Kevin M. Carter
An Information Geometric Framework for Dimensionality Reduction
Sep 29, 2008Abstract:This report concerns the problem of dimensionality reduction through information geometric methods on statistical manifolds. While there has been considerable work recently presented regarding dimensionality reduction for the purposes of learning tasks such as classification, clustering, and visualization, these methods have focused primarily on Riemannian manifolds in Euclidean space. While sufficient for many applications, there are many high-dimensional signals which have no straightforward and meaningful Euclidean representation. In these cases, signals may be more appropriately represented as a realization of some distribution lying on a statistical manifold, or a manifold of probability density functions (PDFs). We present a framework for dimensionality reduction that uses information geometry for both statistical manifold reconstruction as well as dimensionality reduction in the data domain.
Information Preserving Component Analysis: Data Projections for Flow Cytometry Analysis
Apr 17, 2008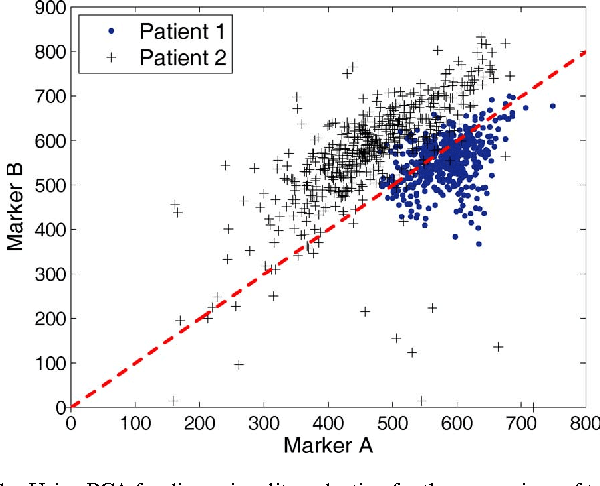
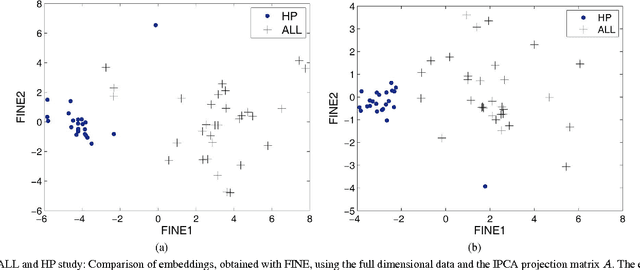
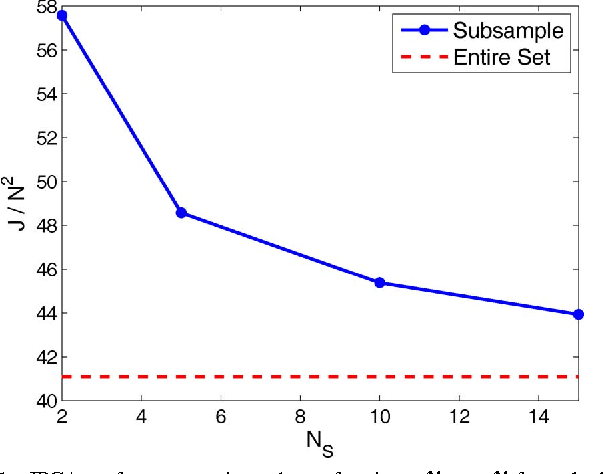
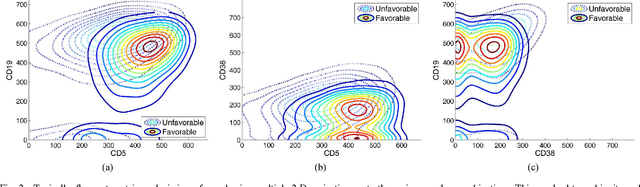
Abstract:Flow cytometry is often used to characterize the malignant cells in leukemia and lymphoma patients, traced to the level of the individual cell. Typically, flow cytometric data analysis is performed through a series of 2-dimensional projections onto the axes of the data set. Through the years, clinicians have determined combinations of different fluorescent markers which generate relatively known expression patterns for specific subtypes of leukemia and lymphoma -- cancers of the hematopoietic system. By only viewing a series of 2-dimensional projections, the high-dimensional nature of the data is rarely exploited. In this paper we present a means of determining a low-dimensional projection which maintains the high-dimensional relationships (i.e. information) between differing oncological data sets. By using machine learning techniques, we allow clinicians to visualize data in a low dimension defined by a linear combination of all of the available markers, rather than just 2 at a time. This provides an aid in diagnosing similar forms of cancer, as well as a means for variable selection in exploratory flow cytometric research. We refer to our method as Information Preserving Component Analysis (IPCA).
FINE: Fisher Information Non-parametric Embedding
Feb 14, 2008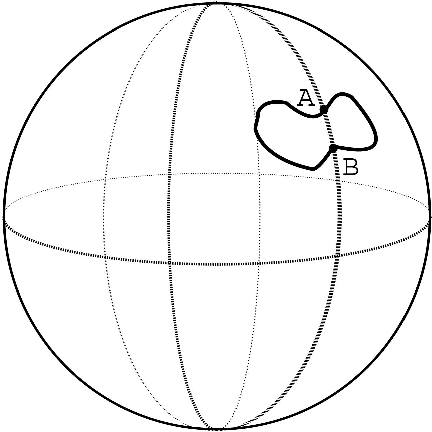
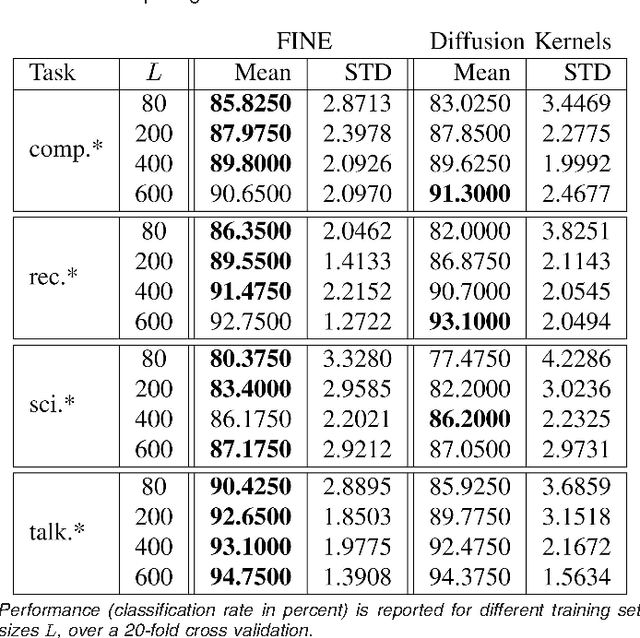
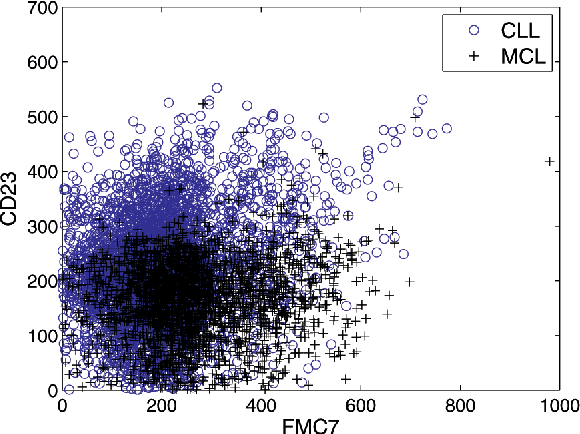
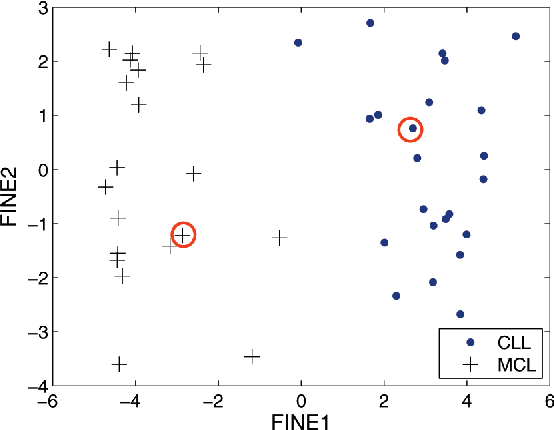
Abstract:We consider the problems of clustering, classification, and visualization of high-dimensional data when no straightforward Euclidean representation exists. Typically, these tasks are performed by first reducing the high-dimensional data to some lower dimensional Euclidean space, as many manifold learning methods have been developed for this task. In many practical problems however, the assumption of a Euclidean manifold cannot be justified. In these cases, a more appropriate assumption would be that the data lies on a statistical manifold, or a manifold of probability density functions (PDFs). In this paper we propose using the properties of information geometry in order to define similarities between data sets using the Fisher information metric. We will show this metric can be approximated using entirely non-parametric methods, as the parameterization of the manifold is generally unknown. Furthermore, by using multi-dimensional scaling methods, we are able to embed the corresponding PDFs into a low-dimensional Euclidean space. This not only allows for classification of the data, but also visualization of the manifold. As a whole, we refer to our framework as Fisher Information Non-parametric Embedding (FINE), and illustrate its uses on a variety of practical problems, including bio-medical applications and document classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge