Gardave S Bhumbra
Quantal synaptic dilution enhances sparse encoding and dropout regularisation in deep networks
Sep 28, 2020
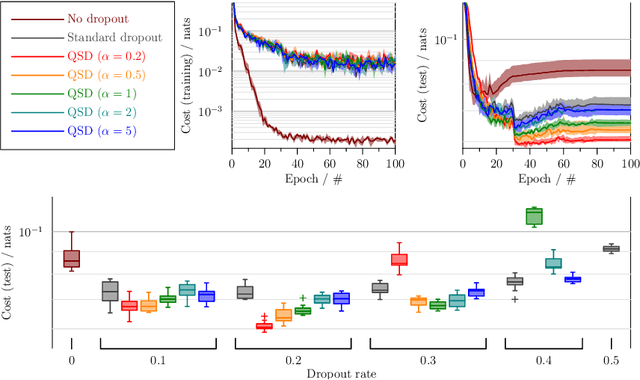
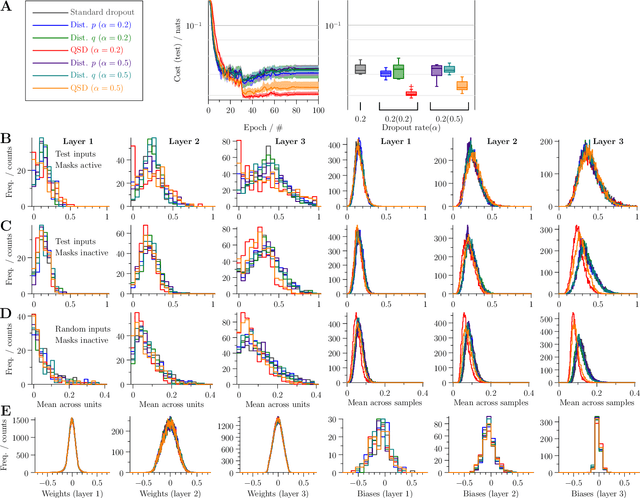
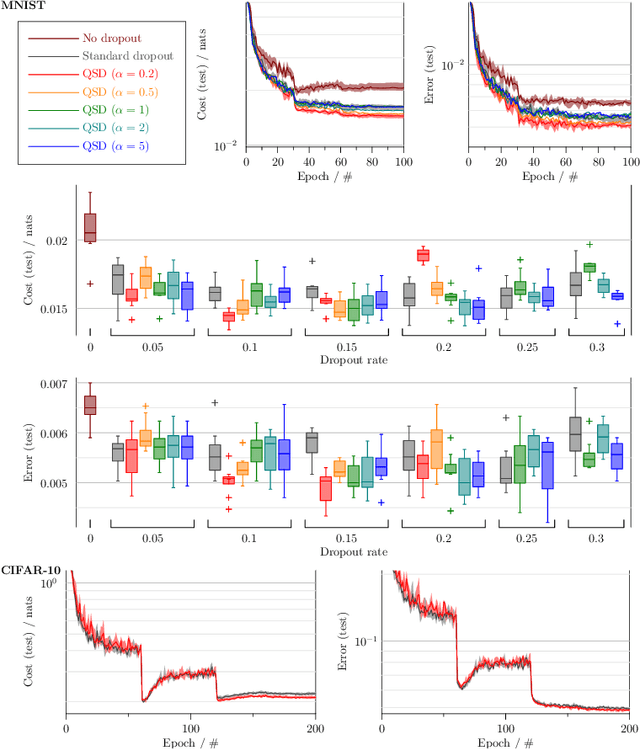
Abstract:Dropout is a technique that silences the activity of units stochastically while training deep networks to reduce overfitting. Here we introduce Quantal Synaptic Dilution (QSD), a biologically plausible model of dropout regularisation based on the quantal properties of neuronal synapses, that incorporates heterogeneities in response magnitudes and release probabilities for vesicular quanta. QSD outperforms standard dropout in ReLU multilayer perceptrons, with enhanced sparse encoding at test time when dropout masks are replaced with identity functions, without shifts in trainable weight or bias distributions. For convolutional networks, the method also improves generalisation in computer vision tasks with and without inclusion of additional forms of regularisation. QSD also outperforms standard dropout in recurrent networks for language modelling and sentiment analysis. An advantage of QSD over many variations of dropout is that it can be implemented generally in all conventional deep networks where standard dropout is applicable.
Deep learning improved by biological activation functions
May 18, 2018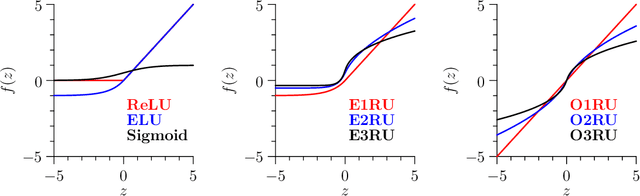
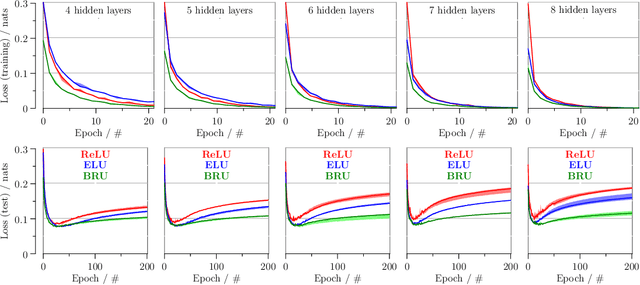
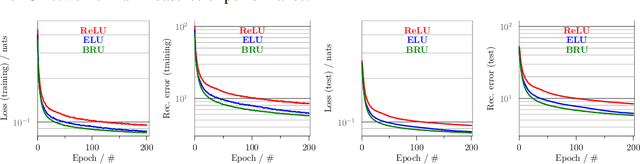
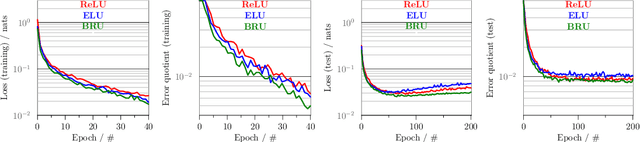
Abstract:`Biologically inspired' activation functions, such as the logistic sigmoid, have been instrumental in the historical advancement of machine learning. However in the field of deep learning, they have been largely displaced by rectified linear units (ReLU) or similar functions, such as its exponential linear unit (ELU) variant, to mitigate the effects of vanishing gradients associated with error back-propagation. The logistic sigmoid however does not represent the true input-output relation in neuronal cells under physiological conditions. Here, bionodal root unit (BRU) activation functions are introduced, exhibiting input-output non-linearities that are substantially more biologically plausible since their functional form is based on known biophysical properties of neuronal cells. In order to evaluate the learning performance of BRU activations, deep networks are constructed with identical architectures except differing in their transfer functions (ReLU, ELU, and BRU). Multilayer perceptrons, stacked auto-encoders, and convolutional networks are used to test supervised and unsupervised learning based on the MNIST and CIFAR-10/100 datasets. Comparisons of learning performance, quantified using loss and error measurements, demonstrate that bionodal networks both train faster than their ReLU and ELU counterparts and result in the best generalised models even in the absence of formal regularisation. These results therefore suggest that revisiting the detailed properties of biological neurones and their circuitry might prove invaluable in the field of deep learning for the future.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge