Korbinian Strimmer
Optimal whitening and decorrelation
Dec 18, 2016

Abstract:Whitening, or sphering, is a common preprocessing step in statistical analysis to transform random variables to orthogonality. However, due to rotational freedom there are infinitely many possible whitening procedures. Consequently, there is a diverse range of sphering methods in use, for example based on principal component analysis (PCA), Cholesky matrix decomposition and zero-phase component analysis (ZCA), among others. Here we provide an overview of the underlying theory and discuss five natural whitening procedures. Subsequently, we demonstrate that investigating the cross-covariance and the cross-correlation matrix between sphered and original variables allows to break the rotational invariance and to identify optimal whitening transformations. As a result we recommend two particular approaches: ZCA-cor whitening to produce sphered variables that are maximally similar to the original variables, and PCA-cor whitening to obtain sphered variables that maximally compress the original variables.
Introduction to Graphical Modelling
Jun 28, 2011



Abstract:The aim of this chapter is twofold. In the first part we will provide a brief overview of the mathematical and statistical foundations of graphical models, along with their fundamental properties, estimation and basic inference procedures. In particular we will develop Markov networks (also known as Markov random fields) and Bayesian networks, which comprise most past and current literature on graphical models. In the second part we will review some applications of graphical models in systems biology.
Feature selection in omics prediction problems using cat scores and false nondiscovery rate control
Oct 08, 2010
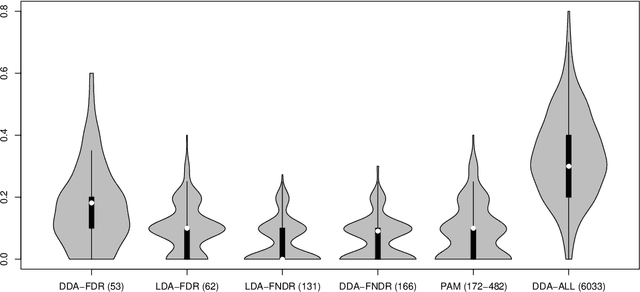


Abstract:We revisit the problem of feature selection in linear discriminant analysis (LDA), that is, when features are correlated. First, we introduce a pooled centroids formulation of the multiclass LDA predictor function, in which the relative weights of Mahalanobis-transformed predictors are given by correlation-adjusted $t$-scores (cat scores). Second, for feature selection we propose thresholding cat scores by controlling false nondiscovery rates (FNDR). Third, training of the classifier is based on James--Stein shrinkage estimates of correlations and variances, where regularization parameters are chosen analytically without resampling. Overall, this results in an effective and computationally inexpensive framework for high-dimensional prediction with natural feature selection. The proposed shrinkage discriminant procedures are implemented in the R package ``sda'' available from the R repository CRAN.
* Published in at http://dx.doi.org/10.1214/09-AOAS277 the Annals of Applied Statistics (http://www.imstat.org/aoas/) by the Institute of Mathematical Statistics (http://www.imstat.org)
Entropy inference and the James-Stein estimator, with application to nonlinear gene association networks
Jul 22, 2009



Abstract:We present a procedure for effective estimation of entropy and mutual information from small-sample data, and apply it to the problem of inferring high-dimensional gene association networks. Specifically, we develop a James-Stein-type shrinkage estimator, resulting in a procedure that is highly efficient statistically as well as computationally. Despite its simplicity, we show that it outperforms eight other entropy estimation procedures across a diverse range of sampling scenarios and data-generating models, even in cases of severe undersampling. We illustrate the approach by analyzing E. coli gene expression data and computing an entropy-based gene-association network from gene expression data. A computer program is available that implements the proposed shrinkage estimator.
* 18 pages, 3 figures, 1 table
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge