Gabriel F. Barros
Enhancing Dynamic Mode Decomposition Workflow with In-Situ Visualization and Data Compression
Aug 16, 2022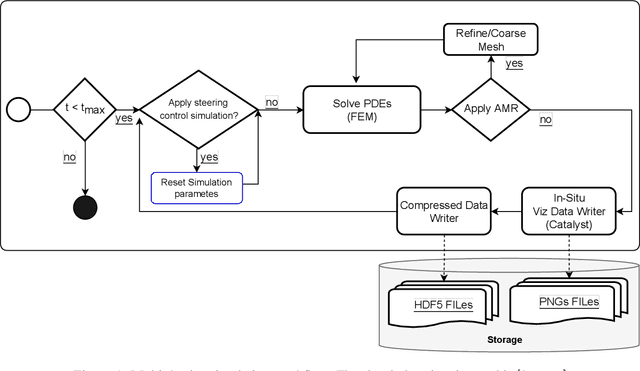

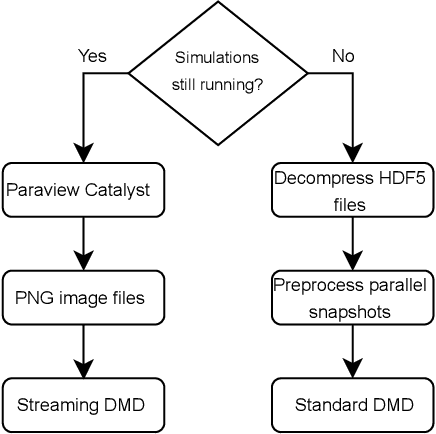
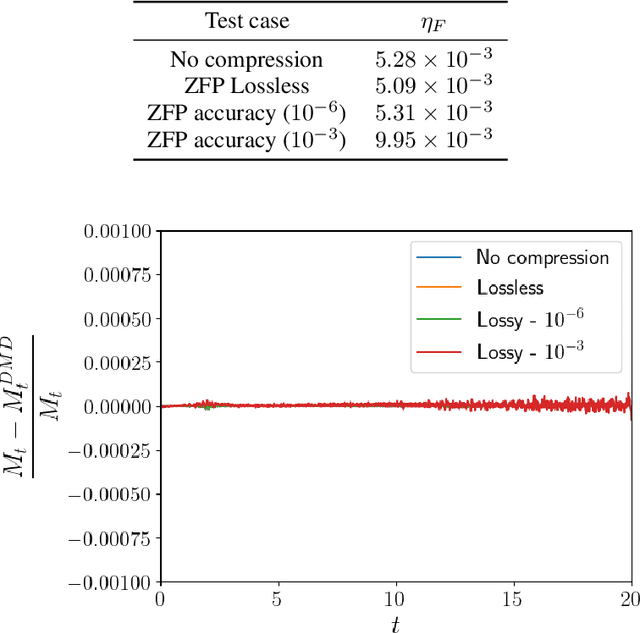
Abstract:Modern computational science and engineering applications are being improved by the advances in scientific machine learning. Data-driven methods such as Dynamic Mode Decomposition (DMD) can extract coherent structures from spatio-temporal data generated from dynamical systems and infer different scenarios for said systems. The spatio-temporal data comes as snapshots containing spatial information for each time instant. In modern engineering applications, the generation of high-dimensional snapshots can be time and/or resource-demanding. In the present study, we consider two strategies for enhancing DMD workflow in large numerical simulations: (i) snapshots compression to relieve disk pressure; (ii) the use of in situ visualization images to reconstruct the dynamics (or part of) in runtime. We evaluate our approaches with two 3D fluid dynamics simulations and consider DMD to reconstruct the solutions. Results reveal that snapshot compression considerably reduces the required disk space. We have observed that lossy compression reduces storage by almost $50\%$ with low relative errors in the signal reconstructions and other quantities of interest. We also extend our analysis to data generated on-the-fly, using in-situ visualization tools to generate image files of our state vectors during runtime. On large simulations, the generation of snapshots may be slow enough to use batch algorithms for inference. Streaming DMD takes advantage of the incremental SVD algorithm and updates the modes with the arrival of each new snapshot. We use streaming DMD to reconstruct the dynamics from in-situ generated images. We show that this process is efficient, and the reconstructed dynamics are accurate.
Coupled and Uncoupled Dynamic Mode Decomposition in Multi-Compartmental Systems with Applications to Epidemiological and Additive Manufacturing Problems
Oct 12, 2021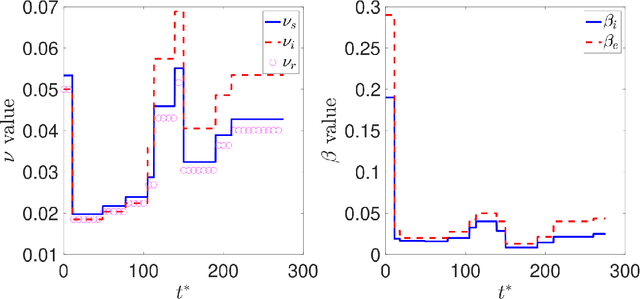
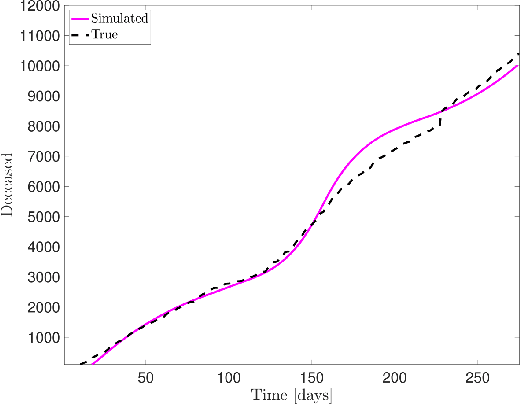
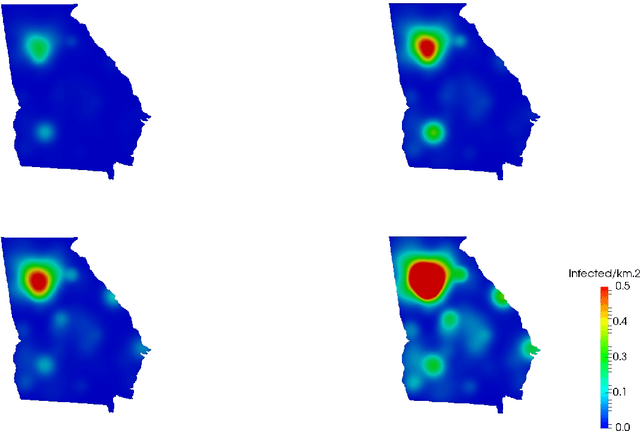
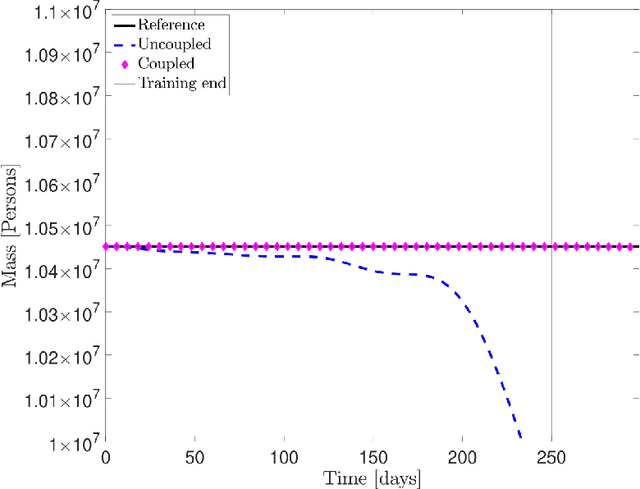
Abstract:Dynamic Mode Decomposition (DMD) is an unsupervised machine learning method that has attracted considerable attention in recent years owing to its equation-free structure, ability to easily identify coherent spatio-temporal structures in data, and effectiveness in providing reasonably accurate predictions for certain problems. Despite these successes, the application of DMD to certain problems featuring highly nonlinear transient dynamics remains challenging. In such cases, DMD may not only fail to provide acceptable predictions but may indeed fail to recreate the data in which it was trained, restricting its application to diagnostic purposes. For many problems in the biological and physical sciences, the structure of the system obeys a compartmental framework, in which the transfer of mass within the system moves within states. In these cases, the behavior of the system may not be accurately recreated by applying DMD to a single quantity within the system, as proper knowledge of the system dynamics, even for a single compartment, requires that the behavior of other compartments is taken into account in the DMD process. In this work, we demonstrate, theoretically and numerically, that, when performing DMD on a fully coupled PDE system with compartmental structure, one may recover useful predictive behavior, even when DMD performs poorly when acting compartment-wise. We also establish that important physical quantities, as mass conservation, are maintained in the coupled-DMD extrapolation. The mathematical and numerical analysis suggests that DMD may be a powerful tool when applied to this common class of problems. In particular, we show interesting numerical applications to a continuous delayed-SIRD model for Covid-19, and to a problem from additive manufacturing considering a nonlinear temperature field and the resulting change of material phase from powder, liquid, and solid states.
Dynamic Mode Decomposition in Adaptive Mesh Refinement and Coarsening Simulations
Apr 28, 2021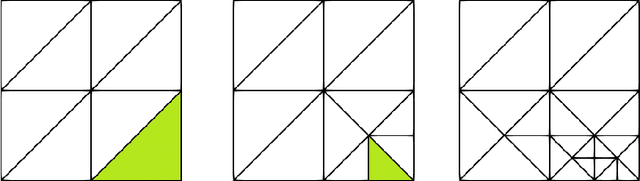
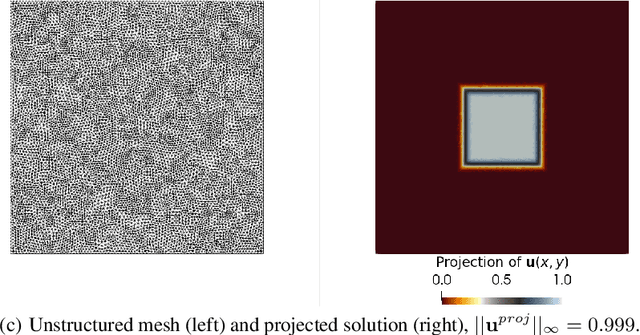

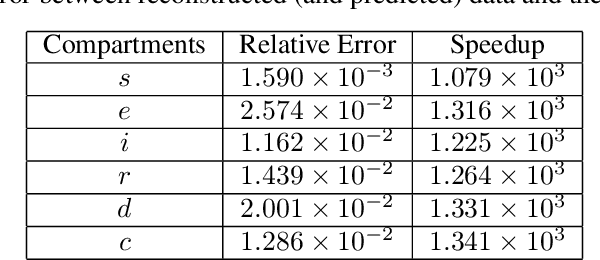
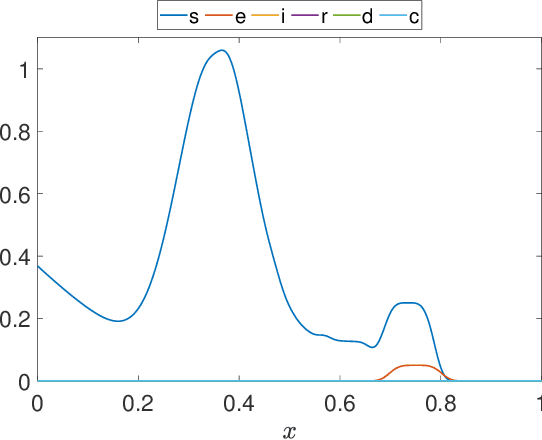
Abstract:Dynamic Mode Decomposition (DMD) is a powerful data-driven method used to extract spatio-temporal coherent structures that dictate a given dynamical system. The method consists of stacking collected temporal snapshots into a matrix and mapping the nonlinear dynamics using a linear operator. The standard procedure considers that snapshots possess the same dimensionality for all the observable data. However, this often does not occur in numerical simulations with adaptive mesh refinement/coarsening schemes (AMR/C). This paper proposes a strategy to enable DMD to extract features from observations with different mesh topologies and dimensions, such as those found in AMR/C simulations. For this purpose, the adaptive snapshots are projected onto the same reference function space, enabling the use of snapshot-based methods such as DMD. The present strategy is applied to challenging AMR/C simulations: a continuous diffusion-reaction epidemiological model for COVID-19, a density-driven gravity current simulation, and a bubble rising problem. We also evaluate the DMD efficiency to reconstruct the dynamics and some relevant quantities of interest. In particular, for the SEIRD model and the bubble rising problem, we evaluate DMD's ability to extrapolate in time (short-time future estimates).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge