Kyriakos Schwarz
DDoS: A Graph Neural Network based Drug Synergy Prediction Algorithm
Oct 10, 2022
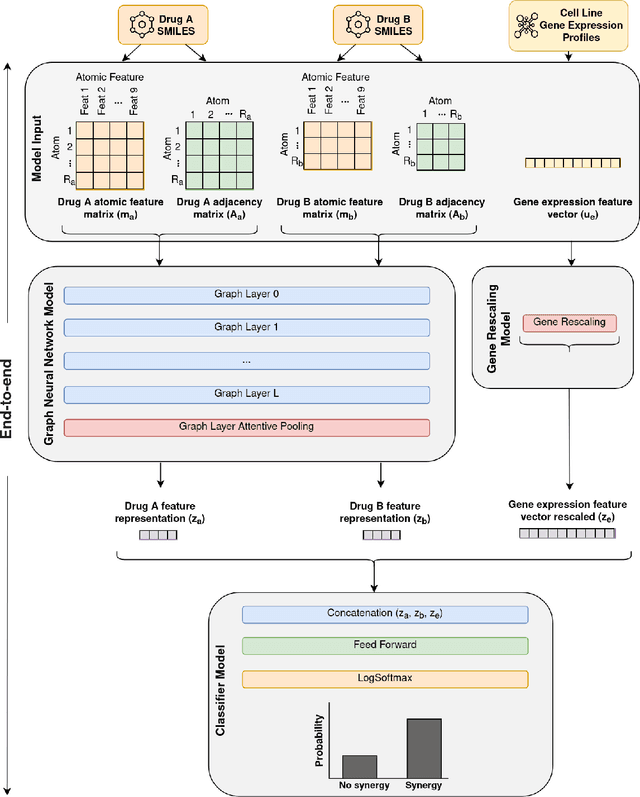
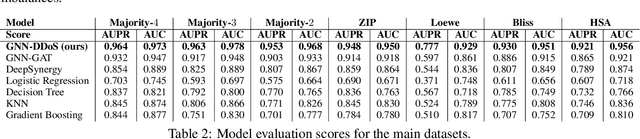
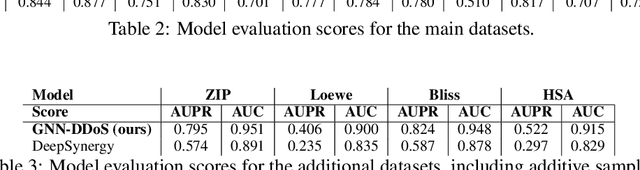
Abstract:Background: Drug synergy occurs when the combined effect of two drugs is greater than the sum of the individual drugs' effect. While cell line data measuring the effect of single drugs are readily available, there is relatively less comparable data on drug synergy given the vast amount of possible drug combinations. Thus, there is interest to use computational approaches to predict drug synergy for untested pairs of drugs. Methods: We introduce a Graph Neural Network (GNN) based model for drug synergy prediction, which utilizes drug chemical structures and cell line gene expression data. We use information from the largest drug combination database available (DrugComb), combining drug synergy scores in order to construct high confidence benchmark datasets. Results: Our proposed solution for drug synergy predictions offers a number of benefits: 1) It utilizes a combination of 34 distinct drug synergy datasets to learn on a wide variety of drugs and cell lines representations. 2) It is trained on constructed high confidence benchmark datasets. 3) It learns task-specific drug representations, instead of relying on generalized and pre-computed chemical drug features. 4) It achieves similar or better prediction performance (AUPR scores ranging from 0.777 to 0.964) compared to state-of-the-art baseline models when tested on various benchmark datasets. Conclusions: We demonstrate that a GNN based model can provide state-of-the-art drug synergy predictions by learning task-specific representations of drugs.
AttentionDDI: Siamese Attention-based Deep Learning method for drug-drug interaction predictions
Dec 24, 2020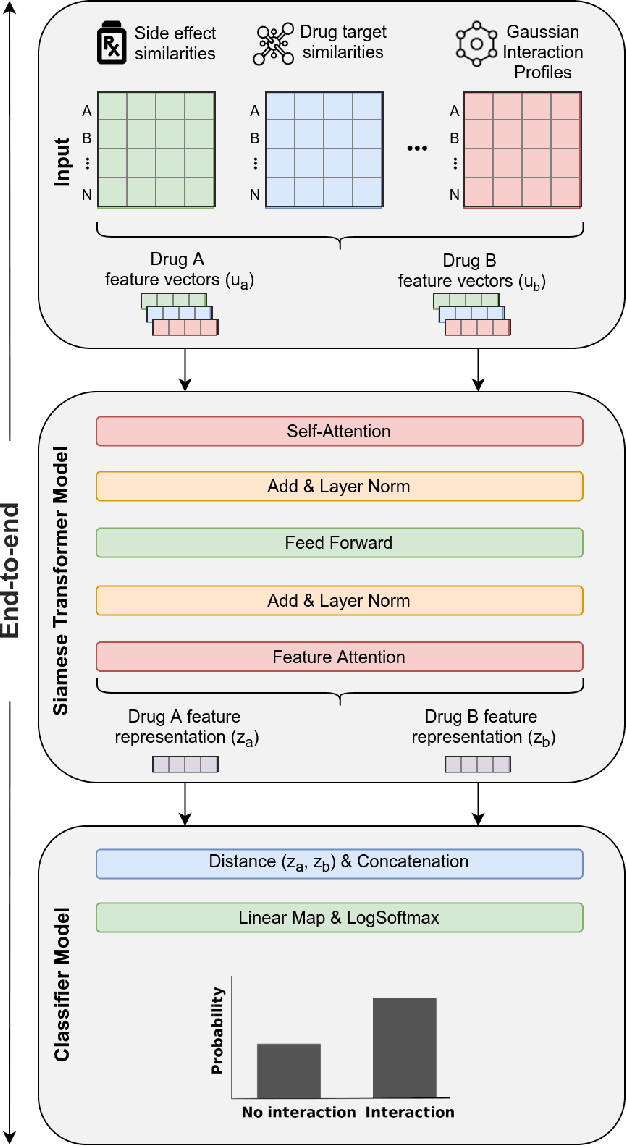
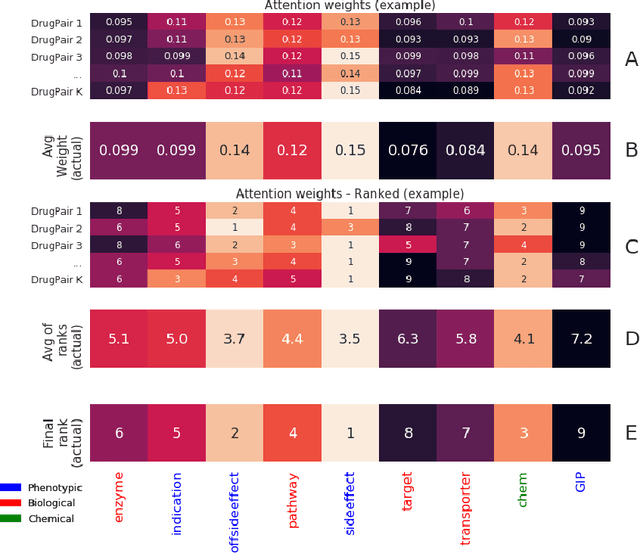
Abstract:Background: Drug-drug interactions (DDIs) refer to processes triggered by the administration of two or more drugs leading to side effects beyond those observed when drugs are administered by themselves. Due to the massive number of possible drug pairs, it is nearly impossible to experimentally test all combinations and discover previously unobserved side effects. Therefore, machine learning based methods are being used to address this issue. Methods: We propose a Siamese self-attention multi-modal neural network for DDI prediction that integrates multiple drug similarity measures that have been derived from a comparison of drug characteristics including drug targets, pathways and gene expression profiles. Results: Our proposed DDI prediction model provides multiple advantages: 1) It is trained end-to-end, overcoming limitations of models composed of multiple separate steps, 2) it offers model explainability via an Attention mechanism for identifying salient input features and 3) it achieves similar or better prediction performance (AUPR scores ranging from 0.77 to 0.92) compared to state-of-the-art DDI models when tested on various benchmark datasets. Novel DDI predictions are further validated using independent data resources. Conclusions: We find that a Siamese multi-modal neural network is able to accurately predict DDIs and that an Attention mechanism, typically used in the Natural Language Processing domain, can be beneficially applied to aid in DDI model explainability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge