Hayit Greenspan
Nodule2vec: a 3D Deep Learning System for Pulmonary Nodule Retrieval Using Semantic Representation
Jul 11, 2020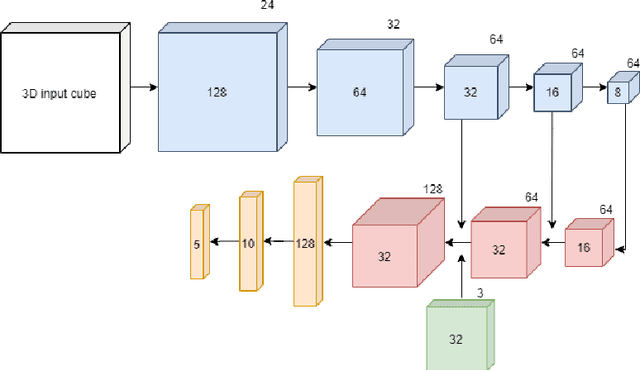
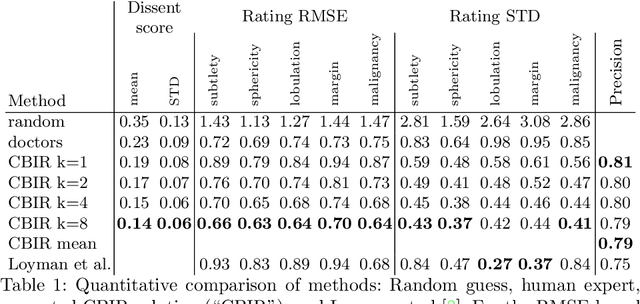
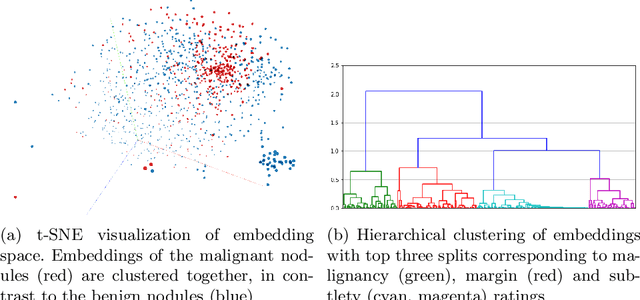
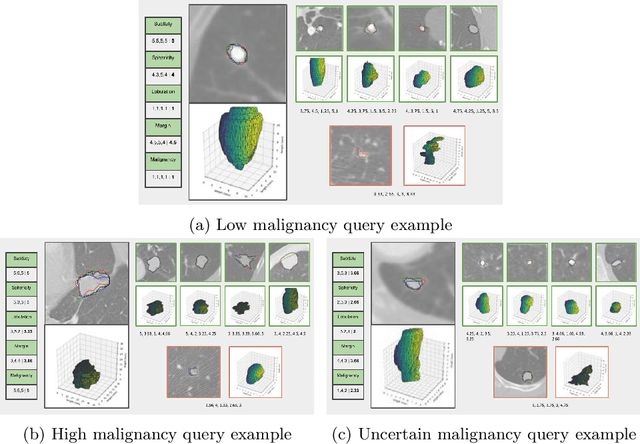
Abstract:Content-based retrieval supports a radiologist decision making process by presenting the doctor the most similar cases from the database containing both historical diagnosis and further disease development history. We present a deep learning system that transforms a 3D image of a pulmonary nodule from a CT scan into a low-dimensional embedding vector. We demonstrate that such a vector representation preserves semantic information about the nodule and offers a viable approach for content-based image retrieval (CBIR). We discuss the theoretical limitations of the available datasets and overcome them by applying transfer learning of the state-of-the-art lung nodule detection model. We evaluate the system using the LIDC-IDRI dataset of thoracic CT scans. We devise a similarity score and show that it can be utilized to measure similarity 1) between annotations of the same nodule by different radiologists and 2) between the query nodule and the top four CBIR results. A comparison between doctors and algorithm scores suggests that the benefit provided by the system to the radiologist end-user is comparable to obtaining a second radiologist's opinion.
Joint Liver Lesion Segmentation and Classification via Transfer Learning
May 24, 2020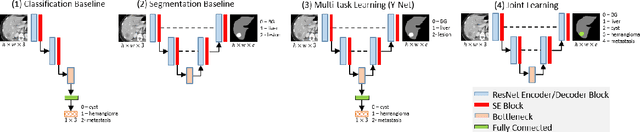
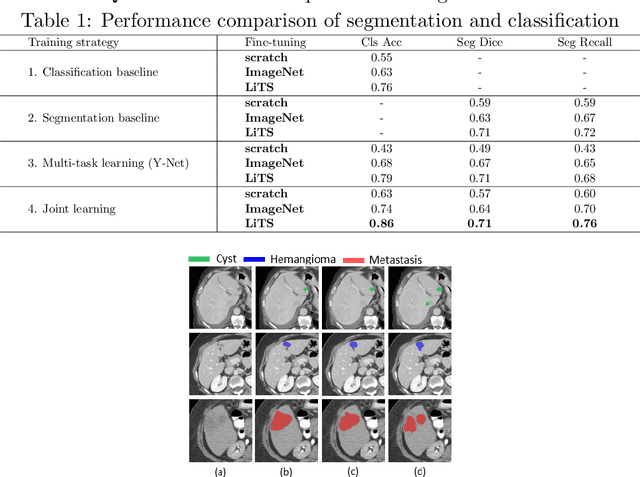
Abstract:Transfer learning and joint learning approaches are extensively used to improve the performance of Convolutional Neural Networks (CNNs). In medical imaging applications in which the target dataset is typically very small, transfer learning improves feature learning while joint learning has shown effectiveness in improving the network's generalization and robustness. In this work, we study the combination of these two approaches for the problem of liver lesion segmentation and classification. For this purpose, 332 abdominal CT slices containing lesion segmentation and classification of three lesion types are evaluated. For feature learning, the dataset of MICCAI 2017 Liver Tumor Segmentation (LiTS) Challenge is used. Joint learning shows improvement in both segmentation and classification results. We show that a simple joint framework outperforms the commonly used multi-task architecture (Y-Net), achieving an improvement of 10% in classification accuracy, compared to a 3% improvement with Y-Net.
Semi-supervised lung nodule retrieval
May 04, 2020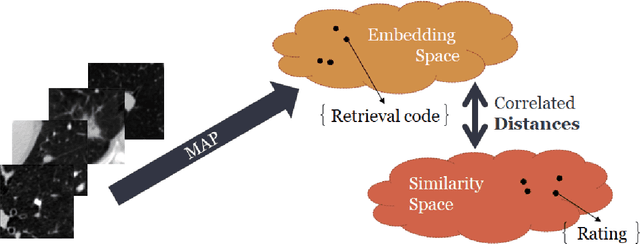
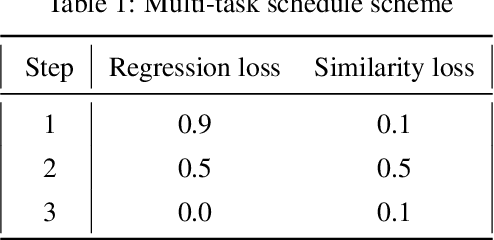
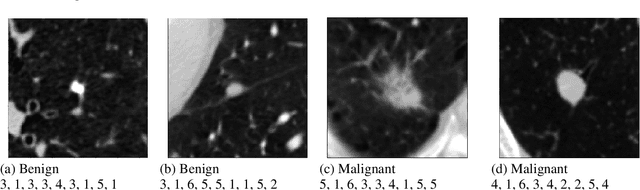
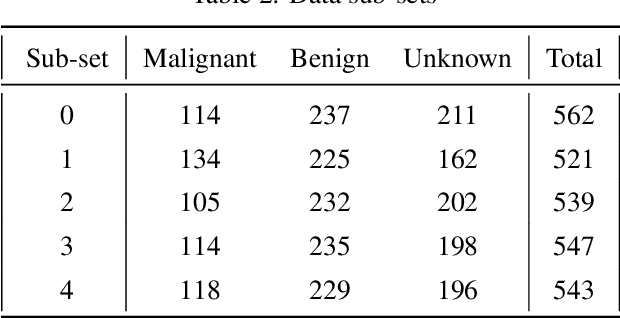
Abstract:Content based image retrieval (CBIR) provides the clinician with visual information that can support, and hopefully improve, his or her decision making process. Given an input query image, a CBIR system provides as its output a set of images, ranked by similarity to the query image. Retrieved images may come with relevant information, such as biopsy-based malignancy labeling, or categorization. Ground truth on similarity between dataset elements (e.g. between nodules) is not readily available, thus greatly challenging machine learning methods. Such annotations are particularly difficult to obtain, due to the subjective nature of the task, with high inter-observer variability requiring multiple expert annotators. Consequently, past approaches have focused on manual feature extraction, while current approaches use auxiliary tasks, such as a binary classification task (e.g. malignancy), for which ground-true is more readily accessible. However, in a previous study, we have shown that binary auxiliary tasks are inferior to the usage of a rough similarity estimate that are derived from data annotations. The current study suggests a semi-supervised approach that involves two steps: 1) Automatic annotation of a given partially labeled dataset; 2) Learning a semantic similarity metric space based on the predicated annotations. The proposed system is demonstrated in lung nodule retrieval using the LIDC dataset, and shows that it is feasible to learn embedding from predicted ratings. The semi-supervised approach has demonstrated a significantly higher discriminative ability than the fully-unsupervised reference.
Coronavirus Detection and Analysis on Chest CT with Deep Learning
Apr 06, 2020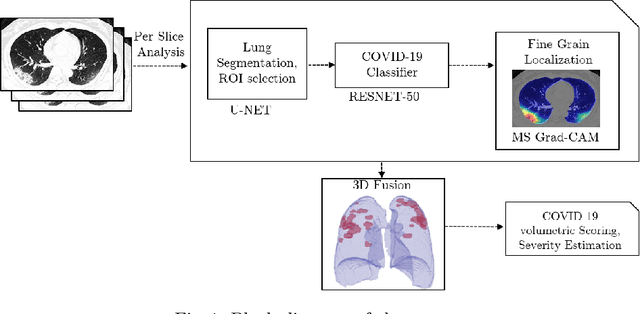
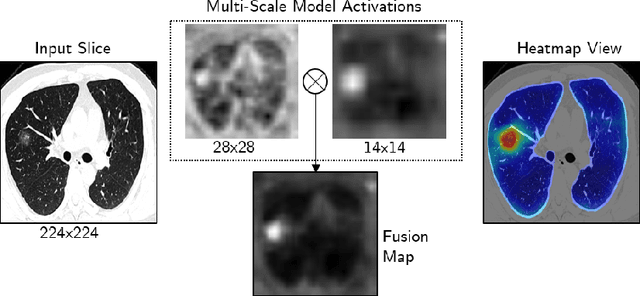


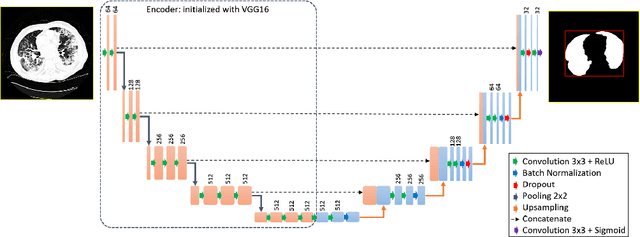
Abstract:The outbreak of the novel coronavirus, officially declared a global pandemic, has a severe impact on our daily lives. As of this writing there are approximately 197,188 confirmed cases of which 80,881 are in "Mainland China" with 7,949 deaths, a mortality rate of 3.4%. In order to support radiologists in this overwhelming challenge, we develop a deep learning based algorithm that can detect, localize and quantify severity of COVID-19 manifestation from chest CT scans. The algorithm is comprised of a pipeline of image processing algorithms which includes lung segmentation, 2D slice classification and fine grain localization. In order to further understand the manifestations of the disease, we perform unsupervised clustering of abnormal slices. We present our results on a dataset comprised of 110 confirmed COVID-19 patients from Zhejiang province, China.
Rapid AI Development Cycle for the Coronavirus (COVID-19) Pandemic: Initial Results for Automated Detection & Patient Monitoring using Deep Learning CT Image Analysis
Mar 24, 2020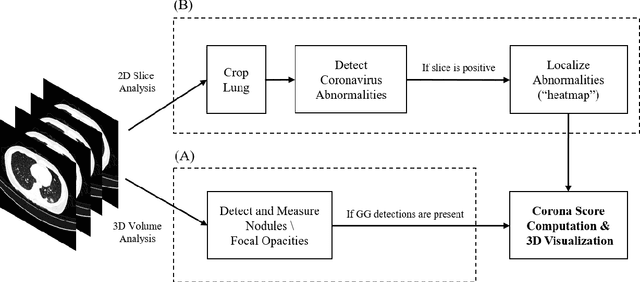
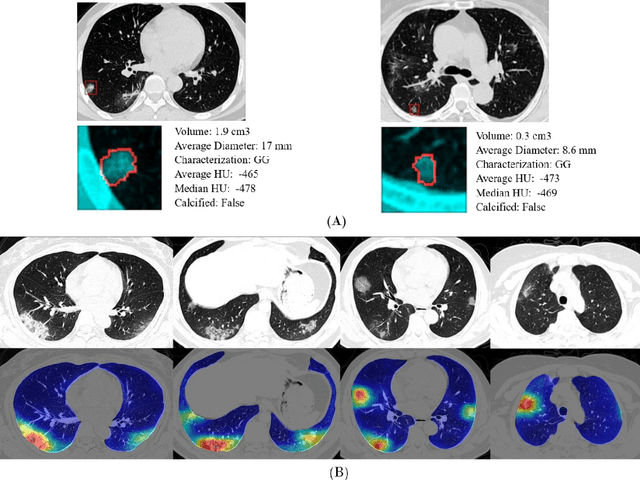
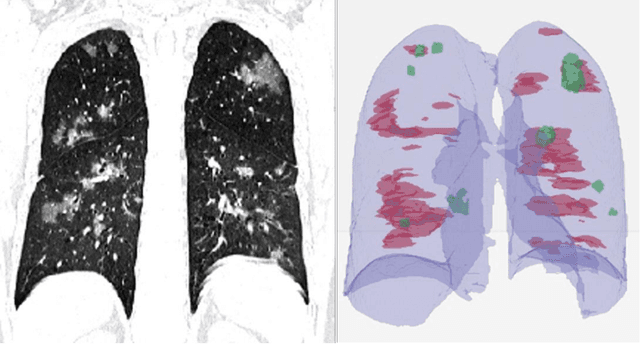
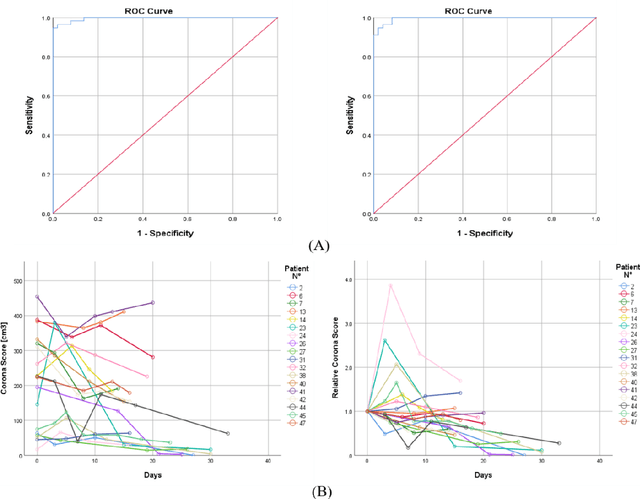
Abstract:Purpose: Develop AI-based automated CT image analysis tools for detection, quantification, and tracking of Coronavirus; demonstrate they can differentiate coronavirus patients from non-patients. Materials and Methods: Multiple international datasets, including from Chinese disease-infected areas were included. We present a system that utilizes robust 2D and 3D deep learning models, modifying and adapting existing AI models and combining them with clinical understanding. We conducted multiple retrospective experiments to analyze the performance of the system in the detection of suspected COVID-19 thoracic CT features and to evaluate evolution of the disease in each patient over time using a 3D volume review, generating a Corona score. The study includes a testing set of 157 international patients (China and U.S). Results: Classification results for Coronavirus vs Non-coronavirus cases per thoracic CT studies were 0.996 AUC (95%CI: 0.989-1.00) ; on datasets of Chinese control and infected patients. Possible working point: 98.2% sensitivity, 92.2% specificity. For time analysis of Coronavirus patients, the system output enables quantitative measurements for smaller opacities (volume, diameter) and visualization of the larger opacities in a slice-based heat map or a 3D volume display. Our suggested Corona score measures the progression of disease over time. Conclusion: This initial study, which is currently being expanded to a larger population, demonstrated that rapidly developed AI-based image analysis can achieve high accuracy in detection of Coronavirus as well as quantification and tracking of disease burden.
Bone Structures Extraction and Enhancement in Chest Radiographs via CNN Trained on Synthetic Data
Mar 20, 2020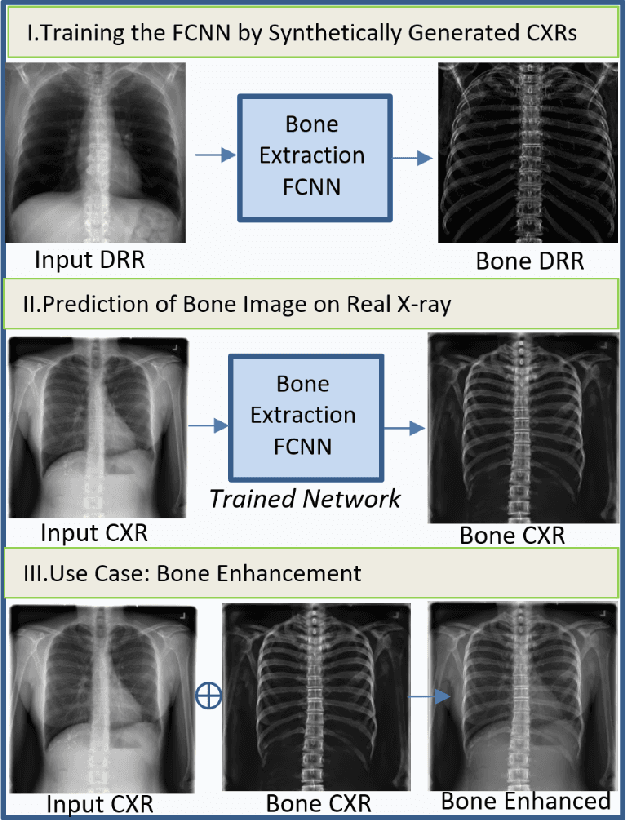
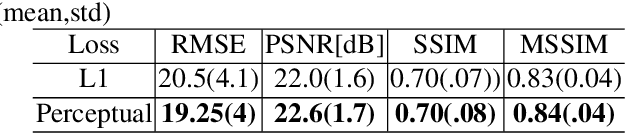

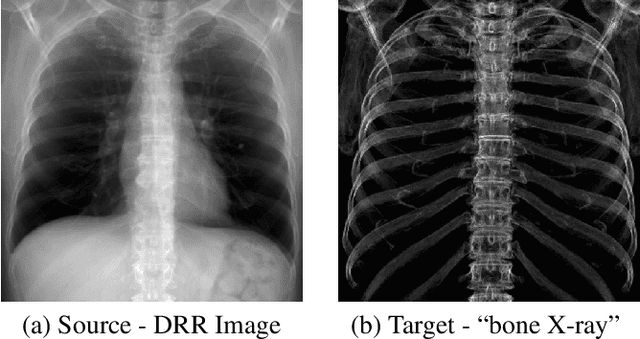
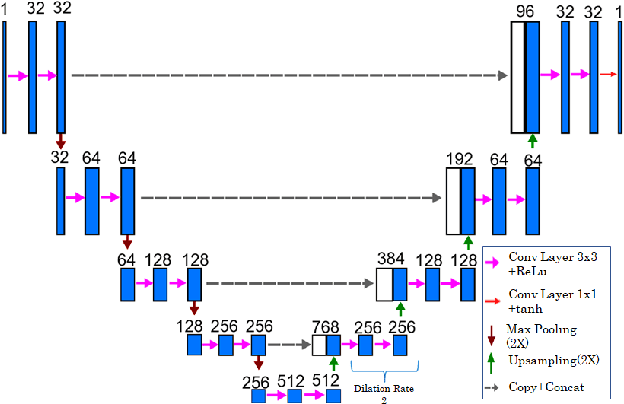
Abstract:In this paper, we present a deep learning-based image processing technique for extraction of bone structures in chest radiographs using a U-Net FCNN. The U-Net was trained to accomplish the task in a fully supervised setting. To create the training image pairs, we employed simulated X-Ray or Digitally Reconstructed Radiographs (DRR), derived from 664 CT scans belonging to the LIDC-IDRI dataset. Using HU based segmentation of bone structures in the CT domain, a synthetic 2D "Bone x-ray" DRR is produced and used for training the network. For the reconstruction loss, we utilize two loss functions- L1 Loss and perceptual loss. Once the bone structures are extracted, the original image can be enhanced by fusing the original input x-ray and the synthesized "Bone X-ray". We show that our enhancement technique is applicable to real x-ray data, and display our results on the NIH Chest X-Ray-14 dataset.
A Soft STAPLE Algorithm Combined with Anatomical Knowledge
Oct 26, 2019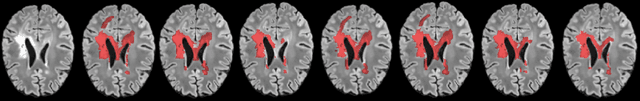
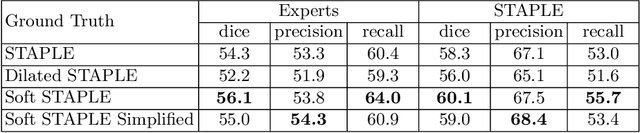
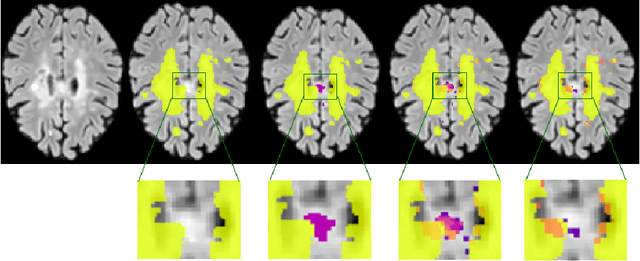
Abstract:Supervised machine learning algorithms, especially in the medical domain, are affected by considerable ambiguity in expert markings. In this study we address the case where the experts' opinion is obtained as a distribution over the possible values. We propose a soft version of the STAPLE algorithm for experts' markings fusion that can handle soft values. The algorithm was applied to obtain consensus from soft Multiple Sclerosis (MS) segmentation masks. Soft MS segmentations are constructed from manual binary delineations by including lesion surrounding voxels in the segmentation mask with a reduced confidence weight. We suggest that these voxels contain additional anatomical information about the lesion structure. The fused masks are utilized as ground truth mask to train a Fully Convolutional Neural Network (FCNN). The proposed method was evaluated on the MICCAI 2016 challenge dataset, and yields improved precision-recall tradeoff and a higher average Dice similarity coefficient.
Automatic Segmentation of Muscle Tissue and Inter-muscular Fat in Thigh and Calf MRI Images
Oct 01, 2019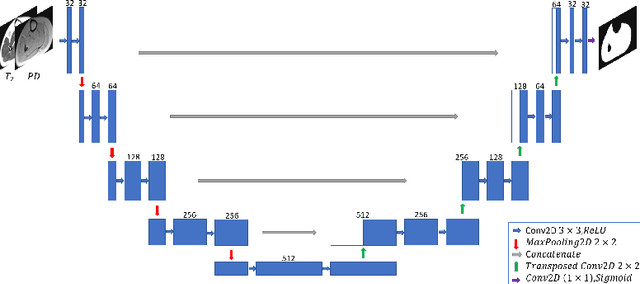
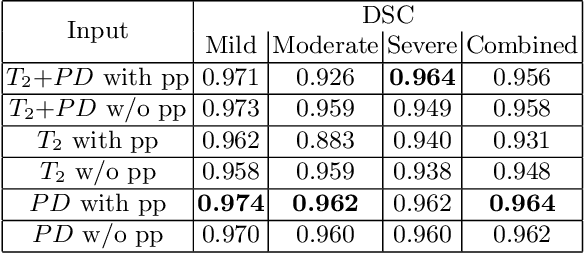

Abstract:Magnetic resonance imaging (MRI) of thigh and calf muscles is one of the most effective techniques for estimating fat infiltration into muscular dystrophies. The infiltration of adipose tissue into the diseased muscle region varies in its severity across, and within, patients. In order to efficiently quantify the infiltration of fat, accurate segmentation of muscle and fat is needed. An estimation of the amount of infiltrated fat is typically done visually by experts. Several algorithmic solutions have been proposed for automatic segmentation. While these methods may work well in mild cases, they struggle in moderate and severe cases due to the high variability in the intensity of infiltration, and the tissue's heterogeneous nature. To address these challenges, we propose a deep-learning approach, producing robust results with high Dice Similarity Coefficient (DSC) of 0.964, 0.917 and 0.933 for muscle-region, healthy muscle and inter-muscular adipose tissue (IMAT) segmentation, respectively.
Endotracheal Tube Detection and Segmentation in Chest Radiographs using Synthetic Data
Aug 20, 2019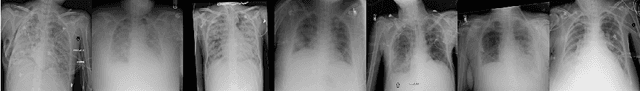
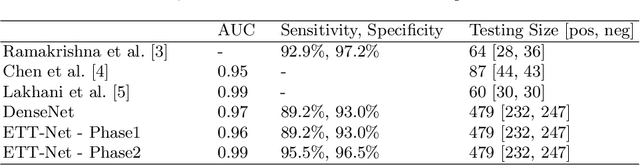
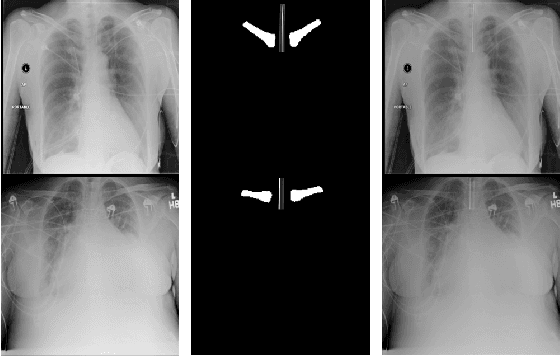
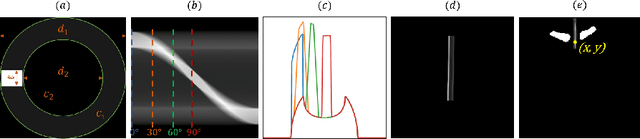
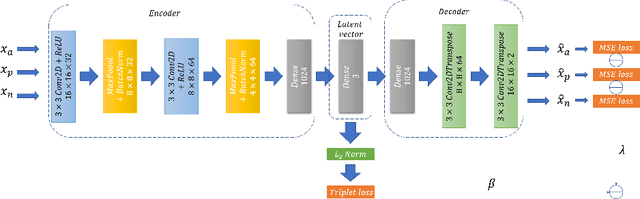
Abstract:Chest radiographs are frequently used to verify the correct intubation of patients in the emergency room. Fast and accurate identification and localization of the endotracheal (ET) tube is critical for the patient. In this study we propose a novel automated deep learning scheme for accurate detection and segmentation of the ET tubes. Development of automatic systems using deep learning networks for classification and segmentation require large annotated data which is not always available. Here we present an approach for synthesizing ET tubes in real X-ray images. We suggest a method for training the network, first with synthetic data and then with real X-ray images in a fine-tuning phase, which allows the network to train on thousands of cases without annotating any data. The proposed method was tested on 477 real chest radiographs from a public dataset and reached AUC of 0.99 in classifying the presence vs. absence of the ET tube, along with outputting high quality ET tube segmentation maps.
Deep Feature Learning from a Hospital-Scale Chest X-ray Dataset with Application to TB Detection on a Small-Scale Dataset
Jun 03, 2019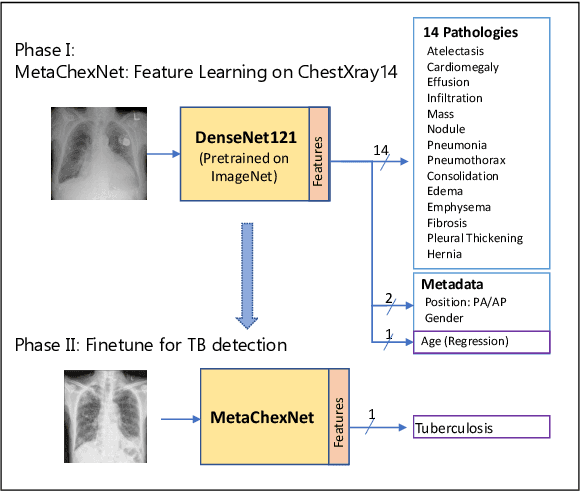
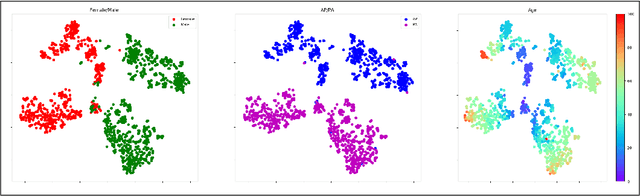
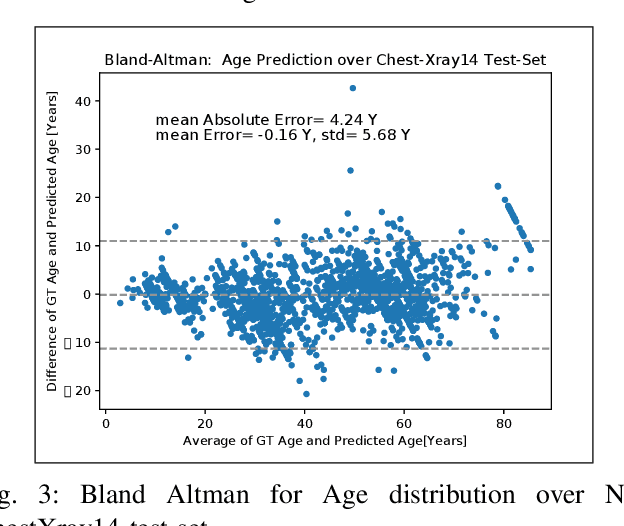
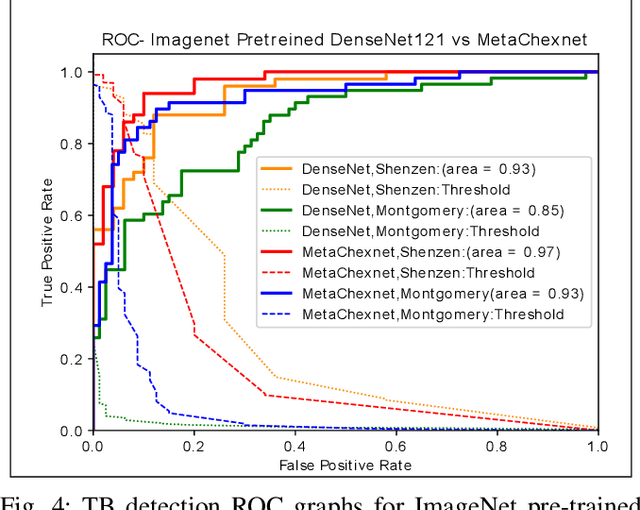
Abstract:The use of ImageNet pre-trained networks is becoming widespread in the medical imaging community. It enables training on small datasets, commonly available in medical imaging tasks. The recent emergence of a large Chest X-ray dataset opened the possibility for learning features that are specific to the X-ray analysis task. In this work, we demonstrate that the features learned allow for better classification results for the problem of Tuberculosis detection and enable generalization to an unseen dataset. To accomplish the task of feature learning, we train a DenseNet-121 CNN on 112K images from the ChestXray14 dataset which includes labels of 14 common thoracic pathologies. In addition to the pathology labels, we incorporate metadata which is available in the dataset: Patient Positioning, Gender and Patient Age. We term this architecture MetaChexNet. As a by-product of the feature learning, we demonstrate state of the art performance on the task of patient Age \& Gender estimation using CNN's. Finally, we show the features learned using ChestXray14 allow for better transfer learning on small-scale datasets for Tuberculosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge