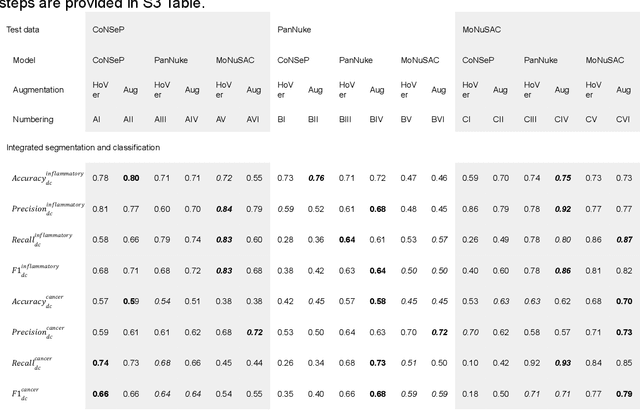
Abstract:Increased levels of tumor infiltrating lymphocytes (TILs) in cancer tissue indicate favourable outcomes in many types of cancer. Manual quantification of immune cells is inaccurate and time consuming for pathologists. Our aim is to leverage a computational solution to automatically quantify TILs in whole slide images (WSIs) of standard diagnostic haematoxylin and eosin stained sections (H&E slides) from lung cancer patients. Our approach is to transfer an open source machine learning method for segmentation and classification of nuclei in H&E slides trained on public data to TIL quantification without manual labeling of our data. Our results show that additional augmentation improves model transferability when training on few samples/limited tissue types. Models trained with sufficient samples/tissue types do not benefit from our additional augmentation policy. Further, the resulting TIL quantification correlates to patient prognosis and compares favorably to the current state-of-the-art method for immune cell detection in non-small lung cancer (current standard CD8 cells in DAB stained TMAs HR 0.34 95% CI 0.17-0.68 vs TILs in HE WSIs: HoVer-Net PanNuke Aug Model HR 0.30 95% CI 0.15-0.60, HoVer-Net MoNuSAC Aug model HR 0.27 95% CI 0.14-0.53). Moreover, we implemented a cloud based system to train, deploy and visually inspect machine learning based annotation for H&E slides. Our pragmatic approach bridges the gap between machine learning research, translational clinical research and clinical implementation. However, validation in prospective studies is needed to assert that the method works in a clinical setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge