Frosti Palsson
Multispectral and Hyperspectral Image Fusion Using a 3-D-Convolutional Neural Network
Jun 16, 2017
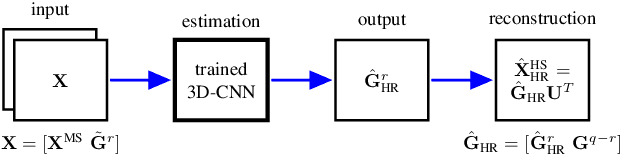
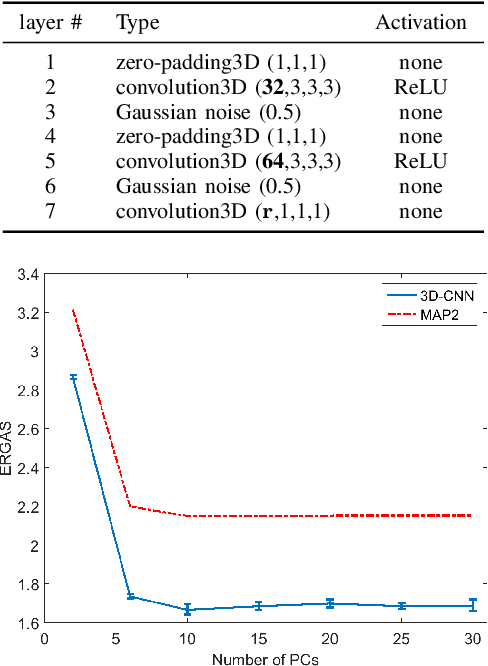
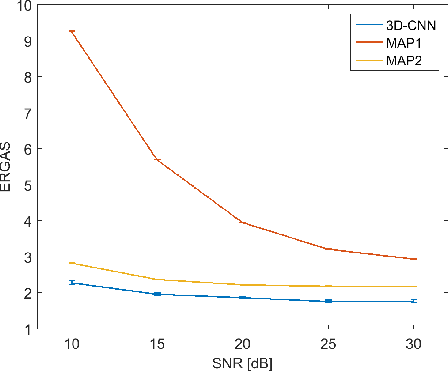
Abstract:In this paper, we propose a method using a three dimensional convolutional neural network (3-D-CNN) to fuse together multispectral (MS) and hyperspectral (HS) images to obtain a high resolution hyperspectral image. Dimensionality reduction of the hyperspectral image is performed prior to fusion in order to significantly reduce the computational time and make the method more robust to noise. Experiments are performed on a data set simulated using a real hyperspectral image. The results obtained show that the proposed approach is very promising when compared to conventional methods. This is especially true when the hyperspectral image is corrupted by additive noise.
Classification of Big Data with Application to Imaging Genetics
May 16, 2016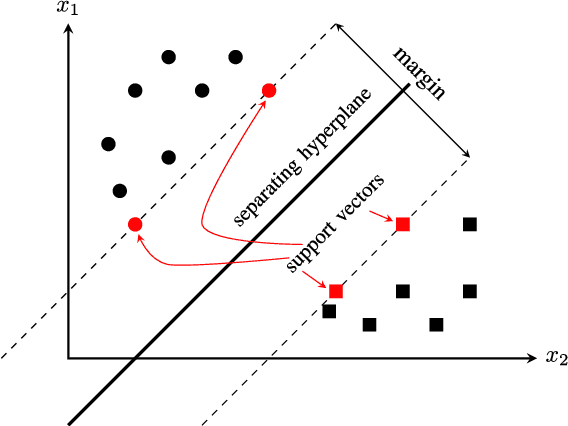
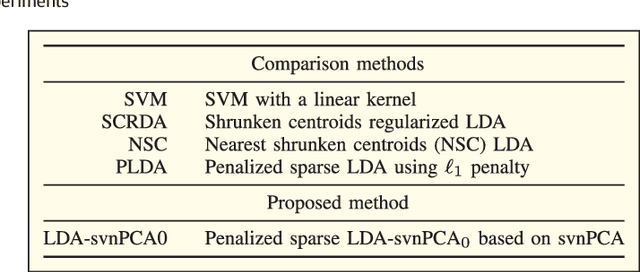
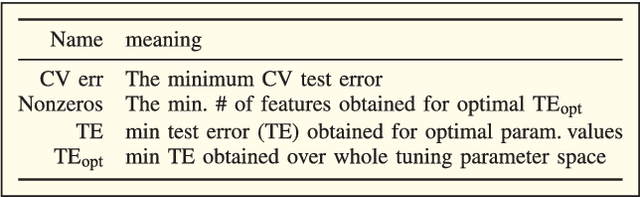


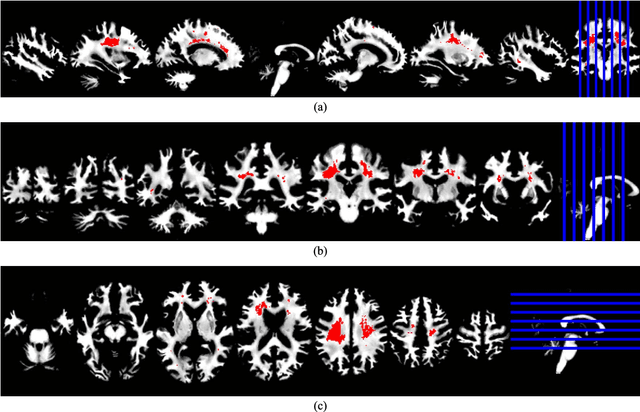
Abstract:Big data applications, such as medical imaging and genetics, typically generate datasets that consist of few observations n on many more variables p, a scenario that we denote as p>>n. Traditional data processing methods are often insufficient for extracting information out of big data. This calls for the development of new algorithms that can deal with the size, complexity, and the special structure of such datasets. In this paper, we consider the problem of classifying p>>n data and propose a classification method based on linear discriminant analysis (LDA). Traditional LDA depends on the covariance estimate of the data, but when p>>n the sample covariance estimate is singular. The proposed method estimates the covariance by using a sparse version of noisy principal component analysis (nPCA). The use of sparsity in this setting aims at automatically selecting variables that are relevant for classification. In experiments, the new method is compared to state-of-the art methods for big data problems using both simulated datasets and imaging genetics datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge