James Newling
A Sub-Quadratic Exact Medoid Algorithm
Apr 12, 2017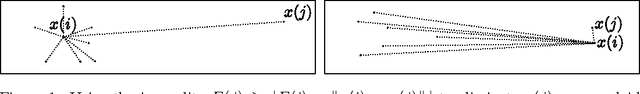
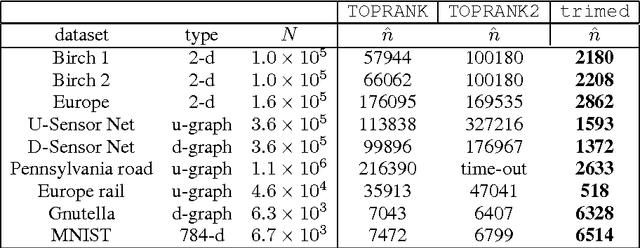
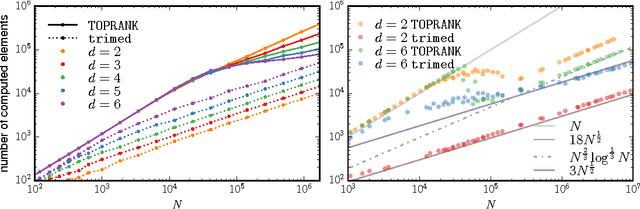
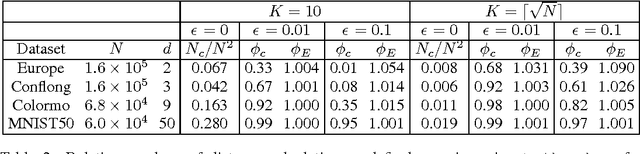
Abstract:We present a new algorithm, trimed, for obtaining the medoid of a set, that is the element of the set which minimises the mean distance to all other elements. The algorithm is shown to have, under certain assumptions, expected run time O(N^(3/2)) in R^d where N is the set size, making it the first sub-quadratic exact medoid algorithm for d>1. Experiments show that it performs very well on spatial network data, frequently requiring two orders of magnitude fewer distance calculations than state-of-the-art approximate algorithms. As an application, we show how trimed can be used as a component in an accelerated K-medoids algorithm, and then how it can be relaxed to obtain further computational gains with only a minor loss in cluster quality.
Nested Mini-Batch K-Means
Sep 12, 2016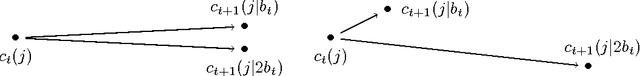
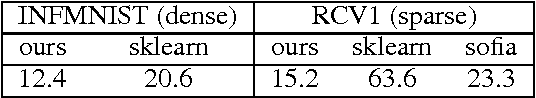
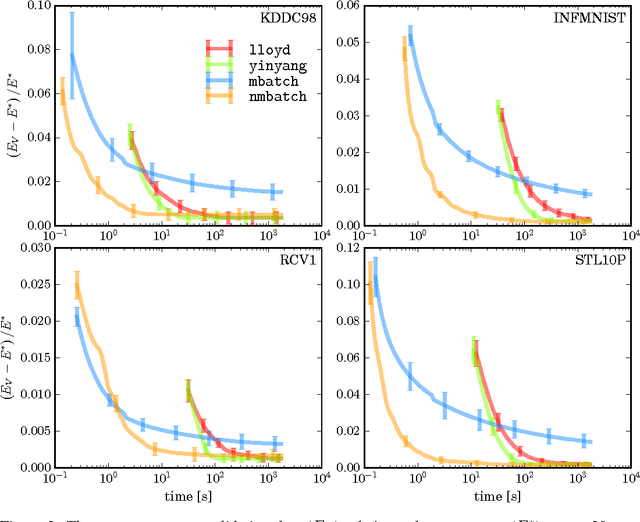
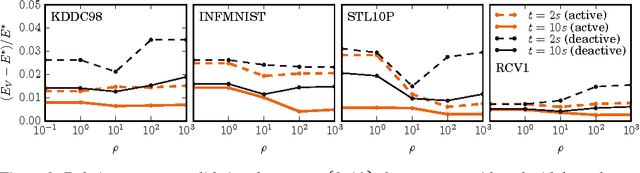
Abstract:A new algorithm is proposed which accelerates the mini-batch k-means algorithm of Sculley (2010) by using the distance bounding approach of Elkan (2003). We argue that, when incorporating distance bounds into a mini-batch algorithm, already used data should preferentially be reused. To this end we propose using nested mini-batches, whereby data in a mini-batch at iteration t is automatically reused at iteration t+1. Using nested mini-batches presents two difficulties. The first is that unbalanced use of data can bias estimates, which we resolve by ensuring that each data sample contributes exactly once to centroids. The second is in choosing mini-batch sizes, which we address by balancing premature fine-tuning of centroids with redundancy induced slow-down. Experiments show that the resulting nmbatch algorithm is very effective, often arriving within 1% of the empirical minimum 100 times earlier than the standard mini-batch algorithm.
* 8 pages + Supplementary Material. Version 2 : new experiments added. Version 3 : Add acknowledgments, upper case in title. Version 4 : Correct spelling of Acknowledgements, change title. Version 5: camera ready NIPS
Fast K-Means with Accurate Bounds
Sep 11, 2016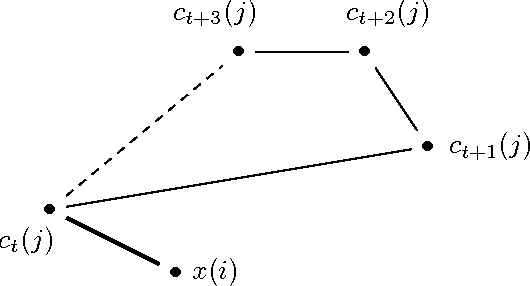
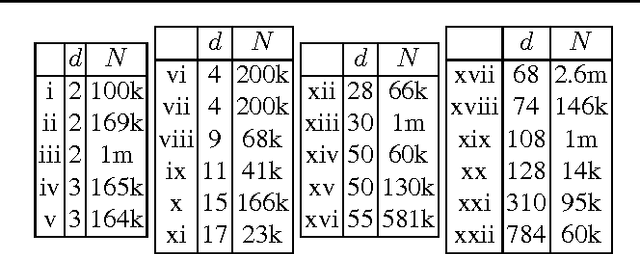
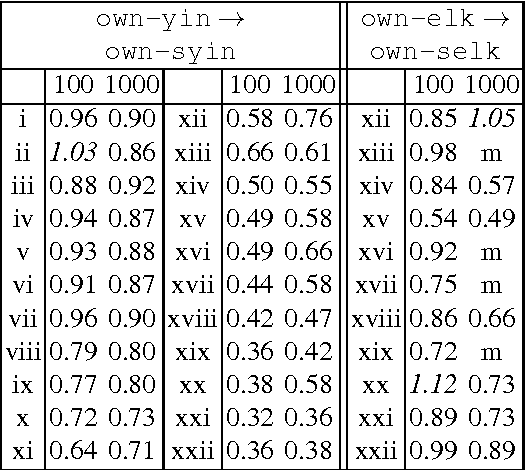
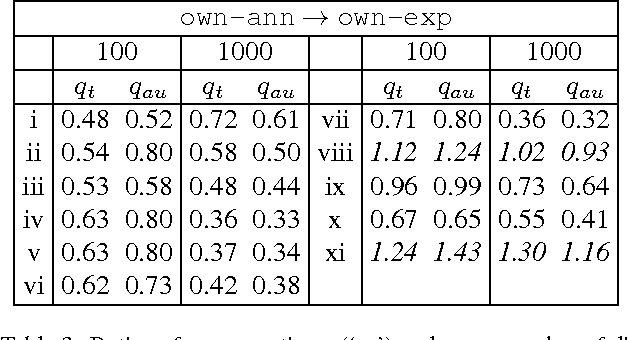
Abstract:We propose a novel accelerated exact k-means algorithm, which performs better than the current state-of-the-art low-dimensional algorithm in 18 of 22 experiments, running up to 3 times faster. We also propose a general improvement of existing state-of-the-art accelerated exact k-means algorithms through better estimates of the distance bounds used to reduce the number of distance calculations, and get a speedup in 36 of 44 experiments, up to 1.8 times faster. We have conducted experiments with our own implementations of existing methods to ensure homogeneous evaluation of performance, and we show that our implementations perform as well or better than existing available implementations. Finally, we propose simplified variants of standard approaches and show that they are faster than their fully-fledged counterparts in 59 of 62 experiments.
* 8 pages + supplementary material v2: mlpack installed with optimisation (previously installed in DEBUG) v3: Annulus -> Annular v4: Author affiliation update v5: Synced with version at ICML, now including Suppl. Mat
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge