Kensuke Tanioka
Federated Rule Ensemble Method in Medical Data
Apr 20, 2026Abstract:Machine learning has become integral to medical research and is increasingly applied in clinical settings to support diagnosis and decision-making; however, its effectiveness depends on access to large, diverse datasets, which are limited within single institutions. Although integrating data across institutions can address this limitation, privacy regulations and data ownership constraints hinder these efforts. Federated learning enables collaborative model training without sharing raw data; however, most methods rely on complex architectures that lack interpretability, limiting clinical applicability. Therefore, we proposed a federated RuleFit framework to construct a unified and interpretable global model for distributed environments. It integrates three components: preprocessing based on differentially private histograms to estimate shared cutoff values, enabling consistent rule definitions and reducing heterogeneity across clients; local rule generation using gradient boosting decision trees with shared cutoffs; and coefficient estimation via $\ell_1$-regularized optimization using a Federated Dual Averaging algorithm for sparse and consistent variable selection. In simulation studies, the proposed method achieved a performance comparable to that of centralized RuleFit while outperforming existing federated approaches. Real-world analysis demonstrated its ability to provide interpretable insights with competitive predictive accuracy. Therefore, the proposed framework offers a practical and effective solution for interpretable and reliable modeling in federated learning environments.
Causal rule ensemble approach for multi-arm data
Apr 24, 2025Abstract:Heterogeneous treatment effect (HTE) estimation is critical in medical research. It provides insights into how treatment effects vary among individuals, which can provide statistical evidence for precision medicine. While most existing methods focus on binary treatment situations, real-world applications often involve multiple interventions. However, current HTE estimation methods are primarily designed for binary comparisons and often rely on black-box models, which limit their applicability and interpretability in multi-arm settings. To address these challenges, we propose an interpretable machine learning framework for HTE estimation in multi-arm trials. Our method employs a rule-based ensemble approach consisting of rule generation, rule ensemble, and HTE estimation, ensuring both predictive accuracy and interpretability. Through extensive simulation studies and real data applications, the performance of our method was evaluated against state-of-the-art multi-arm HTE estimation approaches. The results indicate that our approach achieved lower bias and higher estimation accuracy compared with those of existing methods. Furthermore, the interpretability of our framework allows clearer insights into how covariates influence treatment effects, facilitating clinical decision making. By bridging the gap between accuracy and interpretability, our study contributes a valuable tool for multi-arm HTE estimation, supporting precision medicine.
Deep-learning models in medical image analysis: Detection of esophagitis from the Kvasir Dataset
Jan 06, 2023


Abstract:Early detection of esophagitis is important because this condition can progress to cancer if left untreated. However, the accuracies of different deep learning models in detecting esophagitis have yet to be compared. Thus, this study aimed to compare the accuracies of convolutional neural network models (GoogLeNet, ResNet-50, MobileNet V2, and MobileNet V3) in detecting esophagitis from the open Kvasir dataset of endoscopic images. Results showed that among the models, GoogLeNet achieved the highest F1-scores. Based on the average of true positive rate, MobileNet V3 predicted esophagitis more confidently than the other models. The results obtained using the models were also compared with those obtained using SHapley Additive exPlanations and Gradient-weighted Class Activation Mapping.
Performance Comparison of Deep Learning Architectures for Artifact Removal in Gastrointestinal Endoscopic Imaging
Jan 01, 2022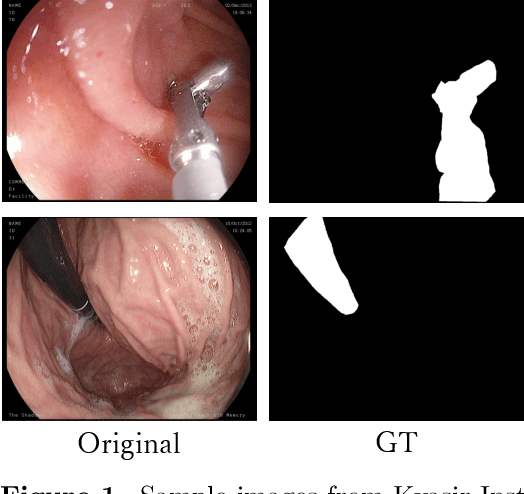
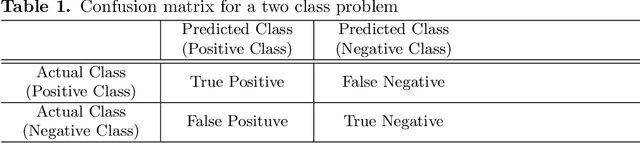
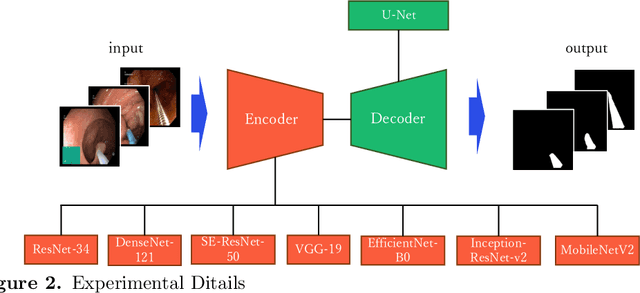
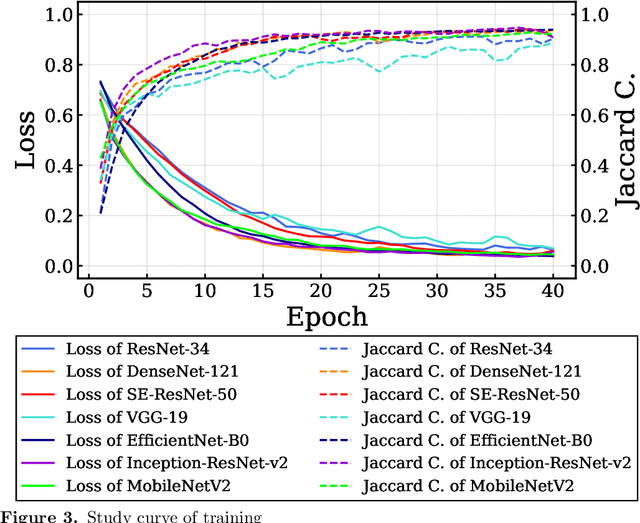
Abstract:Endoscopic images typically contain several artifacts. The artifacts significantly impact image analysis result in computer-aided diagnosis. Convolutional neural networks (CNNs), a type of deep learning, can removes such artifacts. Various architectures have been proposed for the CNNs, and the accuracy of artifact removal varies depending on the choice of architecture. Therefore, it is necessary to determine the artifact removal accuracy, depending on the selected architecture. In this study, we focus on endoscopic surgical instruments as artifacts, and determine and discuss the artifact removal accuracy using seven different CNN architectures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge