Jose Dolz
École de Technologie Supérieure, Montreal, Canada
Class Adaptive Network Calibration
Nov 28, 2022



Abstract:Recent studies have revealed that, beyond conventional accuracy, calibration should also be considered for training modern deep neural networks. To address miscalibration during learning, some methods have explored different penalty functions as part of the learning objective, alongside a standard classification loss, with a hyper-parameter controlling the relative contribution of each term. Nevertheless, these methods share two major drawbacks: 1) the scalar balancing weight is the same for all classes, hindering the ability to address different intrinsic difficulties or imbalance among classes; and 2) the balancing weight is usually fixed without an adaptive strategy, which may prevent from reaching the best compromise between accuracy and calibration, and requires hyper-parameter search for each application. We propose Class Adaptive Label Smoothing (CALS) for calibrating deep networks, which allows to learn class-wise multipliers during training, yielding a powerful alternative to common label smoothing penalties. Our method builds on a general Augmented Lagrangian approach, a well-established technique in constrained optimization, but we introduce several modifications to tailor it for large-scale, class-adaptive training. Comprehensive evaluation and multiple comparisons on a variety of benchmarks, including standard and long-tailed image classification, semantic segmentation, and text classification, demonstrate the superiority of the proposed method. The code is available at https://github.com/by-liu/CALS.
A Strong Baseline for Generalized Few-Shot Semantic Segmentation
Nov 25, 2022Abstract:This paper introduces a generalized few-shot segmentation framework with a straightforward training process and an easy-to-optimize inference phase. In particular, we propose a simple yet effective model based on the well-known InfoMax principle, where the Mutual Information (MI) between the learned feature representations and their corresponding predictions is maximized. In addition, the terms derived from our MI-based formulation are coupled with a knowledge distillation term to retain the knowledge on base classes. With a simple training process, our inference model can be applied on top of any segmentation network trained on base classes. The proposed inference yields substantial improvements on the popular few-shot segmentation benchmarks PASCAL-$5^i$ and COCO-$20^i$. Particularly, for novel classes, the improvement gains range from 5% to 20% (PASCAL-$5^i$) and from 2.5% to 10.5% (COCO-$20^i$) in the 1-shot and 5-shot scenarios, respectively. Furthermore, we propose a more challenging setting, where performance gaps are further exacerbated. Our code is publicly available at https://github.com/sinahmr/DIaM.
Calibrating Segmentation Networks with Margin-based Label Smoothing
Sep 09, 2022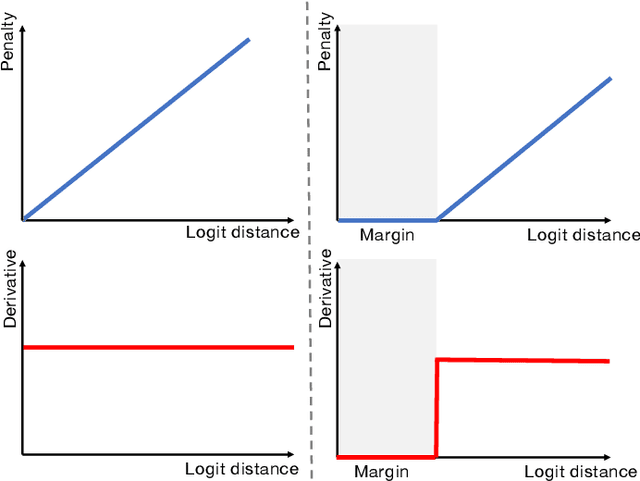
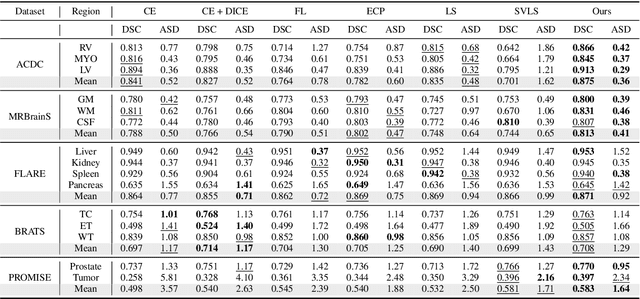
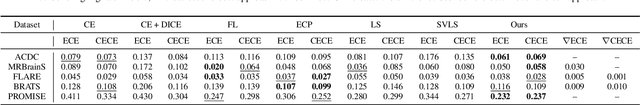
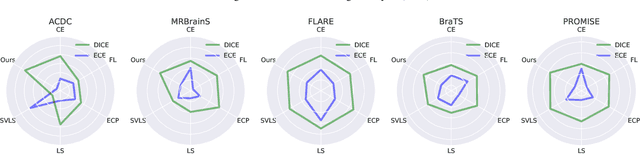
Abstract:Despite the undeniable progress in visual recognition tasks fueled by deep neural networks, there exists recent evidence showing that these models are poorly calibrated, resulting in over-confident predictions. The standard practices of minimizing the cross entropy loss during training promote the predicted softmax probabilities to match the one-hot label assignments. Nevertheless, this yields a pre-softmax activation of the correct class that is significantly larger than the remaining activations, which exacerbates the miscalibration problem. Recent observations from the classification literature suggest that loss functions that embed implicit or explicit maximization of the entropy of predictions yield state-of-the-art calibration performances. Despite these findings, the impact of these losses in the relevant task of calibrating medical image segmentation networks remains unexplored. In this work, we provide a unifying constrained-optimization perspective of current state-of-the-art calibration losses. Specifically, these losses could be viewed as approximations of a linear penalty (or a Lagrangian term) imposing equality constraints on logit distances. This points to an important limitation of such underlying equality constraints, whose ensuing gradients constantly push towards a non-informative solution, which might prevent from reaching the best compromise between the discriminative performance and calibration of the model during gradient-based optimization. Following our observations, we propose a simple and flexible generalization based on inequality constraints, which imposes a controllable margin on logit distances. Comprehensive experiments on a variety of public medical image segmentation benchmarks demonstrate that our method sets novel state-of-the-art results on these tasks in terms of network calibration, whereas the discriminative performance is also improved.
Attention-based Dynamic Subspace Learners for Medical Image Analysis
Jun 18, 2022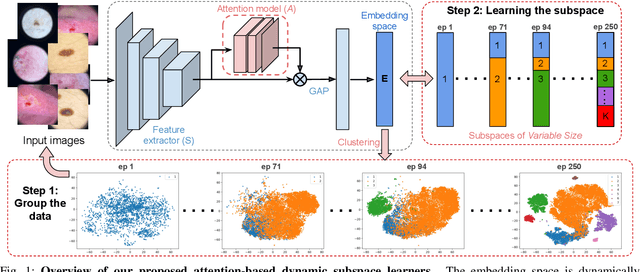
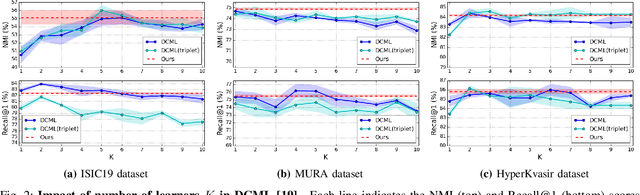
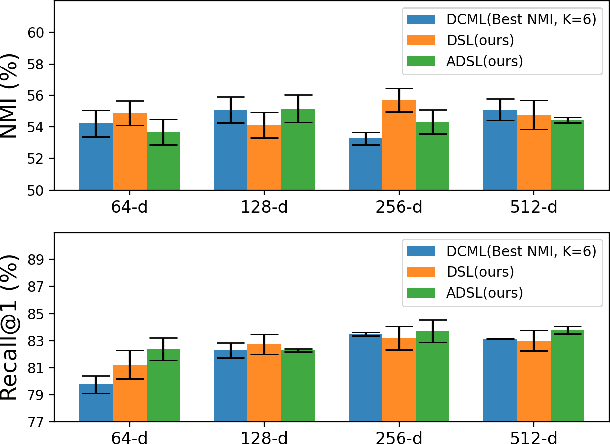
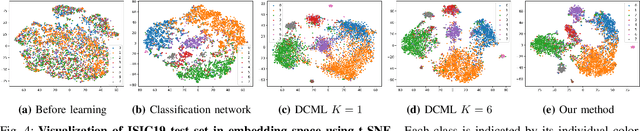
Abstract:Learning similarity is a key aspect in medical image analysis, particularly in recommendation systems or in uncovering the interpretation of anatomical data in images. Most existing methods learn such similarities in the embedding space over image sets using a single metric learner. Images, however, have a variety of object attributes such as color, shape, or artifacts. Encoding such attributes using a single metric learner is inadequate and may fail to generalize. Instead, multiple learners could focus on separate aspects of these attributes in subspaces of an overarching embedding. This, however, implies the number of learners to be found empirically for each new dataset. This work, Dynamic Subspace Learners, proposes to dynamically exploit multiple learners by removing the need of knowing apriori the number of learners and aggregating new subspace learners during training. Furthermore, the visual interpretability of such subspace learning is enforced by integrating an attention module into our method. This integrated attention mechanism provides a visual insight of discriminative image features that contribute to the clustering of image sets and a visual explanation of the embedding features. The benefits of our attention-based dynamic subspace learners are evaluated in the application of image clustering, image retrieval, and weakly supervised segmentation. Our method achieves competitive results with the performances of multiple learners baselines and significantly outperforms the classification network in terms of clustering and retrieval scores on three different public benchmark datasets. Moreover, our attention maps offer a proxy-labels, which improves the segmentation accuracy up to 15% in Dice scores when compared to state-of-the-art interpretation techniques.
Leveraging Uncertainty for Deep Interpretable Classification and Weakly-Supervised Segmentation of Histology Images
May 12, 2022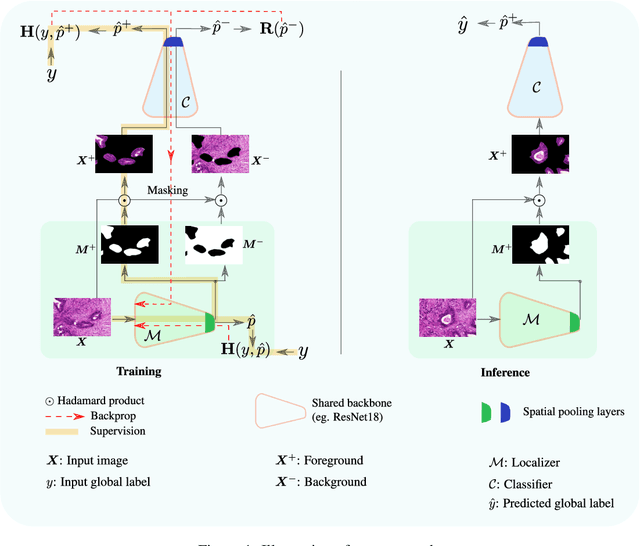
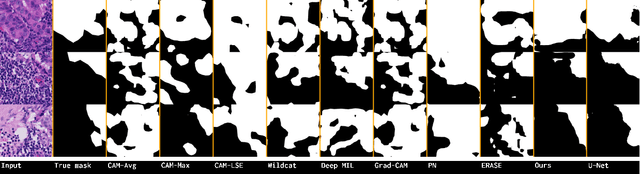
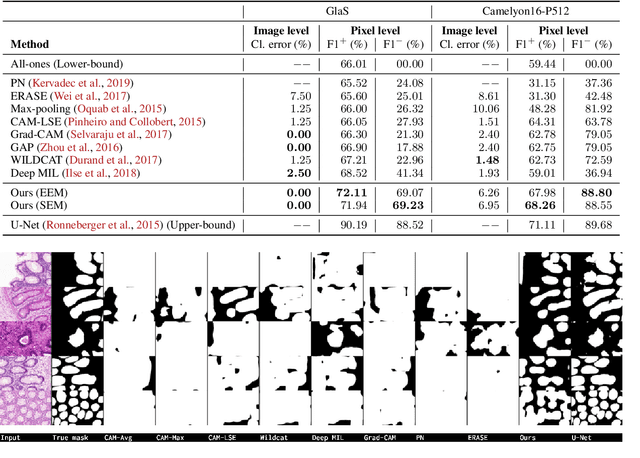


Abstract:Trained using only image class label, deep weakly supervised methods allow image classification and ROI segmentation for interpretability. Despite their success on natural images, they face several challenges over histology data where ROI are visually similar to background making models vulnerable to high pixel-wise false positives. These methods lack mechanisms for modeling explicitly non-discriminative regions which raises false-positive rates. We propose novel regularization terms, which enable the model to seek both non-discriminative and discriminative regions, while discouraging unbalanced segmentations and using only image class label. Our method is composed of two networks: a localizer that yields segmentation mask, followed by a classifier. The training loss pushes the localizer to build a segmentation mask that holds most discrimiantive regions while simultaneously modeling background regions. Comprehensive experiments over two histology datasets showed the merits of our method in reducing false positives and accurately segmenting ROI.
Leveraging Labeling Representations in Uncertainty-based Semi-supervised Segmentation
Mar 10, 2022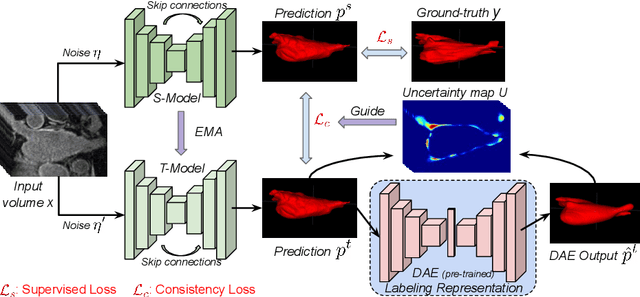
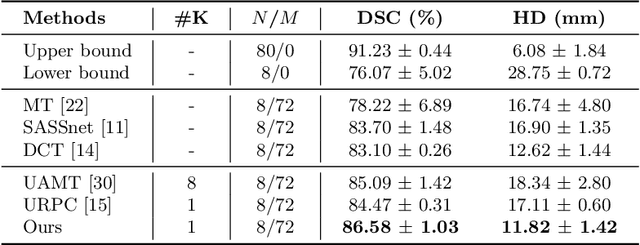
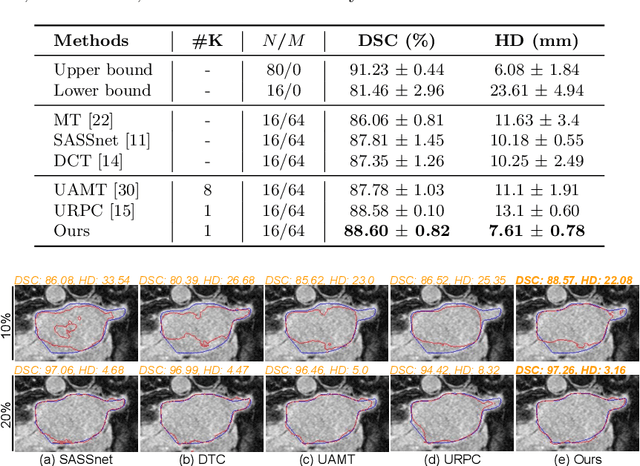
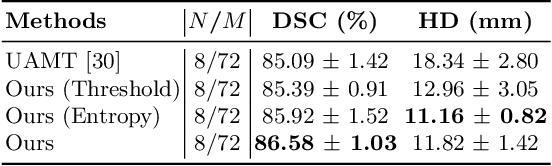
Abstract:Semi-supervised segmentation tackles the scarcity of annotations by leveraging unlabeled data with a small amount of labeled data. A prominent way to utilize the unlabeled data is by consistency training which commonly uses a teacher-student network, where a teacher guides a student segmentation. The predictions of unlabeled data are not reliable, therefore, uncertainty-aware methods have been proposed to gradually learn from meaningful and reliable predictions. Uncertainty estimation, however, relies on multiple inferences from model predictions that need to be computed for each training step, which is computationally expensive. This work proposes a novel method to estimate the pixel-level uncertainty by leveraging the labeling representation of segmentation masks. On the one hand, a labeling representation is learnt to represent the available segmentation masks. The learnt labeling representation is used to map the prediction of the segmentation into a set of plausible masks. Such a reconstructed segmentation mask aids in estimating the pixel-level uncertainty guiding the segmentation network. The proposed method estimates the uncertainty with a single inference from the labeling representation, thereby reducing the total computation. We evaluate our method on the 3D segmentation of left atrium in MRI, and we show that our uncertainty estimates from our labeling representation improve the segmentation accuracy over state-of-the-art methods.
On the pitfalls of entropy-based uncertainty for multi-class semi-supervised segmentation
Mar 07, 2022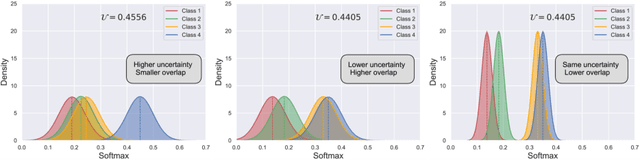
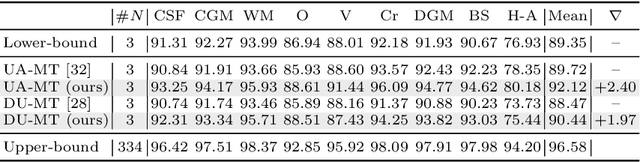
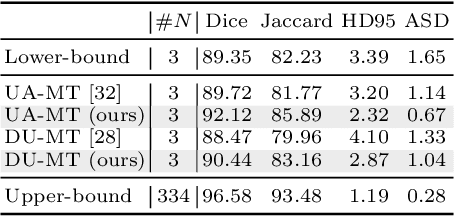
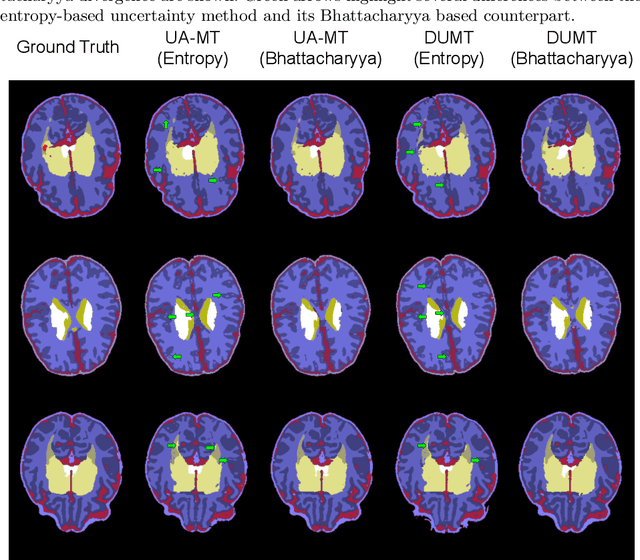
Abstract:Semi-supervised learning has emerged as an appealing strategy to train deep models with limited supervision. Most prior literature under this learning paradigm resorts to dual-based architectures, typically composed of a teacher-student duple. To drive the learning of the student, many of these models leverage the aleatoric uncertainty derived from the entropy of the predictions. While this has shown to work well in a binary scenario, we demonstrate in this work that this strategy leads to suboptimal results in a multi-class context, a more realistic and challenging setting. We argue, indeed, that these approaches underperform due to the erroneous uncertainty approximations in the presence of inter-class overlap. Furthermore, we propose an alternative solution to compute the uncertainty in a multi-class setting, based on divergence distances and which account for inter-class overlap. We evaluate the proposed solution on a challenging multi-class segmentation dataset and in two well-known uncertainty-based segmentation methods. The reported results demonstrate that by simply replacing the mechanism used to compute the uncertainty, our proposed solution brings substantial improvement on tested setups.
Constrained unsupervised anomaly segmentation
Mar 03, 2022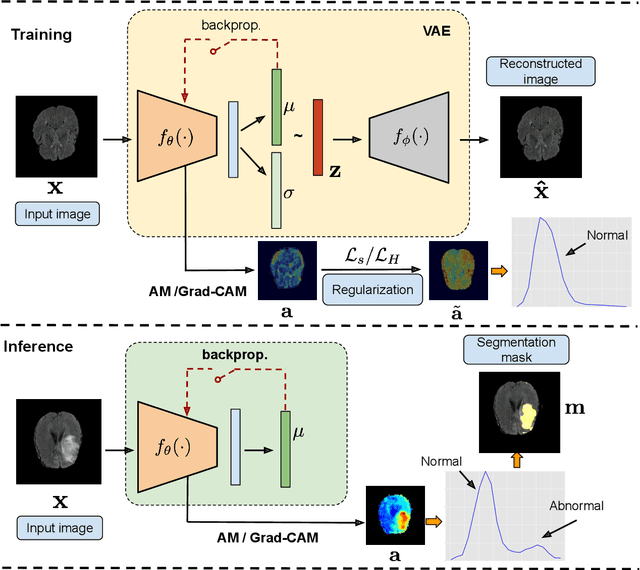
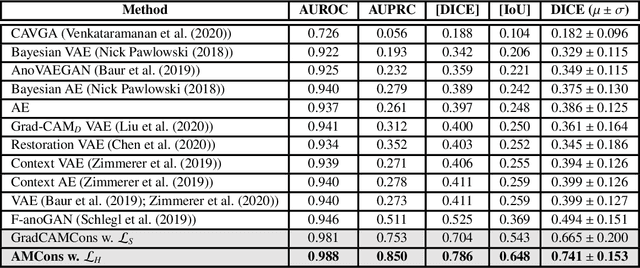
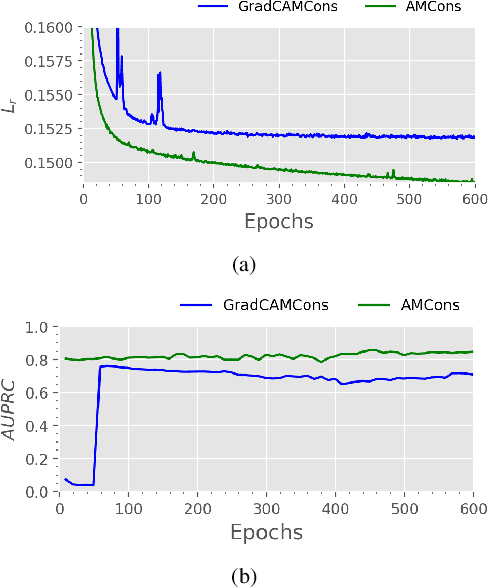
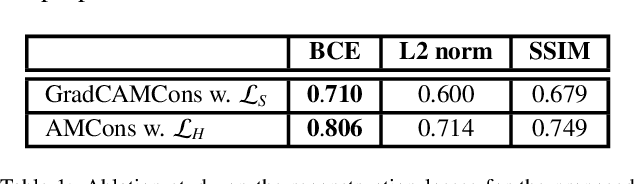
Abstract:Current unsupervised anomaly localization approaches rely on generative models to learn the distribution of normal images, which is later used to identify potential anomalous regions derived from errors on the reconstructed images. However, a main limitation of nearly all prior literature is the need of employing anomalous images to set a class-specific threshold to locate the anomalies. This limits their usability in realistic scenarios, where only normal data is typically accessible. Despite this major drawback, only a handful of works have addressed this limitation, by integrating supervision on attention maps during training. In this work, we propose a novel formulation that does not require accessing images with abnormalities to define the threshold. Furthermore, and in contrast to very recent work, the proposed constraint is formulated in a more principled manner, leveraging well-known knowledge in constrained optimization. In particular, the equality constraint on the attention maps in prior work is replaced by an inequality constraint, which allows more flexibility. In addition, to address the limitations of penalty-based functions we employ an extension of the popular log-barrier methods to handle the constraint. Last, we propose an alternative regularization term that maximizes the Shannon entropy of the attention maps, reducing the amount of hyperparameters of the proposed model. Comprehensive experiments on two publicly available datasets on brain lesion segmentation demonstrate that the proposed approach substantially outperforms relevant literature, establishing new state-of-the-art results for unsupervised lesion segmentation, and without the need to access anomalous images.
Maximum Entropy on Erroneous Predictions (MEEP): Improving model calibration for medical image segmentation
Dec 22, 2021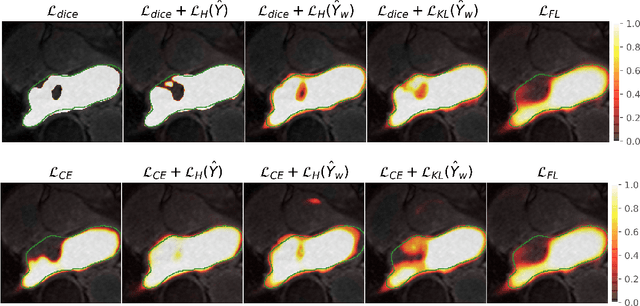
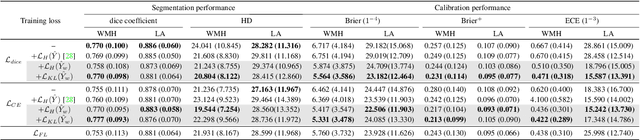
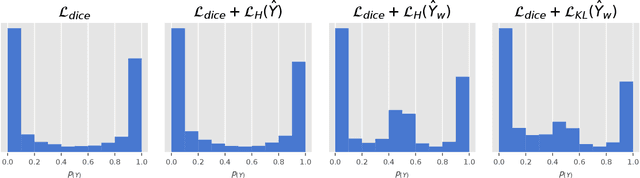
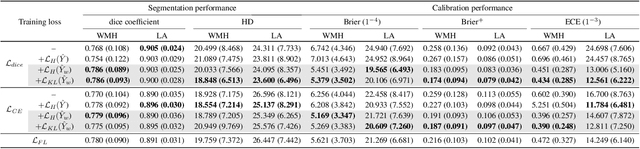
Abstract:Modern deep neural networks have achieved remarkable progress in medical image segmentation tasks. However, it has recently been observed that they tend to produce overconfident estimates, even in situations of high uncertainty, leading to poorly calibrated and unreliable models. In this work we introduce Maximum Entropy on Erroneous Predictions (MEEP), a training strategy for segmentation networks which selectively penalizes overconfident predictions, focusing only on misclassified pixels. In particular, we design a regularization term that encourages high entropy posteriors for wrong predictions, increasing the network uncertainty in complex scenarios. Our method is agnostic to the neural architecture, does not increase model complexity and can be coupled with multiple segmentation loss functions. We benchmark the proposed strategy in two challenging medical image segmentation tasks: white matter hyperintensity lesions in magnetic resonance images (MRI) of the brain, and atrial segmentation in cardiac MRI. The experimental results demonstrate that coupling MEEP with standard segmentation losses leads to improvements not only in terms of model calibration, but also in segmentation quality.
The Devil is in the Margin: Margin-based Label Smoothing for Network Calibration
Nov 30, 2021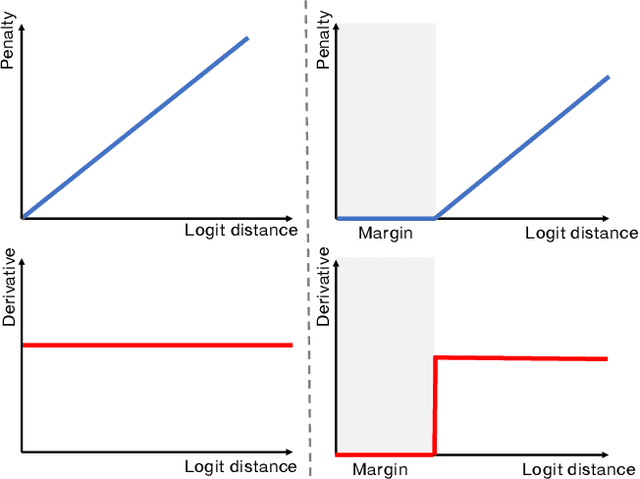
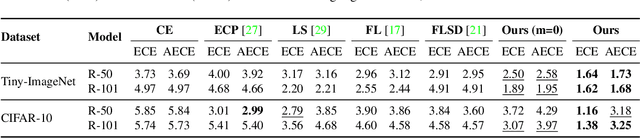
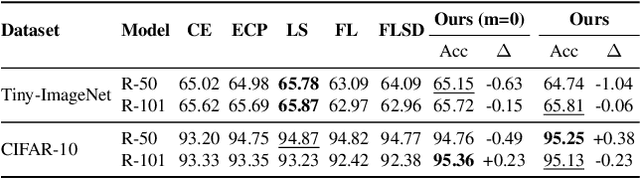
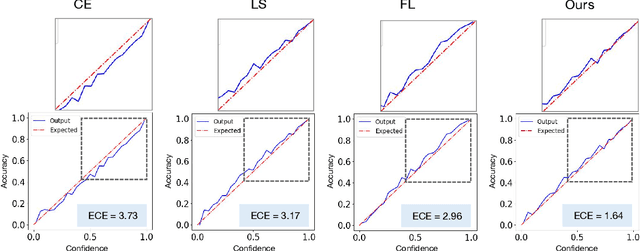
Abstract:In spite of the dominant performances of deep neural networks, recent works have shown that they are poorly calibrated, resulting in over-confident predictions. Miscalibration can be exacerbated by overfitting due to the minimization of the cross-entropy during training, as it promotes the predicted softmax probabilities to match the one-hot label assignments. This yields a pre-softmax activation of the correct class that is significantly larger than the remaining activations. Recent evidence from the literature suggests that loss functions that embed implicit or explicit maximization of the entropy of predictions yield state-of-the-art calibration performances. We provide a unifying constrained-optimization perspective of current state-of-the-art calibration losses. Specifically, these losses could be viewed as approximations of a linear penalty (or a Lagrangian) imposing equality constraints on logit distances. This points to an important limitation of such underlying equality constraints, whose ensuing gradients constantly push towards a non-informative solution, which might prevent from reaching the best compromise between the discriminative performance and calibration of the model during gradient-based optimization. Following our observations, we propose a simple and flexible generalization based on inequality constraints, which imposes a controllable margin on logit distances. Comprehensive experiments on a variety of image classification, semantic segmentation and NLP benchmarks demonstrate that our method sets novel state-of-the-art results on these tasks in terms of network calibration, without affecting the discriminative performance. The code is available at https://github.com/by-liu/MbLS .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge