Gonzalo Maso Talou
Synthesizing Affective Neurophysiological Signals Using Generative Models: A Review Paper
Jun 05, 2023



Abstract:The integration of emotional intelligence in machines is an important step in advancing human-computer interaction. This demands the development of reliable end-to-end emotion recognition systems. However, the scarcity of public affective datasets presents a challenge. In this literature review, we emphasize the use of generative models to address this issue in neurophysiological signals, particularly Electroencephalogram (EEG) and Functional Near-Infrared Spectroscopy (fNIRS). We provide a comprehensive analysis of different generative models used in the field, examining their input formulation, deployment strategies, and methodologies for evaluating the quality of synthesized data. This review serves as a comprehensive overview, offering insights into the advantages, challenges, and promising future directions in the application of generative models in emotion recognition systems. Through this review, we aim to facilitate the progression of neurophysiological data augmentation, thereby supporting the development of more efficient and reliable emotion recognition systems.
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021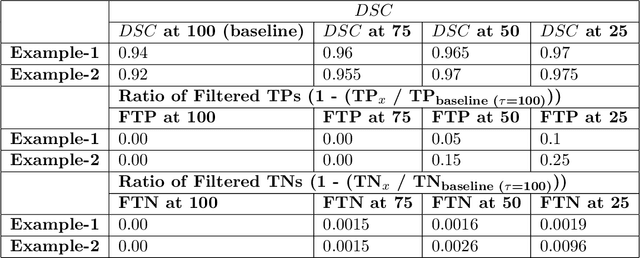
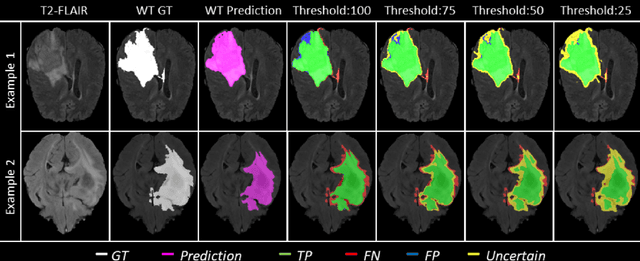
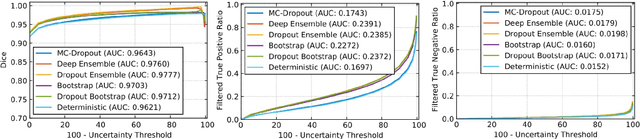
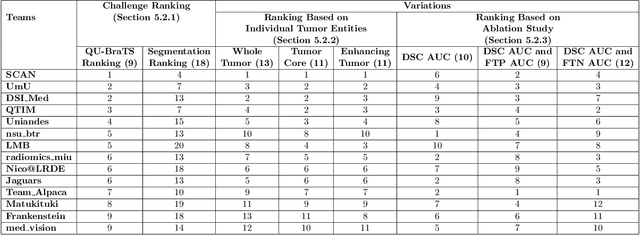
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge