Julien Grosjean
A randomized simulation trial evaluating ABiMed, a clinical decision support system for medication reviews and polypharmacy management
Sep 03, 2024



Abstract:Background: Medication review is a structured interview of the patient, performed by the pharmacist and aimed at optimizing drug treatments. In practice, medication review is a long and cognitively-demanding task that requires specific knowledge. Clinical practice guidelines have been proposed, but their application is tedious. Methods: We designed ABiMed, a clinical decision support system for medication reviews, based on the implementation of the STOPP/START v2 guidelines and on the visual presentation of aggregated drug knowledge using tables, graphs and flower glyphs. We evaluated ABiMed with 39 community pharmacists during a randomized simulation trial, each pharmacist performing a medication review for two fictitious patients without ABiMed, and two others with ABiMed. We recorded the problems identified by the pharmacists, the interventions proposed, the response time, the perceived usability and the comments. Pharmacists' medication reviews were compared to an expert-designed gold standard. Results: With ABiMed, pharmacists found 1.6 times more relevant drug-related problems during the medication review (p=1.1e-12) and proposed better interventions (p=9.8e-9), without needing more time (p=0.56). The System Usability Scale score is 82.7, which is ranked "excellent". In their comments, pharmacists appreciated the visual aspect of ABiMed and its ability to compare the current treatment with the proposed one. A multifactor analysis showed no difference in the support offered by ABiMed according to the pharmacist's age or sex, in terms of percentage of problems identified or quality of the proposed interventions. Conclusions: The use of an intelligent and visual clinical decision support system can help pharmacists when they perform medication reviews. Our main perspective is the validation of the system in clinical conditions.
ABiMed: An intelligent and visual clinical decision support system for medication reviews and polypharmacy management
Dec 13, 2023Abstract:Background: Polypharmacy, i.e. taking five drugs or more, is both a public health and an economic issue. Medication reviews are structured interviews of the patient by the community pharmacist, aiming at optimizing the drug treatment and deprescribing useless, redundant or dangerous drugs. However, they remain difficult to perform and time-consuming. Several clinical decision support systems were developed for helping clinicians to manage polypharmacy. However, most were limited to the implementation of clinical practice guidelines. In this work, our objective is to design an innovative clinical decision support system for medication reviews and polypharmacy management, named ABiMed. Methods: ABiMed associates several approaches: guidelines implementation, but the automatic extraction of patient data from the GP's electronic health record and its transfer to the pharmacist, and the visual presentation of contextualized drug knowledge using visual analytics. We performed an ergonomic assessment and qualitative evaluations involving pharmacists and GPs during focus groups and workshops. Results: We describe the proposed architecture, which allows a collaborative multi-user usage. We present the various screens of ABiMed for entering or verifying patient data, for accessing drug knowledge (posology, adverse effects, interactions), for viewing STOPP/START rules and for suggesting modification to the treatment. Qualitative evaluations showed that health professionals were highly interested by our approach, associating the automatic guidelines execution with the visual presentation of drug knowledge. Conclusions: The association of guidelines implementation with visual presentation of knowledge is a promising approach for managing polypharmacy. Future works will focus on the improvement and the evaluation of ABiMed.
Doc2Vec on the PubMed corpus: study of a new approach to generate related articles
Nov 26, 2019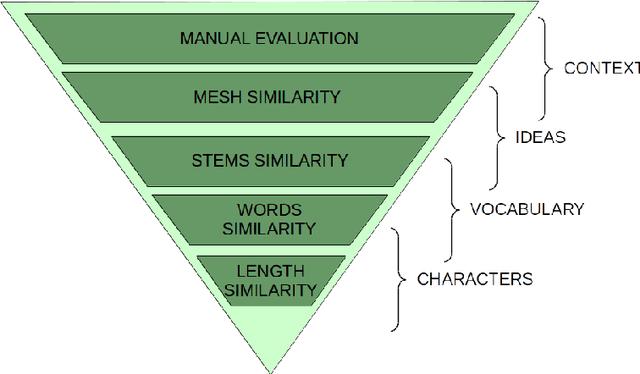
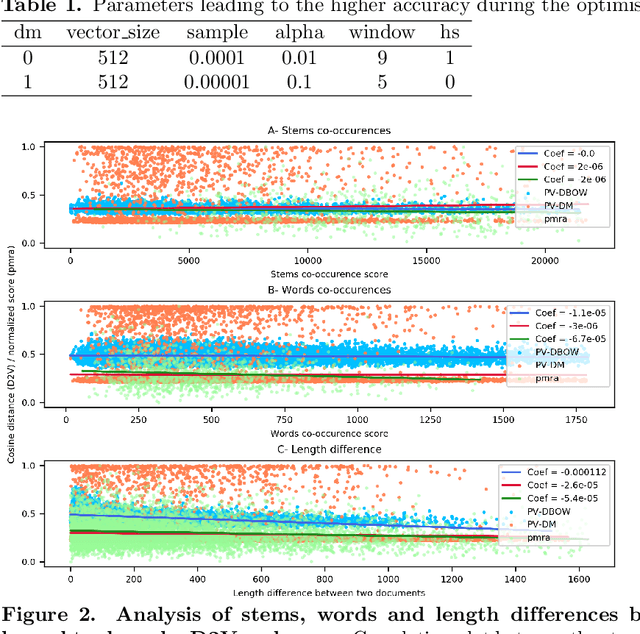
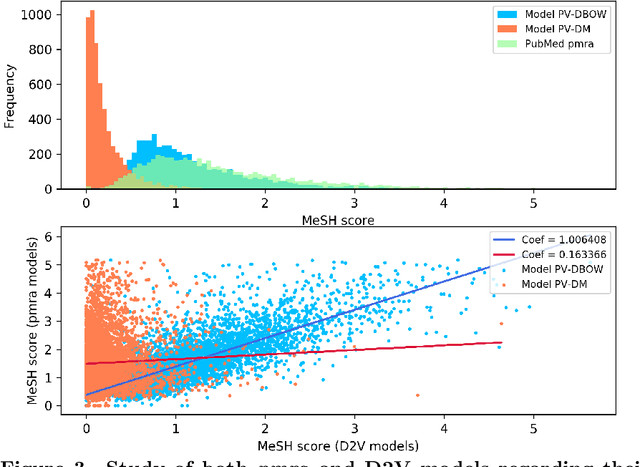
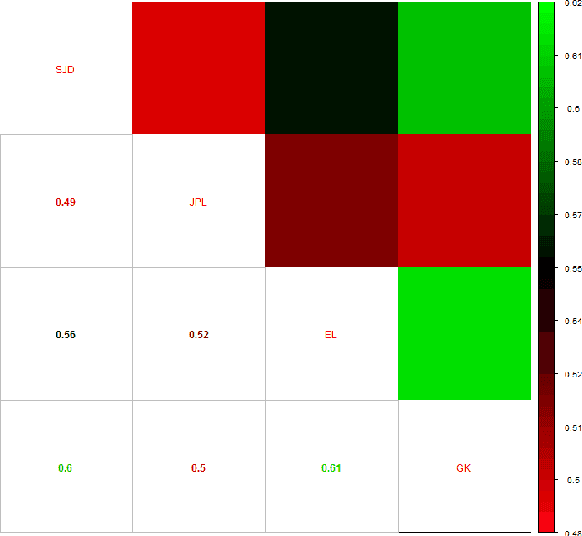
Abstract:PubMed is the biggest and most used bibliographic database worldwide, hosting more than 26M biomedical publications. One of its useful features is the "similar articles" section, allowing the end-user to find scientific articles linked to the consulted document in term of context. The aim of this study is to analyze whether it is possible to replace the statistic model PubMed Related Articles (pmra) with a document embedding method. Doc2Vec algorithm was used to train models allowing to vectorize documents. Six of its parameters were optimised by following a grid-search strategy to train more than 1,900 models. Parameters combination leading to the best accuracy was used to train models on abstracts from the PubMed database. Four evaluations tasks were defined to determine what does or does not influence the proximity between documents for both Doc2Vec and pmra. The two different Doc2Vec architectures have different abilities to link documents about a common context. The terminological indexing, words and stems contents of linked documents are highly similar between pmra and Doc2Vec PV-DBOW architecture. These algorithms are also more likely to bring closer documents having a similar size. In contrary, the manual evaluation shows much better results for the pmra algorithm. While the pmra algorithm links documents by explicitly using terminological indexing in its formula, Doc2Vec does not need a prior indexing. It can infer relations between documents sharing a similar indexing, without any knowledge about them, particularly regarding the PV-DBOW architecture. In contrary, the human evaluation, without any clear agreement between evaluators, implies future studies to better understand this difference between PV-DBOW and pmra algorithm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge