Kaya Turgut
PLANesT-3D: A new annotated dataset for segmentation of 3D plant point clouds
Jul 30, 2024



Abstract:Creation of new annotated public datasets is crucial in helping advances in 3D computer vision and machine learning meet their full potential for automatic interpretation of 3D plant models. In this paper, we introduce PLANesT-3D; a new annotated dataset of 3D color point clouds of plants. PLANesT-3D is composed of 34 point cloud models representing 34 real plants from three different plant species: \textit{Capsicum annuum}, \textit{Rosa kordana}, and \textit{Ribes rubrum}. Both semantic labels in terms of "leaf" and "stem", and organ instance labels were manually annotated for the full point clouds. As an additional contribution, SP-LSCnet, a novel semantic segmentation method that is a combination of unsupervised superpoint extraction and a 3D point-based deep learning approach is introduced and evaluated on the new dataset. Two existing deep neural network architectures, PointNet++ and RoseSegNet were also tested on the point clouds of PLANesT-3D for semantic segmentation.
Star+: A New Multi-Domain Model for CTR Prediction
Jun 24, 2024



Abstract:In this paper, we introduce Star+, a novel multi-domain model for click-through rate (CTR) prediction inspired by the Star model. Traditional single-domain approaches and existing multi-task learning techniques face challenges in multi-domain environments due to their inability to capture domain-specific data distributions and complex inter-domain relationships. Star+ addresses these limitations by enhancing the interaction between shared and domain-specific information through various fusion strategies, such as add, adaptive add, concatenation, and gating fusions, to find the optimal balance between domain-specific and shared information. We also investigate the impact of different normalization techniques, including layer normalization, batch normalization, and partition normalization, on the performance of our model. Our extensive experiments on both industrial and public datasets demonstrate that Star+ significantly improves prediction accuracy and efficiency. This work contributes to the advancement of recommendation systems by providing a robust, scalable, and adaptive solution for multi-domain environments.
Local region-learning modules for point cloud classification
Mar 30, 2023Abstract:Data organization via forming local regions is an integral part of deep learning networks that process 3D point clouds in a hierarchical manner. At each level, the point cloud is sampled to extract representative points and these points are used to be centers of local regions. The organization of local regions is of considerable importance since it determines the location and size of the receptive field at a particular layer of feature aggregation. In this paper, we present two local region-learning modules: Center Shift Module to infer the appropriate shift for each center point, and Radius Update Module to alter the radius of each local region. The parameters of the modules are learned through optimizing the loss associated with the particular task within an end-to-end network. We present alternatives for these modules through various ways of modeling the interactions of the features and locations of 3D points in the point cloud. We integrated both modules independently and together to the PointNet++ object classification architecture, and demonstrated that the modules contributed to a significant increase in classification accuracy for the ScanObjectNN data set.
Segmentation of structural parts of rosebush plants with 3D point-based deep learning methods
Dec 21, 2020
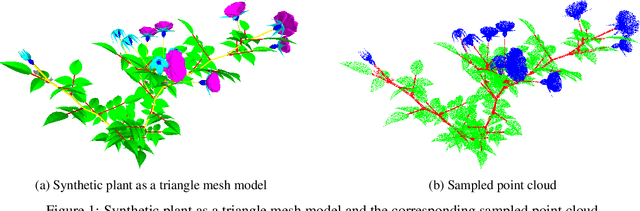
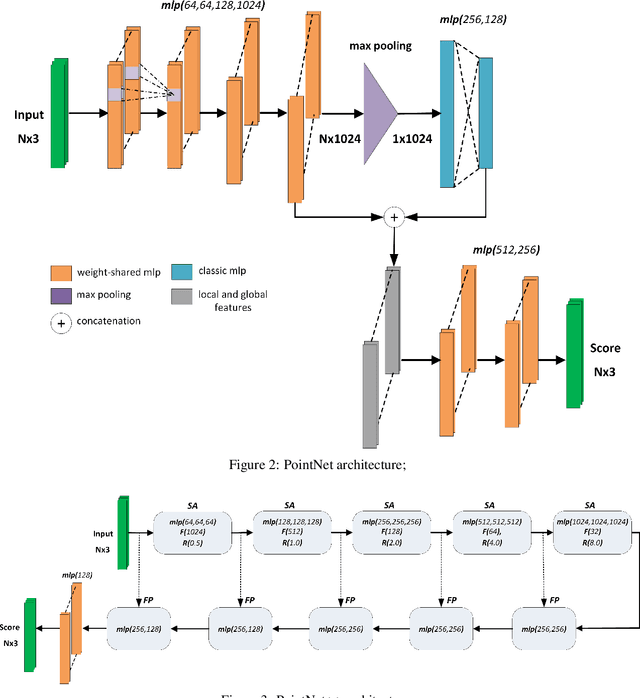
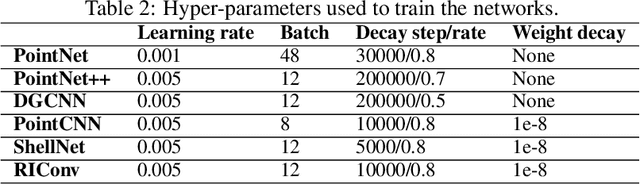
Abstract:Segmentation of structural parts of 3D models of plants is an important step for plant phenotyping, especially for monitoring architectural and morphological traits. This work introduces a benchmark for assessing the performance of 3D point-based deep learning methods on organ segmentation of 3D plant models, specifically rosebush models. Six recent deep learning architectures that segment 3D point clouds into semantic parts were adapted and compared. The methods were tested on the ROSE-X data set, containing fully annotated 3D models of real rosebush plants. The contribution of incorporating synthetic 3D models generated through Lindenmayer systems into training data was also investigated.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge