John P. Cunningham
Predicting post-operative right ventricular failure using video-based deep learning
Feb 28, 2021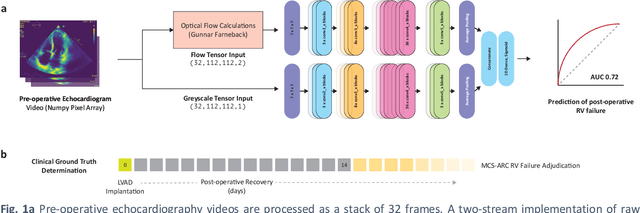
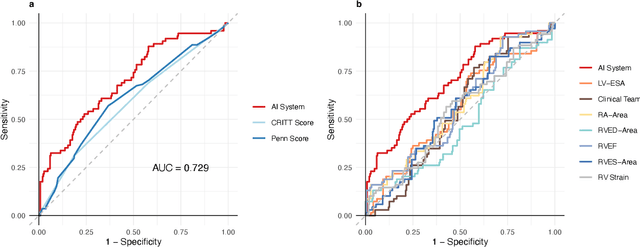
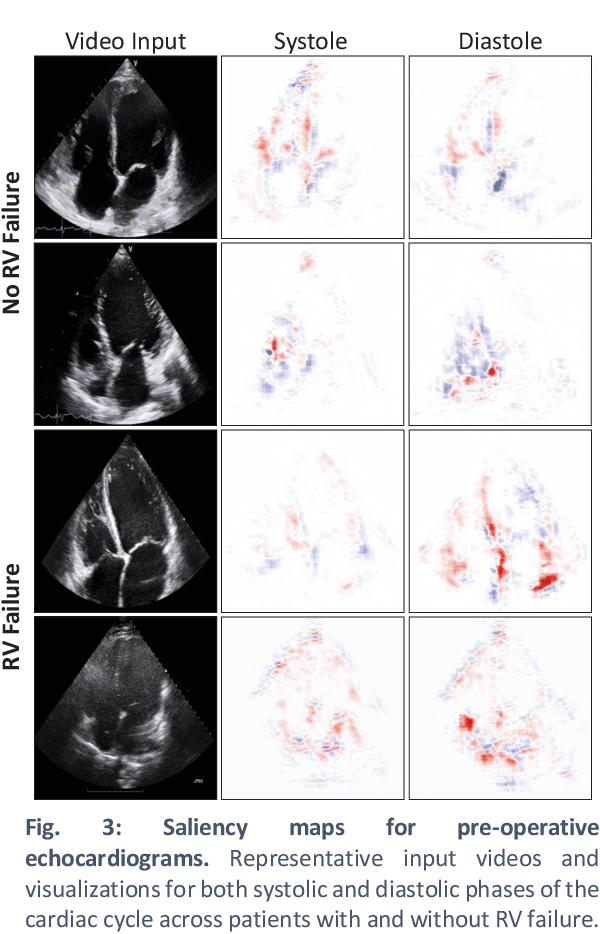
Abstract:Non-invasive and cost effective in nature, the echocardiogram allows for a comprehensive assessment of the cardiac musculature and valves. Despite progressive improvements over the decades, the rich temporally resolved data in echocardiography videos remain underutilized. Human reads of echocardiograms reduce the complex patterns of cardiac wall motion, to a small list of measurements of heart function. Furthermore, all modern echocardiography artificial intelligence (AI) systems are similarly limited by design - automating measurements of the same reductionist metrics rather than utilizing the wealth of data embedded within each echo study. This underutilization is most evident in situations where clinical decision making is guided by subjective assessments of disease acuity, and tools that predict disease onset within clinically actionable timeframes are unavailable. Predicting the likelihood of developing post-operative right ventricular failure (RV failure) in the setting of mechanical circulatory support is one such clinical example. To address this, we developed a novel video AI system trained to predict post-operative right ventricular failure (RV failure), using the full spatiotemporal density of information from pre-operative echocardiography scans. We achieve an AUC of 0.729, specificity of 52% at 80% sensitivity and 46% sensitivity at 80% specificity. Furthermore, we show that our ML system significantly outperforms a team of human experts tasked with predicting RV failure on independent clinical evaluation. Finally, the methods we describe are generalizable to any cardiac clinical decision support application where treatment or patient selection is guided by qualitative echocardiography assessments.
Bias-Free Scalable Gaussian Processes via Randomized Truncations
Feb 12, 2021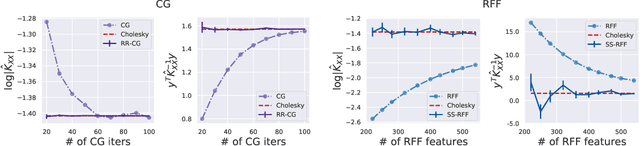
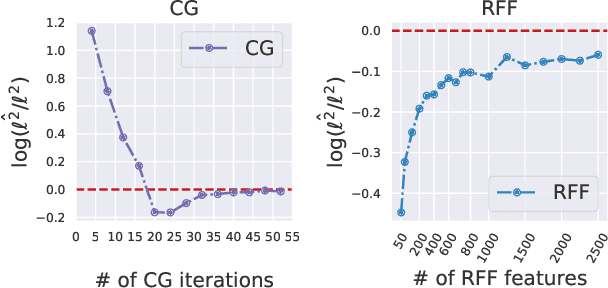
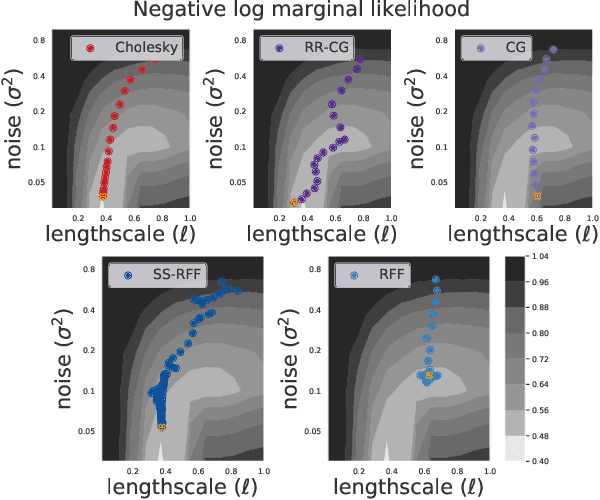
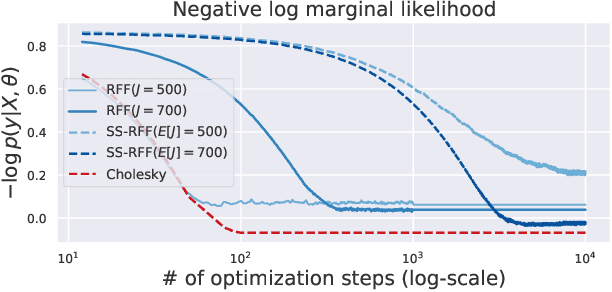
Abstract:Scalable Gaussian Process methods are computationally attractive, yet introduce modeling biases that require rigorous study. This paper analyzes two common techniques: early truncated conjugate gradients (CG) and random Fourier features (RFF). We find that both methods introduce a systematic bias on the learned hyperparameters: CG tends to underfit while RFF tends to overfit. We address these issues using randomized truncation estimators that eliminate bias in exchange for increased variance. In the case of RFF, we show that the bias-to-variance conversion is indeed a trade-off: the additional variance proves detrimental to optimization. However, in the case of CG, our unbiased learning procedure meaningfully outperforms its biased counterpart with minimal additional computation.
Uses and Abuses of the Cross-Entropy Loss: Case Studies in Modern Deep Learning
Nov 10, 2020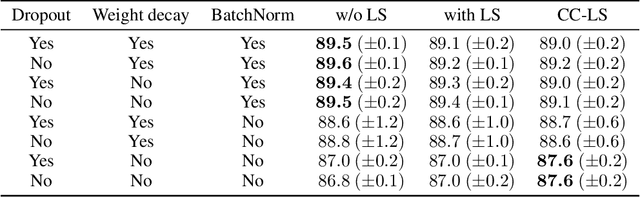
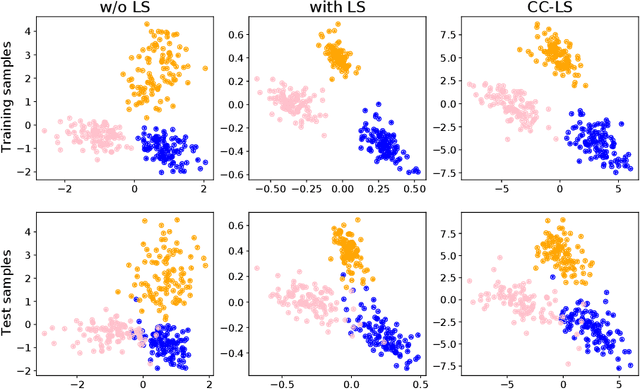

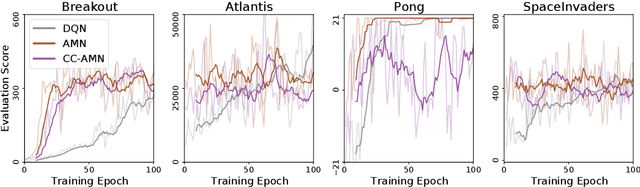
Abstract:Modern deep learning is primarily an experimental science, in which empirical advances occasionally come at the expense of probabilistic rigor. Here we focus on one such example; namely the use of the categorical cross-entropy loss to model data that is not strictly categorical, but rather takes values on the simplex. This practice is standard in neural network architectures with label smoothing and actor-mimic reinforcement learning, amongst others. Drawing on the recently discovered continuous-categorical distribution, we propose probabilistically-inspired alternatives to these models, providing an approach that is more principled and theoretically appealing. Through careful experimentation, including an ablation study, we identify the potential for outperformance in these models, thereby highlighting the importance of a proper probabilistic treatment, as well as illustrating some of the failure modes thereof.
General linear-time inference for Gaussian Processes on one dimension
Mar 11, 2020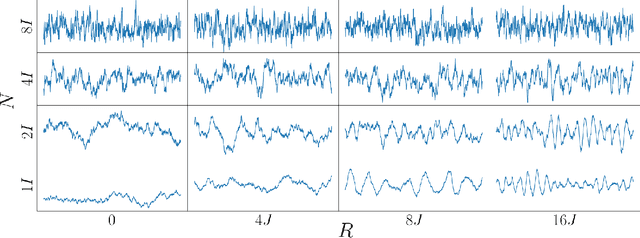
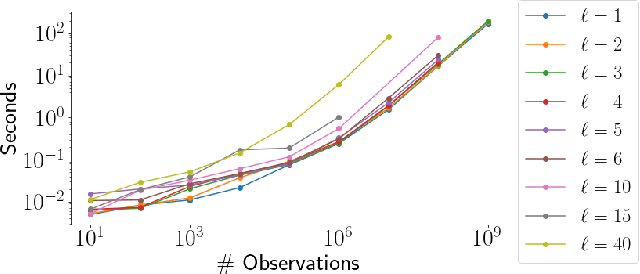
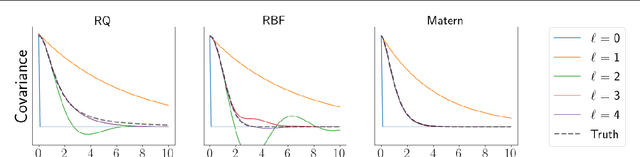
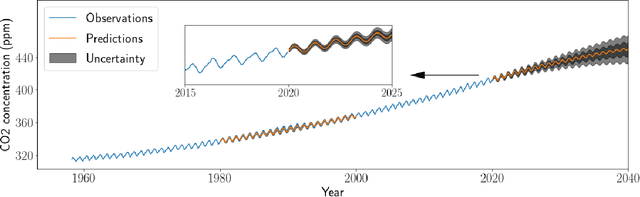
Abstract:Gaussian Processes (GPs) provide a powerful probabilistic framework for interpolation, forecasting, and smoothing, but have been hampered by computational scaling issues. Here we prove that for data sampled on one dimension (e.g., a time series sampled at arbitrarily-spaced intervals), approximate GP inference at any desired level of accuracy requires computational effort that scales linearly with the number of observations; this new theorem enables inference on much larger datasets than was previously feasible. To achieve this improved scaling we propose a new family of stationary covariance kernels: the Latent Exponentially Generated (LEG) family, which admits a convenient stable state-space representation that allows linear-time inference. We prove that any continuous integrable stationary kernel can be approximated arbitrarily well by some member of the LEG family. The proof draws connections to Spectral Mixture Kernels, providing new insight about the flexibility of this popular family of kernels. We propose parallelized algorithms for performing inference and learning in the LEG model, test the algorithm on real and synthetic data, and demonstrate scaling to datasets with billions of samples.
The continuous categorical: a novel simplex-valued exponential family
Feb 20, 2020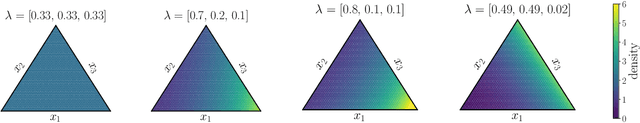
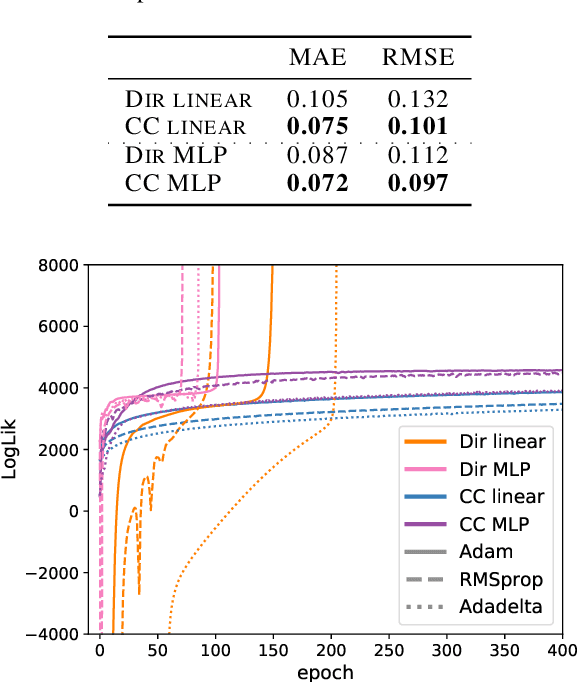
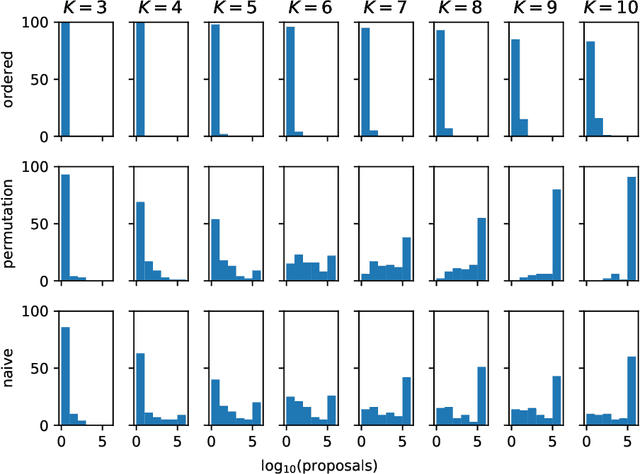
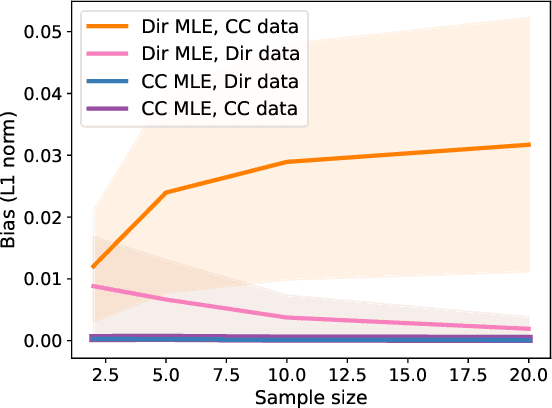
Abstract:Simplex-valued data appear throughout statistics and machine learning, for example in the context of transfer learning and compression of deep networks. Existing models for this class of data rely on the Dirichlet distribution or other related loss functions; here we show these standard choices suffer systematically from a number of limitations, including bias and numerical issues that frustrate the use of flexible network models upstream of these distributions. We resolve these limitations by introducing a novel exponential family of distributions for modeling simplex-valued data - the continuous categorical, which arises as a nontrivial multivariate generalization of the recently discovered continuous Bernoulli. Unlike the Dirichlet and other typical choices, the continuous categorical results in a well-behaved probabilistic loss function that produces unbiased estimators, while preserving the mathematical simplicity of the Dirichlet. As well as exploring its theoretical properties, we introduce sampling methods for this distribution that are amenable to the reparameterization trick, and evaluate their performance. Lastly, we demonstrate that the continuous categorical outperforms standard choices empirically, across a simulation study, an applied example on multi-party elections, and a neural network compression task.
Invertible Gaussian Reparameterization: Revisiting the Gumbel-Softmax
Feb 07, 2020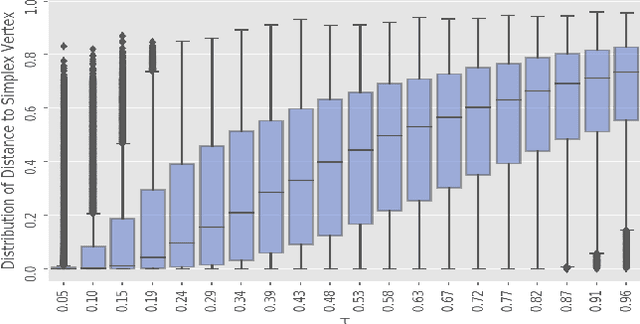
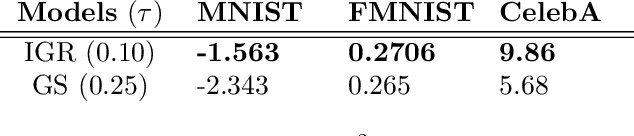
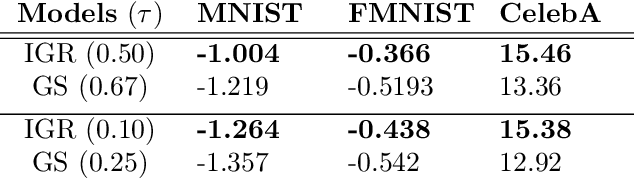
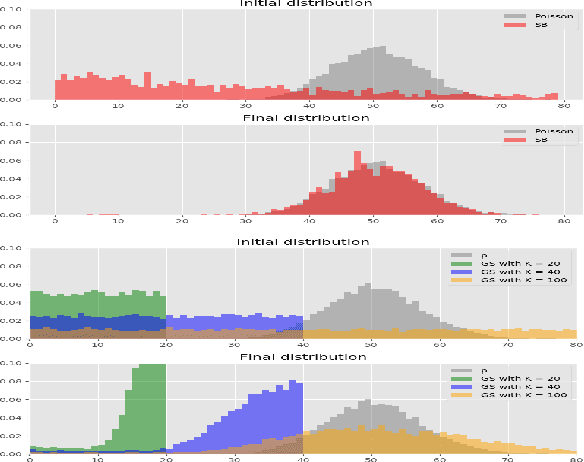

Abstract:The Gumbel-Softmax is a continuous distribution over the simplex that is often used as a relaxation of discrete distributions. Because it can be readily interpreted and easily reparameterized, it enjoys widespread use. Unfortunately, we show that the cost of this aesthetic interpretability is material: the temperature hyperparameter must be set too high, KL estimates are noisy, and as a result, performance suffers. We circumvent the previous issues by proposing a much simpler and more flexible reparameterizable family of distributions that transforms Gaussian noise into a one-hot approximation through an invertible function. This invertible function is composed of a modified softmax and can incorporate diverse transformations that serve different specific purposes. For example, the stick-breaking procedure allows us to extend the reparameterization trick to distributions with countably infinite support, or normalizing flows let us increase the flexibility of the distribution. Our construction improves numerical stability and outperforms the Gumbel-Softmax in a variety of experiments while generating samples that are closer to their discrete counterparts and achieving lower-variance gradients.
Paraphrase Generation with Latent Bag of Words
Jan 07, 2020
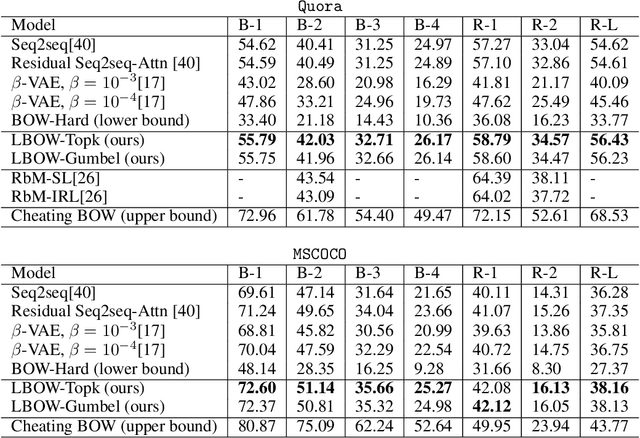
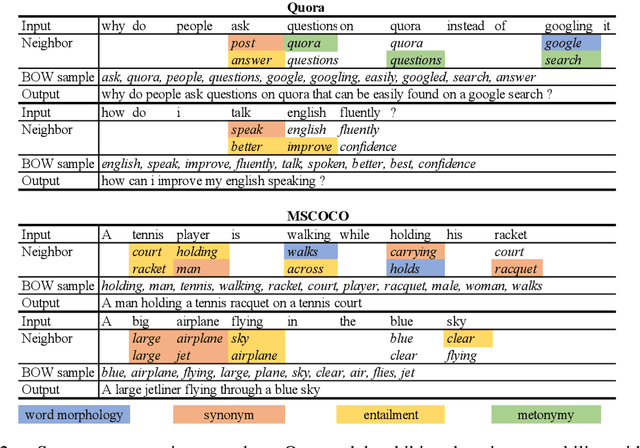
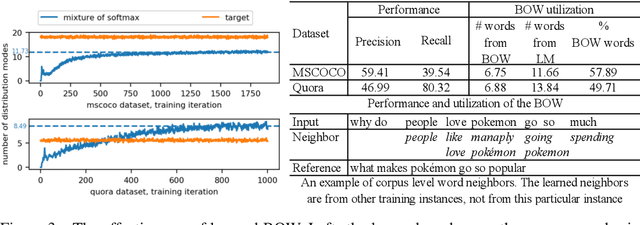
Abstract:Paraphrase generation is a longstanding important problem in natural language processing. In addition, recent progress in deep generative models has shown promising results on discrete latent variables for text generation. Inspired by variational autoencoders with discrete latent structures, in this work, we propose a latent bag of words (BOW) model for paraphrase generation. We ground the semantics of a discrete latent variable by the BOW from the target sentences. We use this latent variable to build a fully differentiable content planning and surface realization model. Specifically, we use source words to predict their neighbors and model the target BOW with a mixture of softmax. We use Gumbel top-k reparameterization to perform differentiable subset sampling from the predicted BOW distribution. We retrieve the sampled word embeddings and use them to augment the decoder and guide its generation search space. Our latent BOW model not only enhances the decoder, but also exhibits clear interpretability. We show the model interpretability with regard to \emph{(i)} unsupervised learning of word neighbors \emph{(ii)} the step-by-step generation procedure. Extensive experiments demonstrate the transparent and effective generation process of this model.\footnote{Our code can be found at \url{https://github.com/FranxYao/dgm_latent_bow}}
The continuous Bernoulli: fixing a pervasive error in variational autoencoders
Jul 23, 2019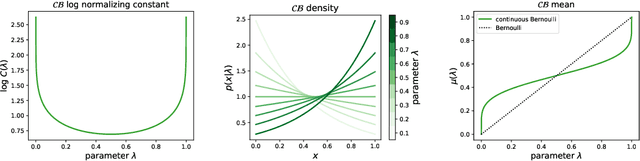
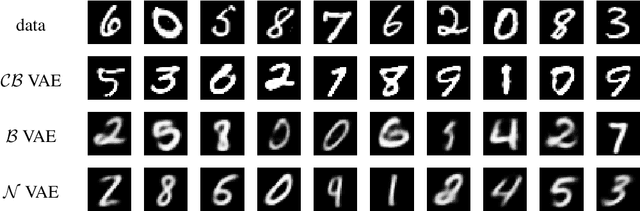
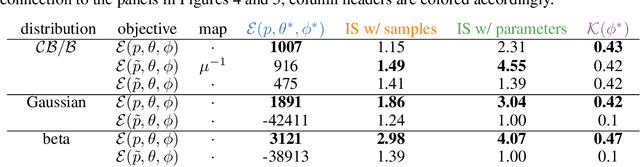

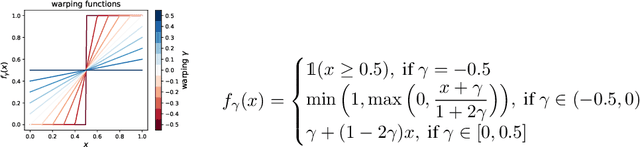
Abstract:Variational autoencoders (VAE) have quickly become a central tool in machine learning, applicable to a broad range of data types and latent variable models. By far the most common first step, taken by seminal papers and by core software libraries alike, is to model MNIST data using a deep network parameterizing a Bernoulli likelihood. This practice contains what appears to be and what is often set aside as a minor inconvenience: the pixel data is [0,1] valued, not {0,1} as supported by the Bernoulli likelihood. Here we show that, far from being a triviality or nuisance that is convenient to ignore, this error has profound importance to VAE, both qualitative and quantitative. We introduce and fully characterize a new [0,1]-supported, single parameter distribution: the continuous Bernoulli, which patches this pervasive bug in VAE. This distribution is not nitpicking; it produces meaningful performance improvements across a range of metrics and datasets, including sharper image samples, and suggests a broader class of performant VAE.
Approximating exponential family models (not single distributions) with a two-network architecture
Mar 18, 2019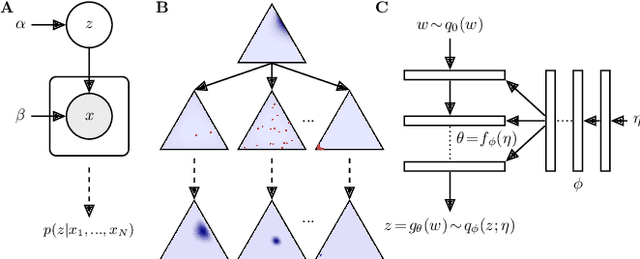
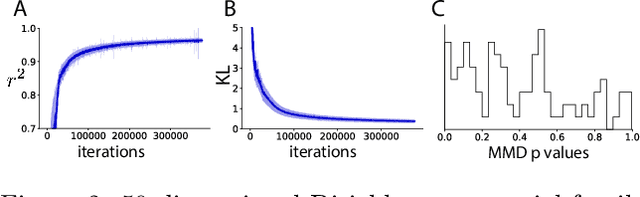
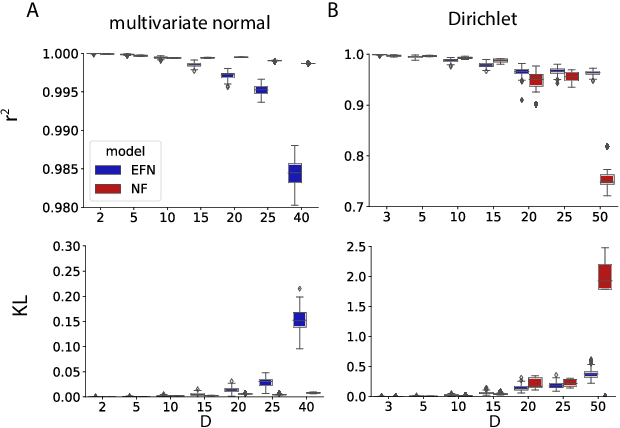
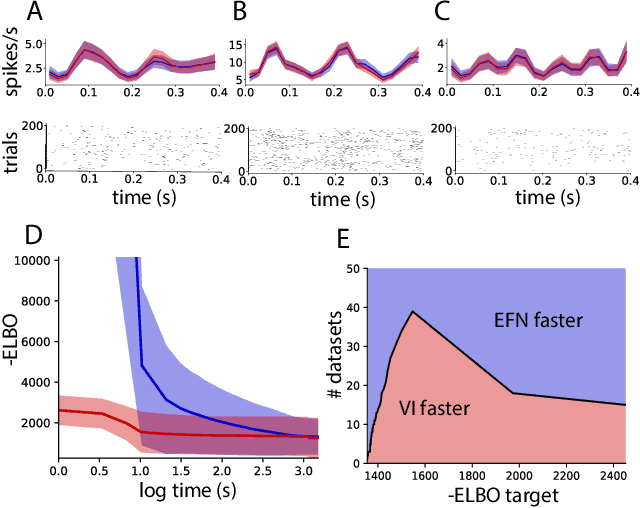
Abstract:Recently much attention has been paid to deep generative models, since they have been used to great success for variational inference, generation of complex data types, and more. In most all of these settings, the goal has been to find a particular member of that model family: optimized parameters index a distribution that is close (via a divergence or classification metric) to a target distribution. Much less attention, however, has been paid to the problem of learning a model itself. Here we introduce a two-network architecture and optimization procedure for learning intractable exponential family models (not a single distribution from those models). These exponential families are learned accurately, allowing operations like posterior inference to be executed directly and generically with an input choice of natural parameters, rather than performing inference via optimization for each particular distribution within that model.
Deep Random Splines for Point Process Intensity Estimation
Mar 06, 2019

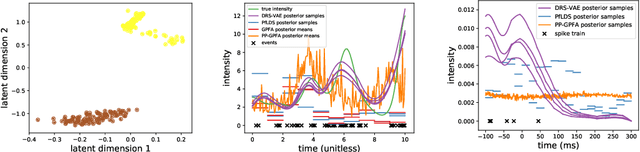
Abstract:Gaussian processes are the leading class of distributions on random functions, but they suffer from well known issues including difficulty scaling and inflexibility with respect to certain shape constraints (such as nonnegativity). Here we propose Deep Random Splines, a flexible class of random functions obtained by transforming Gaussian noise through a deep neural network whose output are the parameters of a spline. Unlike Gaussian processes, Deep Random Splines allow us to readily enforce shape constraints while inheriting the richness and tractability of deep generative models. We also present an observational model for point process data which uses Deep Random Splines to model the intensity function of each point process and apply it to neuroscience data to obtain a low-dimensional representation of spiking activity. Inference is performed via a variational autoencoder that uses a novel recurrent encoder architecture that can handle multiple point processes as input.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge