Jennifer Steeden
Training deep learning based dynamic MR image reconstruction using synthetic fractals
Mar 31, 2026Abstract:Purpose: To investigate whether synthetically generated fractal data can be used to train deep learning (DL) models for dynamic MRI reconstruction, thereby avoiding the privacy, licensing, and availability limitations associated with cardiac MR training datasets. Methods: A training dataset was generated using quaternion Julia fractals to produce 2D+time images. Multi-coil MRI acquisition was simulated to generate paired fully sampled and radially undersampled k-space data. A 3D UNet deep artefact suppression model was trained using these fractal data (F-DL) and compared with an identical model trained on cardiac MRI data (CMR-DL). Both models were evaluated on prospectively acquired radial real-time cardiac MRI from 10 patients. Reconstructions were compared against compressed sensing(CS) and low-rank deep image prior (LR-DIP). All reconstrctuions were ranked for image quality, while ventricular volumes and ejection fraction were compared with reference breath-hold cine MRI. Results: There was no significant difference in qualitative ranking between F-DL and CMR-DL (p=0.9), while both outperformed CS and LR-DIP (p<0.001). Ventricular volumes and function derived from F-DL were similar to CMR-DL, showing no significant bias and accptable limits of agreement compared to reference cine imaging. However, LR-DIP had a signifcant bias (p=0.016) and wider lmits of agreement. Conclusion: DL models trained using synthetic fractal data can reconstruct real-time cardiac MRI with image quality and clinical measurements comparable to models trained on true cardiac MRI data. Fractal training data provide an open, scalable alternative to clinical datasets and may enable development of more generalisable DL reconstruction models for dynamic MRI.
Investigating the use of publicly available natural videos to learn Dynamic MR image reconstruction
Nov 23, 2023Abstract:Purpose: To develop and assess a deep learning (DL) pipeline to learn dynamic MR image reconstruction from publicly available natural videos (Inter4K). Materials and Methods: Learning was performed for a range of DL architectures (VarNet, 3D UNet, FastDVDNet) and corresponding sampling patterns (Cartesian, radial, spiral) either from true multi-coil cardiac MR data (N=692) or from pseudo-MR data simulated from Inter4K natural videos (N=692). Real-time undersampled dynamic MR images were reconstructed using DL networks trained with cardiac data and natural videos, and compressed sensing (CS). Differences were assessed in simulations (N=104 datasets) in terms of MSE, PSNR, and SSIM and prospectively for cardiac (short axis, four chambers, N=20) and speech (N=10) data in terms of subjective image quality ranking, SNR and Edge sharpness. Friedman Chi Square tests with post-hoc Nemenyi analysis were performed to assess statistical significance. Results: For all simulation metrics, DL networks trained with cardiac data outperformed DL networks trained with natural videos, which outperformed CS (p<0.05). However, in prospective experiments DL reconstructions using both training datasets were ranked similarly (and higher than CS) and presented no statistical differences in SNR and Edge Sharpness for most conditions. Additionally, high SSIM was measured between the DL methods with cardiac data and natural videos (SSIM>0.85). Conclusion: The developed pipeline enabled learning dynamic MR reconstruction from natural videos preserving DL reconstruction advantages such as high quality fast and ultra-fast reconstructions while overcoming some limitations (data scarcity or sharing). The natural video dataset, code and pre-trained networks are made readily available on github. Key Words: real-time; dynamic MRI; deep learning; image reconstruction; machine learning;
GLADE: Gradient Loss Augmented Degradation Enhancement for Unpaired Super-Resolution of Anisotropic MRI
Mar 21, 2023Abstract:We present a novel approach to synthesise high-resolution isotropic 3D abdominal MR images, from anisotropic 3D images in an unpaired fashion. Using a modified CycleGAN architecture with a gradient mapping loss, we leverage disjoint patches from the high-resolution (in-plane) data of an anisotropic volume to enforce the network generator to increase the resolution of the low-resolution (through-plane) slices. This will enable accelerated whole-abdomen scanning with high-resolution isotropic images within short breath-hold times.
FReSCO: Flow Reconstruction and Segmentation for low latency Cardiac Output monitoring using deep artifact suppression and segmentation
Mar 25, 2022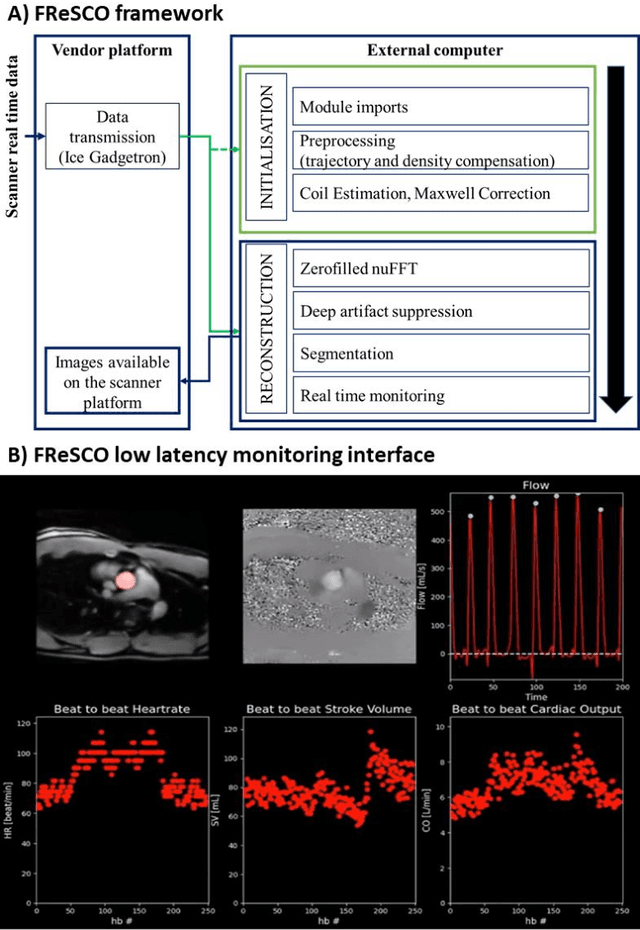
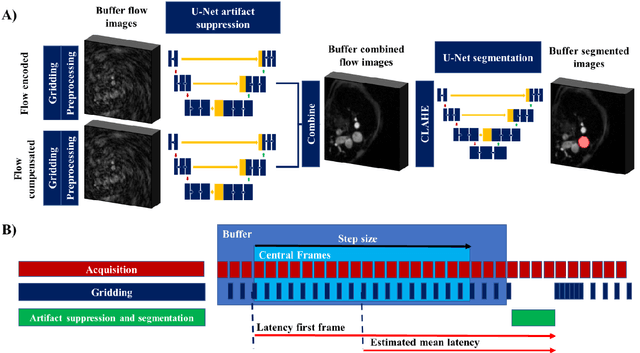
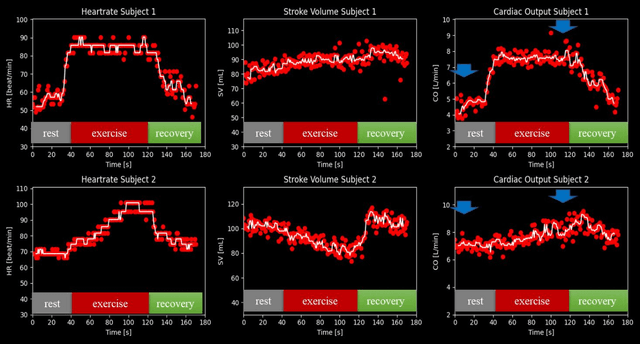
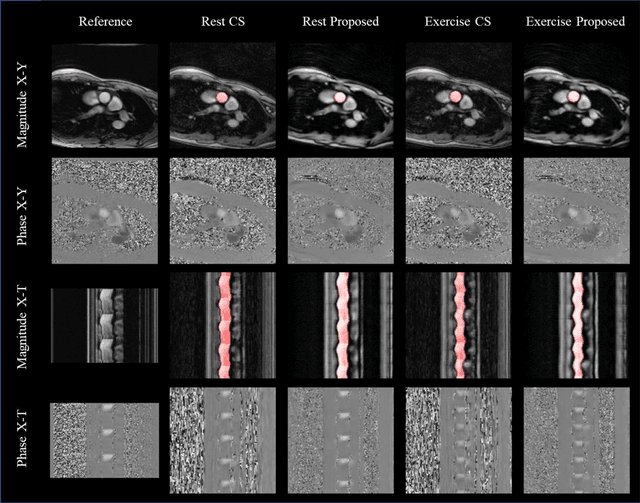
Abstract:Purpose: Real-time monitoring of cardiac output (CO) requires low latency reconstruction and segmentation of real-time phase contrast MR (PCMR), which has previously been difficult to perform. Here we propose a deep learning framework for 'Flow Reconstruction and Segmentation for low latency Cardiac Output monitoring' (FReSCO). Methods: Deep artifact suppression and segmentation U-Nets were independently trained. Breath hold spiral PCMR data (n=516) was synthetically undersampled using a variable density spiral sampling pattern and gridded to create aliased data for training of the artifact suppression U-net. A subset of the data (n=96) was segmented and used to train the segmentation U-net. Real-time spiral PCMR was prospectively acquired and then reconstructed and segmented using the trained models (FReSCO) at low latency at the scanner in 10 healthy subjects during rest, exercise and recovery periods. CO obtained via FReSCO was compared to a reference rest CO and rest and exercise Compressed Sensing (CS) CO. Results: FReSCO was demonstrated prospectively at the scanner. Beat-to-beat heartrate, stroke volume and CO could be visualized with a mean latency of 622ms. No significant differences were noted when compared to reference at rest (Bias = -0.21+-0.50 L/min, p=0.246) or CS at peak exercise (Bias=0.12+-0.48 L/min, p=0.458). Conclusion: FReSCO was successfully demonstrated for real-time monitoring of CO during exercise and could provide a convenient tool for assessment of the hemodynamic response to a range of stressors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge