J. Cohen-Adad
NeoRS: a neonatal resting state fMRI data preprocessing pipeline
Apr 08, 2022
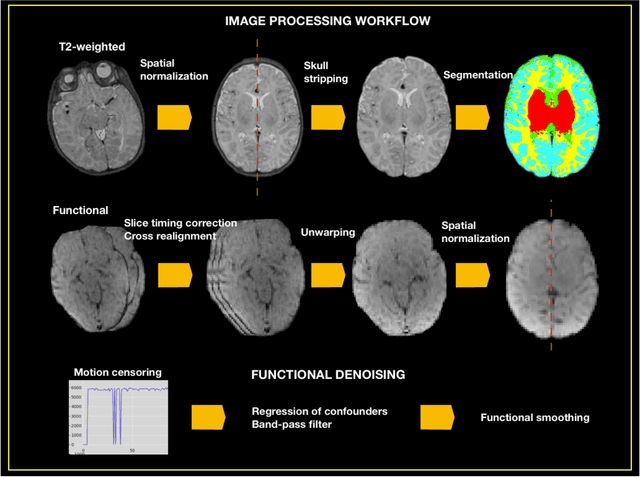
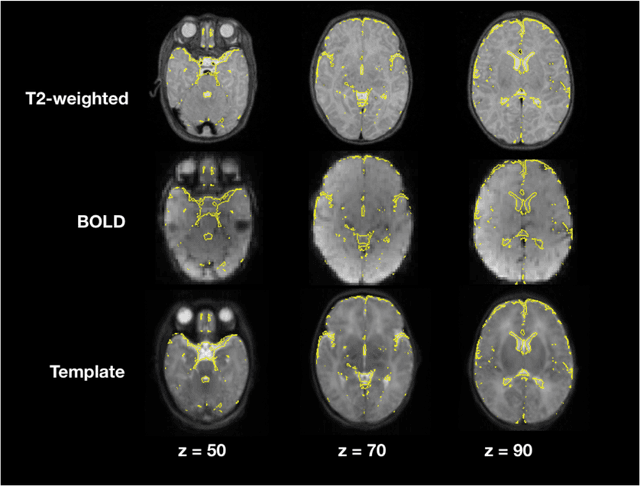
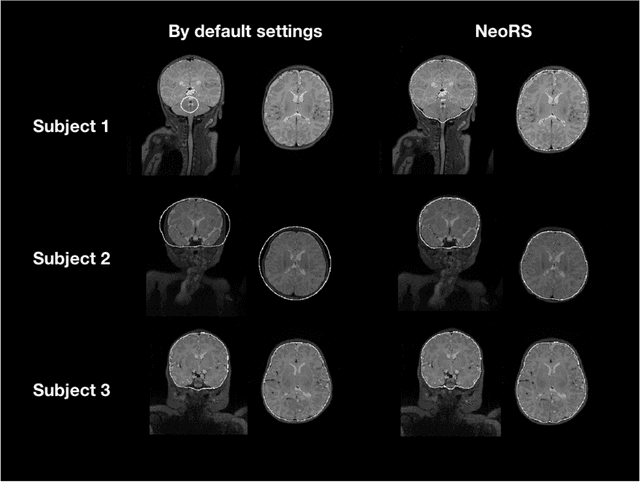
Abstract:Resting state fMRI (rsfMRI) has been shown to be a promising tool to study intrinsic functional connectivity and assess its integrity in cerebral development. In neonates, where fMRI is limited to few paradigms, rsfMRI was shown to be a relevant tool to explore regional interactions of brain networks. However, to identify the resting state networks, data needs to be carefully processed. Because of the non-collaborative nature of the neonates, the differences in brain size and the reversed contrast compared to adults, neonates can't be processed with the existing adult pipelines. Therefore, we developed NeoRS. The main processing steps include atlas registration, skull tripping, segmentation, slice timing and head motion correction and confounds regression. To address the specificity of neonatal brain imaging, particular attention was given to registration including neonatal atlas type and parameters, such as brain size variations, and contrast differences compared to adults. Furthermore, head motion was scrutinized and optimized, as it is a major issue when processing neonatal data. The pipeline includes visual quality control assessment checkpoints. To assess its effectiveness, we used the data from the Baby Connectome Project including 10 neonates. NeoRS was designed to work on both multi-band and single-band acquisitions and is applicable on smaller datasets. It also includes popular functional connectivity analysis features such as seed based correlations. Language, default mode, dorsal attention, visual, ventral attention, motor and fronto parietal networks were evaluated. The different analyzed networks were in agreement with previously published studies in the neonate. NeoRS is coded in Matlab, it is open-source and available on https://github.com/venguix/NeoRS. NeoRS allows robust image processing of the neonatal rsfMRI data that can be readily customized to different datasets.
Dynamic Realtime z-Shimming: A Feasibility Study
Jul 21, 2021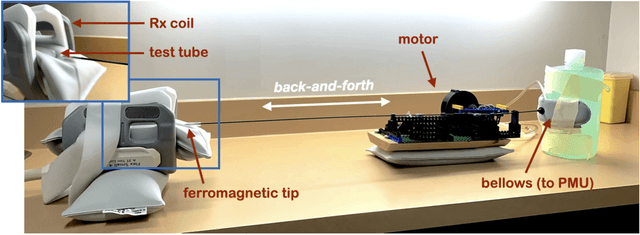
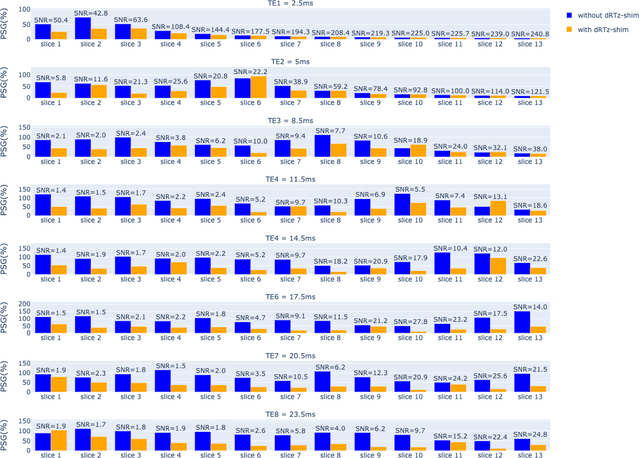
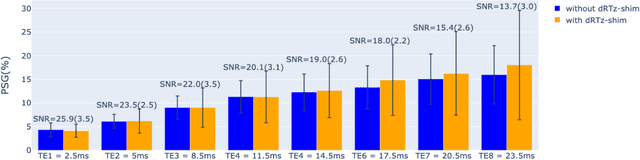
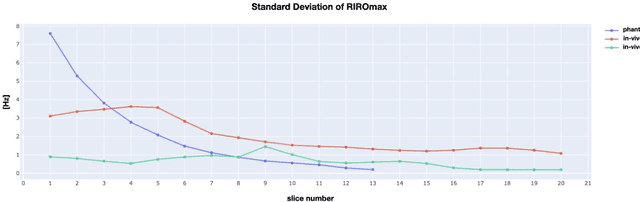
Abstract:Respiration causes time-varying frequency offsets that can result in ghosting artifacts. We propose a solution, which we term dynamic realtime z-shimming, wherein linear gradients are adjusted dynamically (slice-wise) and in real-time, to reflect magnetic field inhomogeneities that arise during image acquisition. In dynamic z-shimming, a method that is commonly used to reduce static frequency offsets in MR images of the spinal cord and brain, in-plane (static) frequency offsets are assumed to be homogeneous. Here we investigate whether or not that same assumption can be made for time-varying frequency offsets in the cervical spinal cord region. In order to explore the feasibility of dynamic realtime z-shimming, we acquired images using a pneumatic phantom setup, as well as in-vivo. We then simulated the effects of time-varying frequency offsets on MR images acquired with and without dynamic realtime z-shimming in different scenarios. We found that dynamic realtime z-shimming can reduce ghosting if the time-varying frequency offsets have an in-plane variability (standard deviation) of approximately less than 1 Hz. This scenario was achieved in our phantom setup, where we observed a 50.2% reduction in ghosting within multi-echo gradient echo images acquired with dynamic realtime z-shimming, compared to without. On the other hand, we observed that the in-plane variability of the time-varying frequency offsets is too high within the cervical spinal cord region for dynamic realtime z-shimming to be successful. These results can serve as a guideline and starting point for future dynamic realtime z-shimming experiments in which the in-plane variability of frequency offsets are minimized.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge