G. Varoquaux
Confidence intervals for performance estimates in 3D medical image segmentation
Jul 21, 2023Abstract:Medical segmentation models are evaluated empirically. As such an evaluation is based on a limited set of example images, it is unavoidably noisy. Beyond a mean performance measure, reporting confidence intervals is thus crucial. However, this is rarely done in medical image segmentation. The width of the confidence interval depends on the test set size and on the spread of the performance measure (its standard-deviation across of the test set). For classification, many test images are needed to avoid wide confidence intervals. Segmentation, however, has not been studied, and it differs by the amount of information brought by a given test image. In this paper, we study the typical confidence intervals in medical image segmentation. We carry experiments on 3D image segmentation using the standard nnU-net framework, two datasets from the Medical Decathlon challenge and two performance measures: the Dice accuracy and the Hausdorff distance. We show that the parametric confidence intervals are reasonable approximations of the bootstrap estimates for varying test set sizes and spread of the performance metric. Importantly, we show that the test size needed to achieve a given precision is often much lower than for classification tasks. Typically, a 1% wide confidence interval requires about 100-200 test samples when the spread is low (standard-deviation around 3%). More difficult segmentation tasks may lead to higher spreads and require over 1000 samples.
Comparing distributions: $\ell_1$ geometry improves kernel two-sample testing
Oct 01, 2019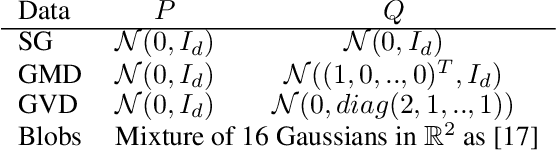
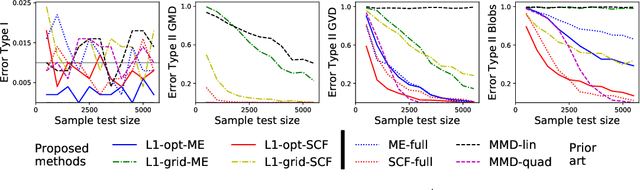
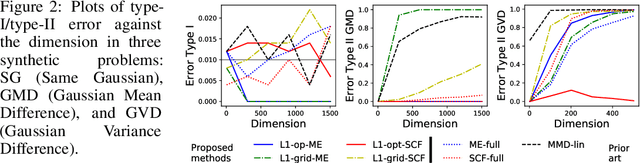
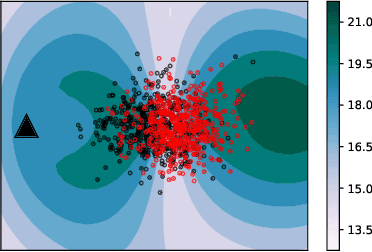
Abstract:Are two sets of observations drawn from the same distribution? This problem is a two-sample test. Kernel methods lead to many appealing properties. Indeed state-of-the-art approaches use the $L^2$ distance between kernel-based distribution representatives to derive their test statistics. Here, we show that $L^p$ distances (with $p\geq 1$) between these distribution representatives give metrics on the space of distributions that are well-behaved to detect differences between distributions as they metrize the weak convergence. Moreover, for analytic kernels, we show that the $L^1$ geometry gives improved testing power for scalable computational procedures. Specifically, we derive a finite dimensional approximation of the metric given as the $\ell_1$ norm of a vector which captures differences of expectations of analytic functions evaluated at spatial locations or frequencies (i.e, features). The features can be chosen to maximize the differences of the distributions and give interpretable indications of how they differs. Using an $\ell_1$ norm gives better detection because differences between representatives are dense as we use analytic kernels (non-zero almost everywhere). The tests are consistent, while much faster than state-of-the-art quadratic-time kernel-based tests. Experiments on artificial and real-world problems demonstrate improved power/time tradeoff than the state of the art, based on $\ell_2$ norms, and in some cases, better outright power than even the most expensive quadratic-time tests.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge