H. A. W. M. Tiddens
Automatic airway segmentation from Computed Tomography using robust and efficient 3-D convolutional neural networks
Mar 30, 2021
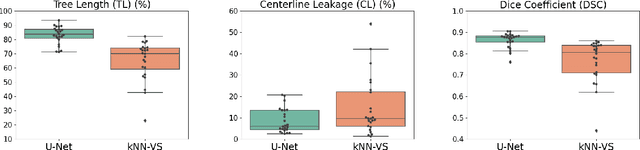
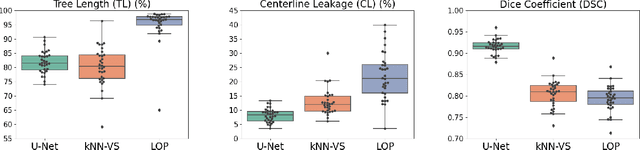
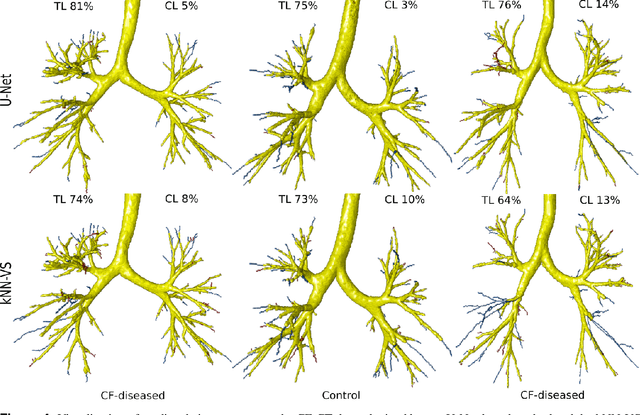
Abstract:This paper presents a fully automatic and end-to-end optimised airway segmentation method for thoracic computed tomography, based on the U-Net architecture. We use a simple and low-memory 3D U-Net as backbone, which allows the method to process large 3D image patches, often comprising full lungs, in a single pass through the network. This makes the method simple, robust and efficient. We validated the proposed method on three datasets with very different characteristics and various airway abnormalities: i) a dataset of pediatric patients including subjects with cystic fibrosis, ii) a subset of the Danish Lung Cancer Screening Trial, including subjects with chronic obstructive pulmonary disease, and iii) the EXACT'09 public dataset. We compared our method with other state-of-the-art airway segmentation methods, including relevant learning-based methods in the literature evaluated on the EXACT'09 data. We show that our method can extract highly complete airway trees with few false positive errors, on scans from both healthy and diseased subjects, and also that the method generalizes well across different datasets. On the EXACT'09 test set, our method achieved the second highest sensitivity score among all methods that reported good specificity.
Automatic Airway Segmentation in chest CT using Convolutional Neural Networks
Aug 14, 2018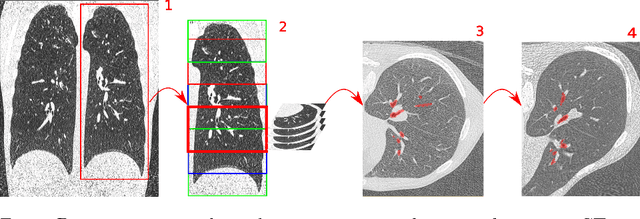

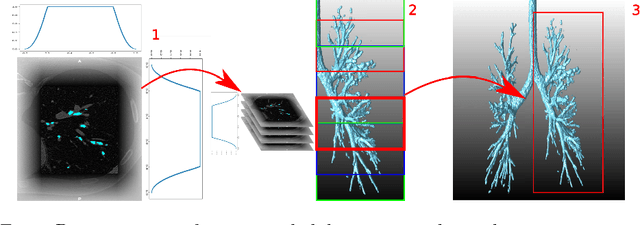
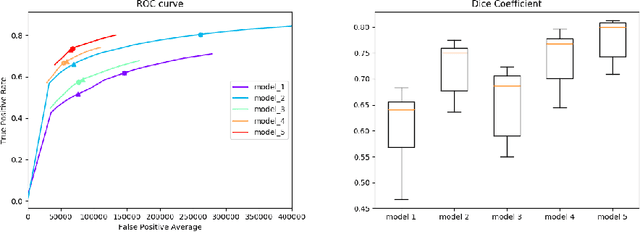
Abstract:Segmentation of the airway tree from chest computed tomography (CT) images is critical for quantitative assessment of airway diseases including bronchiectasis and chronic obstructive pulmonary disease (COPD). However, obtaining an accurate segmentation of airways from CT scans is difficult due to the high complexity of airway structures. Recently, deep convolutional neural networks (CNNs) have become the state-of-the-art for many segmentation tasks, and in particular the so-called Unet architecture for biomedical images. However, its application to the segmentation of airways still remains a challenging task. This work presents a simple but robust approach based on a 3D Unet to perform segmentation of airways from chest CTs. The method is trained on a dataset composed of 12 CTs, and tested on another 6 CTs. We evaluate the influence of different loss functions and data augmentation techniques, and reach an average dice coefficient of 0.8 between the ground-truth and our automated segmentations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge