Karol Mikula
A Graph Diffusion Algorithm for Lexical Similarity Evaluation
Apr 09, 2025Abstract:In this paper, we present an algorithm for evaluating lexical similarity between a given language and several reference language clusters. As an input, we have a list of concepts and the corresponding translations in all considered languages. Moreover, each reference language is assigned to one of $c$ language clusters. For each of the concepts, the algorithm computes the distance between each pair of translations. Based on these distances, it constructs a weighted directed graph, where every vertex represents a language. After, it solves a graph diffusion equation with a Dirichlet boundary condition, where the unknown is a map from the vertex set to $\mathbb{R}^c$. The resulting coordinates are values from the interval $[0,1]$ and they can be interpreted as probabilities of belonging to each of the clusters or as a lexical similarity distribution with respect to the reference clusters. The distances between translations are calculated using phonetic transcriptions and a modification of the Damerau-Levenshtein distance. The algorithm can be useful in analyzing relationships between languages spoken in multilingual territories with a lot of mutual influences. We demonstrate this by presenting a case study regarding various European languages.
Segmentation based tracking of cells in 2D+time microscopy images of macrophages
Jan 02, 2023Abstract:The automated segmentation and tracking of macrophages during their migration are challenging tasks due to their dynamically changing shapes and motions. This paper proposes a new algorithm to achieve automatic cell tracking in time-lapse microscopy macrophage data. First, we design a segmentation method employing space-time filtering, local Otsu's thresholding, and the SUBSURF (subjective surface segmentation) method. Next, the partial trajectories for cells overlapping in the temporal direction are extracted in the segmented images. Finally, the extracted trajectories are linked by considering their direction of movement. The segmented images and the obtained trajectories from the proposed method are compared with those of the semi-automatic segmentation and manual tracking. The proposed tracking achieved 97.4% of accuracy for macrophage data under challenging situations, feeble fluorescent intensity, irregular shapes, and motion of macrophages. We expect that the automatically extracted trajectories of macrophages can provide pieces of evidence of how macrophages migrate depending on their polarization modes in the situation, such as during wound healing.
Direct simple computation of middle surface between 3D point clouds and/or discrete surfaces by tracking sources in distance function calculation algorithms
Dec 17, 2021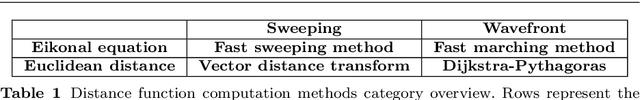
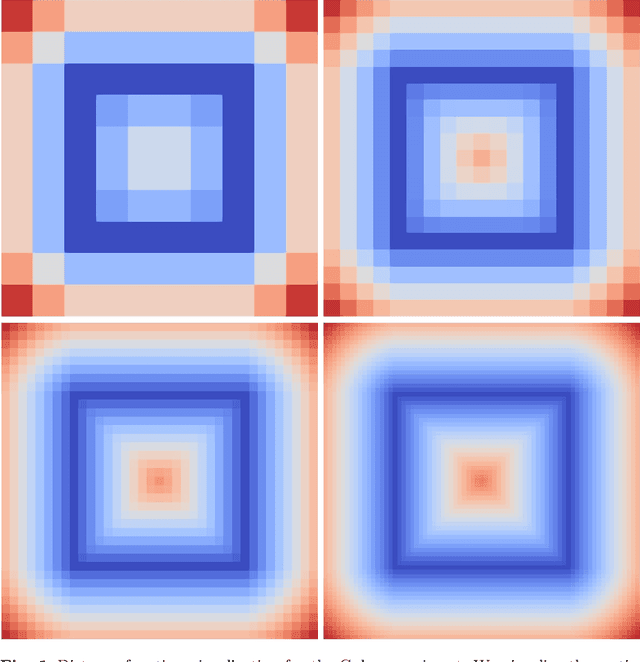
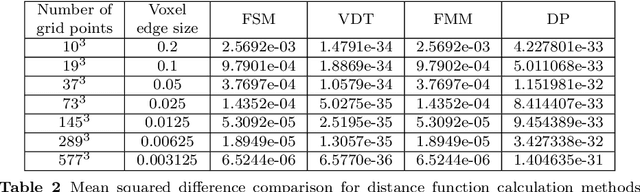
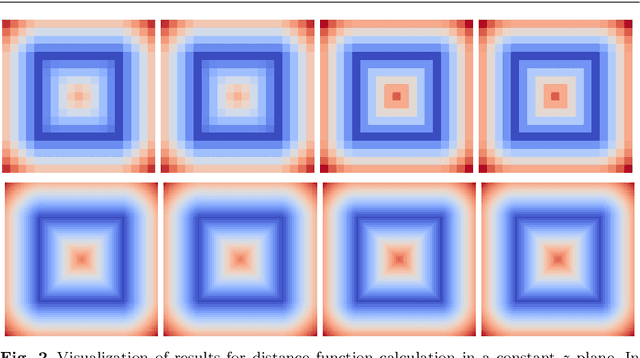
Abstract:In this paper, we introduce novel methods for computing middle surfaces between various 3D data sets such as point clouds and/or discrete surfaces. Traditionally the middle surface is obtained by detecting singularities in computed distance function such as ridges, triple junctions, etc. It requires to compute second order differential characteristics and also some kinds of heuristics must be applied. Opposite to that, we determine the middle surface just from computing the distance function itself which is a fast and simple approach. We present and compare the results of the fast sweeping method, the vector distance transform algorithm, the fast marching method, and the Dijkstra-Pythagoras method in finding the middle surface between 3D data sets.
Natural Numerical Networks for Natura 2000 habitats classification by satellite images
Aug 09, 2021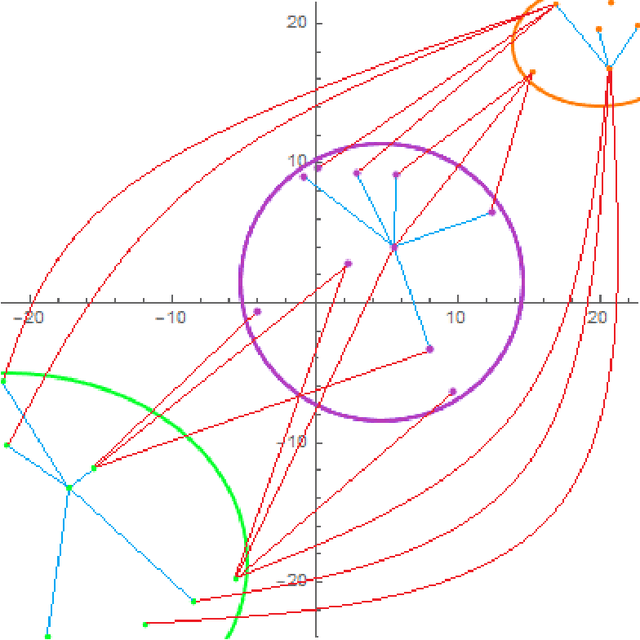

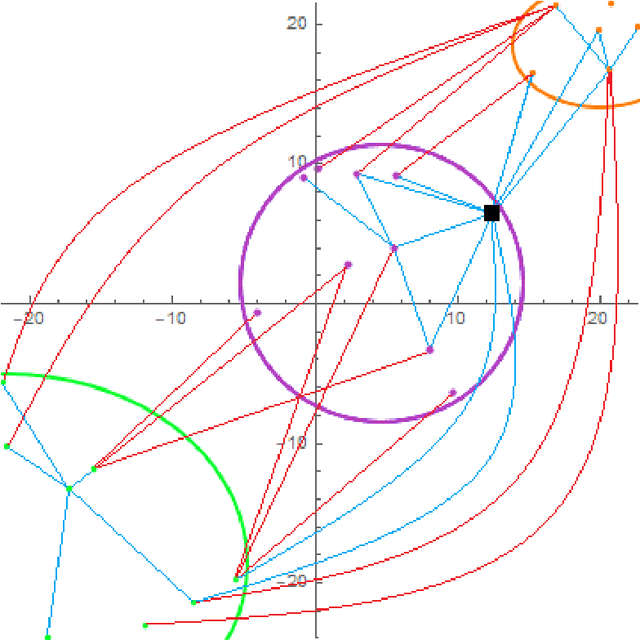

Abstract:Natural numerical networks are introduced as a new classification algorithm based on the numerical solution of nonlinear partial differential equations of forward-backward diffusion type on complete graphs. The proposed natural numerical network is applied to open important environmental and nature conservation task, the automated identification of protected habitats by using satellite images. In the natural numerical network, the forward diffusion causes the movement of points in a feature space toward each other. The opposite effect, keeping the points away from each other, is caused by backward diffusion. This yields the desired classification. The natural numerical network contains a few parameters that are optimized in the learning phase of the method. After learning parameters and optimizing the topology of the network graph, classification necessary for habitat identification is performed. A relevancy map for each habitat is introduced as a tool for validating the classification and finding new Natura 2000 habitat appearances.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge